Genome. Gene Expression Profiling Analysis Reveals Fur Development in Rex Rabbits (Oryctolagus cuniculus)
|
|
|
- Νατάσσα Καραμήτσος
- 5 χρόνια πριν
- Προβολές:
Transcript
1 Gene Expression Profiling Analysis Reveals Fur Development in Rex Rabbits (Oryctolagus cuniculus) Journal: Manuscript ID gen r2 Manuscript Type: Article Date Submitted by the Author: 31-Jul-2017 Complete List of Authors: Zhao, Bohao; Yangzhou University Chen, Yang; Yangzhou University Yan, Xiaorong ; Yangzhou University Hao, Ye; Yangzhou University Zhu, Jie; Yangzhou University Weng, Qiiaoqing; Zhejiang Yuyao Xinnong Rabbit Industry Co., Ltd. Wu, Xinsheng; Yangzhou University, College of Animal Science and Technology Is the invited manuscript for consideration in a Special Issue? : This submission is not invited Keyword: Chinchilla rex rabbit, fur development, key gene, transcriptome
2 Page 1 of Gene Expression Profiling Analysis Reveals Fur Development in Rex Rabbits (Oryctolagus cuniculus) BoHao Zhao 1, Yang Chen 1, XiaoRong Yan 1, Ye Hao 1, Jie Zhu 1, QiaoQing Weng 2, and XinSheng Wu 1 * 1 The Key Laboratory of Animal Genetics & Breeding and Molecular Design of Jiangsu Province, College of Animal Science and Technology, Yangzhou University, Yangzhou , P.R. China.; 2 Zhejiang Yuyao Xinnong Rabbit Industry Co., Ltd., Yuyao, Zhejiang , China *Corresponding author xswu@yzu.edu.cn 10 1
3 Page 2 of Abstract Fur is an important economic trait in rabbits. The identification of genes that influence fur development and knowledge regarding the actions of these genes provides useful tools for improving fur quality. However, the mechanism of fur development is unclear. To obtain candidate genes related to fur development, the transcriptomes of tissues from backs and bellies of Chinchilla rex rabbits were compared. Of the genes analyzed, 336 showed altered expression in the two groups (285 upregulated and 51 downregulated), P 0.05, fold-change 2 or 0.5). Using GO and KEGG to obtain gene classes that were differentially enriched, we found several genes to be involved in many important biological processes. In addition, we identified several signaling pathways involved in fur development, including the Wnt and MAPK signaling pathways, revealing mechanisms of skin and hair follicle development, and epidermal cell and keratinocytes differentiation. The obtained rabbit transcriptome and differentially expressed gene profiling data provided comprehensive gene expression information for SFRP2, FRZB, CACNG1, SLC25A4 and SLC16A3. To validate the RNA-seq data, the expression levels of eight differentially expressed genes involved in fur development were confirmed by qrt-pcr. The results of rabbit transcriptomic profiling provide a basis for understanding the molecular mechanisms of fur development. Keywords Chinchilla rex rabbit, fur development, key gene, transcriptome 32 2
4 Page 3 of Introduction The Chinchilla rex rabbit is an important rabbit breed with varied natural coat colors; consumers highly appreciate the properties of rex furs, such as beauty, softness, color, lightness, and warmth retention (Pan et al. 2015). The characteristics of Chinchilla rex rabbit fur differs between the back and belly, especially the length and diameter of the wools (Tao 2010). In recent years, many studies have revealed the mechanisms of skin and fur development. RNA-seq was used to explore the mechanisms of keratinocyte development in mouse skin, and transcription factor (TF) p63 was found to be highly 42 expressed in stratified epithelia, which affected the epidermal phenotype (Rizzo et al ). Many genes involved in skin development, including those for transcription factors and growth factors, have been identified in rex rabbits with the plaice phenotype (Pan et al. 2015). In cashmere goats, genes related to hair follicle development and cycling were identified in anagen, catagen and telogen stages by transcriptomic investigation of fur development (Geng et al. 2013). It is generally known that fur development is influenced by many factors, including the proliferation of keratinocytes, development of the epidermis and hair follicle (HF) morphogenesis (Danilenko et al. 1995). Multiple genes involved in HF morphogenesis, regulation of proliferation, differentiation and migration of skin are controlled by members of the Wnt signaling pathway, such as the frizzled and secreted frizzled-related protein (SFRP) families (Ehrlund et al. 2013; Kim and Yoon 2014). Epidermal growth factor is regulated by the MAPK/ERK pathway and plays a vital role in the animal skin 3
5 Page 4 of development, enhancing epidermal growth and keratinization, directly stimulating the proliferation of epidermal cells and promoting keratinocyte proliferation and migration. However, fur characteristics are different at different parts of an animal, and the mechanism of fur development regulation is still unclear in rabbits. Rabbit genome sequencing has been used to study the polygene-related phenotypic changes during rabbit domestication (Carneiro et al. 2014) and differential gene expression in animal skin between anagen and telogen was shown by transcriptome sequencing (Xu et al. 2013). In this study, the skin from the backs and bellies of Chinchilla rex rabbits was collected, and gene expression profiling was used to obtain 64 the differentially expressed genes related to the fur development. After functional annotation, enrichment analysis and assessment of biological functions, candidate genes were identified. These key genes were verified by quantitative real-time PCR. The results obtained serve to improve our understanding of fur development and the differential expression profiles of the candidate genes enable us to clarify the mechanisms of fur development, providing a valuable theoretical basis for further research on the hair and fur of animals Materials and methods Ethics statement All animal experiments were reviewed and approved by the Institutional Animal Care and Use Committee of the School of Animal Science and Technology, Yangzhou University, and performed in accordance with the Regulations for the Administration 4
6 Page 5 of of Affairs Concerning Experimental Animals (China, 1988) and the Standards for the administration of experimental practices (Jiangsu, China, 2008). All surgery was performed according to recommendations proposed by the European Commission (1997), and all efforts were made to minimize suffering of the animals. Tissue collection The Chinchilla rex rabbits used in the experiments were obtained from Zhejiang Yuyao Xinnong Rabbit Co., Ltd. During our experiments, rabbits were raised in a controlled environment and had free access to water and food. All rabbits were housed in a suitable, clean and disease free environment, and a secure cage. The health of the 86 rabbits were monitored twice a day (7 am and 6 pm) and recorded. Three healthy, day-old rabbits with the same fur traits were evaluated. Fur on the back (B group) and belly (F group) were different. Three biological samples were taken for each of the two groups (one sample of back and belly fur from each rabbit) to ensure the same genetic background and fur phenotype in each group. After transfer to the laboratory, skin tissue samples (1.5 cm 2 ) were collected from the back and belly of each rabbit. Animals were anesthetized with by injection with 0.7% sodium pentobarbital solution into the ear vein of the rabbits; in order to prevent bacterial infection iodine solution was smeared on the resultant lesion. Fur was removed from the surface, and then the skin was cut into pieces. The pieces were placed in tubes containing RNase, immediately preserved in liquid nitrogen and stored at -70 C until use in subsequent experiments. RNA extraction, cdna library construction, and Illumina sequencing 5
7 Page 6 of Total RNA was extracted following the manufacturer s instructions using the mirvana mirna isolation kit (Ambion); the integrity of the RNA was determined with an Agilent Bioanalyzer 2100 (Agilent technologies, Santa Clara, CA, US) to obtain a RNA Integrity Number (RIN). An RNeasy micro kit (Cat#74004, QIAGEN, GmBH, Germany) was used to further purify the qualified total RNA, and DNA was removed with the RNase-Free DNase set (QIAGEN, GmBH, Germany). RNA quality was monitored using NanoDrop ND-1000 and Agilent Bioanalyzer After RNA extraction and purification, 3 µg RNA was used for construction of the back and belly cdna libraries. Ribosomal RNA (rrna) was depleted from the total RNA and the 108 remaining RNA was subsequently fragmented. These steps were followed by first and second cdna strand synthesis, end repair, 3'-end adenylation, adapter ligation, and enrichment of the cdna templates. Finally, the library concentration was determined using a Qubit 2.0 fluorometer and a Qubit dsdna HS kit (Invitrogen). Cluster generation was completed the sample library, and the first primers hybridized to cbot matched the Illumina HiSeq 2500 platform. After cluster generation, the sequencing reagent was prepared according to the HiSeq 2500 user guide using paired-end technology. Sequencing was controlled by data collection software (Illumina, San Francisco, USA) and the data were analyzed in real time. Transcriptome mapping and analysis of differentially expressed genes The cdna library was sequenced using an Illumina HiSeq 2500 sequencing platform. Original image files were obtained, and bases were called and filtered, after which the results were stored in fastq format. The original sequencing reads were used 6
8 Page 7 of for transcriptome sequencing analysis. As the raw reads contained sequences of low quality, clean reads were obtained using fastx (version: ), which included the removal of low-quality sequence fragments, 3 end bases that were 10 below the quality score of Q=10 (Q =-10log error_ratio ), adapter sequences, reads containing runs of N s blurs, and any sequences shorter than 20 nucleotides with low overall quality. We then used the TopHat algorithm (version:2.0.9) (Trapnell et al. 2009) to map the clean reads to the Oryctolagus cuniculus genome by spliced mapping, allowing two bases of mispairing and multiple hits less than or equal to two, according to Ensembl OryCun2.0. Gene expression was quantified using Cufflinks (version:2.1.1) (Trapnell et al. 2010). Additionally, the fragments per kilobase of exon model per million 131 mapped reads (FPKM) were defined as follows: transcript reads FPKM= transcript length total mapped reads in run The fold-change and Fisher-test were used to analyze the differentially expressed genes that were selected with a false discovery rate (FDR) of less than 0.05 and a fold-change greater than or equal to 2 or less than or equal to 0.5. Gene annotation and network analysis The differentially expressed genes obtained from the two skin types were used for functional annotation and mapped to Gene Ontology (GO; terms and Kyoto Encyclopedia of Genes and s (KEGG; pathways to identify pathways potentially associated with skin development. P values less than or equal to 0.05 and FDR less than or equal to 0.05 were considered statistically significant. After comparing the hypergeometrics 7
9 Page 8 of with the background of the genome, we screened the GO terms looking for significant enrichment of the differentially expressed genes. P values correspond to differential gene expression after Bonferroni correction, and corrected P values less than or equal to 0.05 were the threshold for significance of differences in gene expression. The STRING database was used to perform network analysis and the union of all differentially expressed genes between the B and F groups was used to build the network. Quantitative real-time PCR confirmation of differentially expressed genes To validate the sequencing data, eight known differentially expressed candidate 151 genes were selected for validation by qrt-pcr, which was performed on a real-time PCR system (Applied Biosystems) using the AceQ qpcr SYBR Green Master Mix (Vazyme) according to the manufacturer s instructions. The primer sequences are listed in Table 1. We chose the rabbit glyceraldehyde-3-phosphate dehydrogenase (GAPDH) gene as an internal control. The results of the experiments were normalized to the expression of the constitutive GAPDH gene. Quantitative 157 variation and relative fold-change were calculated based on the 2 Ct method (Schmittgen and Livak 2008). Error bars represent the mean ± S.D. as determined using GraphPad Prism 5, and a paired t-test was performed to test significant differences between the two groups using SPSS Western blotting Protein lysates from six skin samples were obtained using RIPA Lysis Buffer (PPLYGEN). Protein concentrations were determined with the Enhanced BCA Protein 8
10 Page 9 of Kit (Beyotime). The protein lysates were diluted to 0.5µg/µL, and 2.5µg protein was detected and analyzed with the Wes automated Western blotting system (Protein Simple) (Harris 2015). The following antibodies were used: 1:100 Anti-GAPDH mouse monoclonal antibody (Abcam), 1:50 Anti-SFRP2 monoclonal antibody (Sangon Biotech). 169 Results 170 Results of transcriptome sequencing 171 To evaluate whether the RNA-seq data were sufficient for further analysis, their 172 global quality was firstly assessed. After trimming clean reads were generated with a ratio of greater than 85% for each sample (Table 2); and the total clean reads that mapped to the rabbit genome had mapping ratios greater than 80% (Table 3). All reads were deposited in the Short Read Archive (SRA) of the National Center for Biotechnology Information (NCBI) under the accession number SRR We then obtained the mapped read distribution, which showed whether the reads mapped to genes, coding regions, splice sequences, intron sequences, intergenic regions, or non-coding regions (5 UTR and 3 UTR, non-coding RNA regions) (Figure 1). Our results showed that we were able to detect 17,327 expressed genes, including 8445 upregulated genes and 8882 downregulated genes (Table S1). To identify the key differentially expressed genes, the parameters of a fold-change greater than or equal to 2 or less than or equal to 0.5 and FDR less than or equal to 0.05 were used to further select the significantly differentially expressed genes. These 9
11 Page 10 of cut-offs identified 285 upregulated genes and 51 downregulated genes (Table S2). Volcano plots of differentially expressed genes were constructed to explore the relationship between fold-change and significance as shown in Figure 2. This analysis identified 336 genes that were significantly differentially expressed between the back and belly regions of the rex rabbits. 190 Functional annotation of differentially expressed genes GO functional enrichment and KEGG pathway analyses were performed to determine the functions of the differentially expressed genes. According to the GO 193 analysis, categories of biological process (n=563), molecular function (n=204), and cellular component (n=132) contained 214 differentially expressed genes (Table S3), which were all considered statistically significant (Figure S1). In the biological process category, most GO terms focused on ion activity, development, morphogenesis, cellular processes, biological regulation, and metabolic processes. In the molecular function category, a high proportion of the GO terms were related to binding, activity regulation, enzymatic activity, and ionic equilibrium. In the cellular component category, we found the terms Z disc, I band, muscle contraction, myofibril, titin binding, and M band sarcoplasmic reticulum showed the greatest enrichment of differentially expressed genes. These GO terms revealed the biological functions of the genes. According to the GO analysis, many GO terms were related to the functions of skin and epithelial cells, such as epithelial cell proliferation, regulation of cell proliferation, skin development and regulation of cell development. This suggests that 10
12 Page 11 of the genes in these GO categories may play roles in fur development. We then identified the fur development related genes from the GO terms and their distribution in the molecular function categories (Figure 3). Meanwhile, KEGG pathway analysis identified 66 differentially expressed genes (Table S4), and results of the KEGG enrichment are shown in Figure S2. Analysis of all the KEGG signaling pathways showed a definite relationship between several pathways, such as the Wnt and MAPK signaling pathways. We identified the SFRP2 and FRZB genes from the Wnt signaling pathway (Figure S3) and calcium channel, voltage-dependent, beta 1 subunit (CACNB1), voltage-dependent calcium channel 215 gamma-1 subunit (CACNG1), and calcium channel, voltage-dependent, L type, alpha S subunit (CACNA1S) from the MAPK signaling pathway. These genes were found to influence HF and skin development related signaling pathways, suggesting that these genes may play a role in fur development (Chu et al. 2014; Fuchs and Raghavan 2002; Kim and Yoon 2014). 220 Network analysis of differentially expressed gene interactions To explore interactions of the differentially expressed genes, RNA-seq data was used to construct a differentially expressed gene interaction network. We identified interacting partners of the differentially expressed genes using the STRING database, which predicts functional associations between proteins (Figure 4). 225 Validation of differentially expressed genes 226 To confirm our RNA-seq results, qrt-pcr was performed on samples from the 11
13 Page 12 of backs and bellies of the rabbits. Eight target genes were selected and specific primers were designed for validation. As shown in Figure 5A, we found that FRZB, SFRP2, DUSP26, PTP4A3, EN1, and CACNA1S were upregulated, and HBB1 and MRPL36 were downregulated by qrt-pcr. The protein levels of SFRP2 among the six samples were further assessed by Western blotting, and we found the highest expression of the SFRP2 protein in the back group (Figure 5B). These data indicate that the results from the transcriptome study are consistent with the overall changes in expression of the differentially expressed genes Discussion The characteristics of fur are different on different body parts of Chinchilla rex rabbits, especially the back and belly (Tao 2010). Fur development is regulated by many factors, such as rearing conditions, nutrition, environment, and gene regulation (Harkness et al. 2013; McNitt et al. 2013). Our study revealed several fur development related genes by transcriptomics, and the functional enrichment and pathway analysis of these genes will contribute to our understanding of the mechanisms of fur development. The Wnt signaling pathway plays an indispensable role at various stages during tissue regeneration and skin development (Stoick-Cooper et al. 2007). In this pathway, Wnt proteins are important in conveying inductive signals between the mesenchyme of follicles and the follicular epithelium. As a family of secreted glycoproteins, Wnts 12
14 Page 13 of also play roles in embryonic development and maintenance of homeostasis by regulating migration, differentiation, proliferation and apoptosis of cells in mature tissues (Fujimaki et al. 2015; Millar 2002). Several transcription factors have been identified that regulate the differentiation of keratinocytes via the MAPK signaling pathway (Eckert and Welter 1996), which governs many cellular processes, including cell fate, proliferation, differentiation, homeostasis, and survival in all eukaryotes (Whelan et al. 2012). For example, activator protein I is a keratinocyte transcription factor that plays a crucial role in the regulation of epidermal differentiation and the expression of genes in the MAPK pathway (Briata et al. 1993; Eckert and Welter 1996; 257 Smeyne et al. 1992) Many genes that are related to fur development such as Secreted frizzled-related protein 2 (SFRP2) and Frizzled-related protein (FRZB) are glycoproteins that are involved in the processes of development and disease in diverse cells and tissues (Ezan et al. 2004; Hoang et al. 1996; Kim and Yoon 2014). SFRP2 inhibited mouse keratinocyte proliferation in the catagen phase and was regarded as a Wnt inhibitor in HFs. In the back skin of mice, the Wnt target genes Ccnd1 and C-myc, were shown to have an inverse relationship between the two genes and SFRP2 throughout the HF cycle, while SFRP2 may inhibit the proliferation of keratinocytes in the catagen phase (Kim and Yoon 2014). Moreover, SFRP2 can control cell apoptosis and fate, and mediate regulation of the Wnt pathway that has effects on intestinal epithelial cells (Skah et al. 2015). However, the SFRP2 protein was highly expressed in the back group in our study; suggesting that the protein product of SFRP2 acted as an activator 13
15 Page 14 of in skin development. This means that SFRP2 could play a positive role in controlling epidermal cell and keratinocyte differentiation. FRZB (also called SFRP-3 or Fritz) is a member of secreted frizzled-related protein family, which includes secreted proteins (SFRP1-5) that bind and inhibit Wnts (Ehrlund et al. 2013; Ladher et al. 2000). As an antagonist of Wnt proteins in chick development, FRZB inhibited the activity of Xwnt-8 during the gastrula stages (Ladher et al. 2000). With the activation of downstream targets of the pathway, a knockdown of FRZB upregulated the Wnt/β catenin pathway (Qin et al. 2014). In this study, FRZB was up-regulated in the back group, suggesting that FRZB may promote 279 the growth of HFs and skin development. However many previous studies have regarded FRZB as an inhibitor, although it is probably an activator that influences the target genes in the Wnt pathway. In our study, the fur phenotypes, such as the length and diameter of the wool, were different between the back and belly skin of rex rabbits. These differences may be influenced by the structure of HF and skin, especially the development of epidermal cells and keratinocytes. Bioinformatic analysis revealed that genes in the Wnt signaling pathway may have effects on the development of HF and skin. We also found that SFRP2 mrna and protein were highly expressed in the back group, leading to the longer, thicker and greater density than belly group. The regulation of fur development related genes involves the differential expression of genes associated with many biological signaling pathways. The mechanism of HF and skin development will be clarified by functional studies of the candidate genes. 14
16 Page 15 of Therefore, with RNA sequencing and function analysis, the functions of these key genes, such as SFRP2 and FRZB, should be verified by further study including their function in hair cycle regulation, differentiation of keratinocytes and their role in the Wnt pathway Conclusion Gene expression profiling analysis was used to assess fur development in Chinchilla rex rabbits. This study found several genes associated with HF and skin development, including SFRP2, FRZB, CACNG1, CACNB1, CACNA1S, PTPLA, PTP4A3, TTN, 301 DUSP26, EN1, MT3, SLC25A4 and SLC16A3. The mechanisms regulating fur development are complex, and fur development related genes should be studied further Acknowledgments This work was supported by the Modern Agricultural Industrial System Special Funding (CARS-44-1) and the Priority Academic Program Development of Jiangsu Higher Education Institutions ( ) References Briata, P., D'Anna, F., Franzi, A.T., and Gherzi, R AP-1 activity during normal human keratinocyte differentiation: evidence for a cytosolic modulator of AP-1/DNA binding. Experimental cell research 204(1): Carneiro, M., Rubin, C.-J., Di Palma, F., Albert, F.W., Alföldi, J., Barrio, A.M., 15
17 Page 16 of Pielberg, G., Rafati, N., Sayyab, S., and Turner-Maier, J Rabbit genome analysis reveals a polygenic basis for phenotypic change during domestication. Science 345(6200): Chu, Q., Cai, L., Fu, Y., Chen, X., Yan, Z., Lin, X., Zhou, G., Han, H., Widelitz, R.B., and Chuong, C.-m Dkk2/Frzb in the dermal papillae regulates feather regeneration. Dev Biol 387(2): Danilenko, D.M., Ring, B.D., Yanagihara, D., Benson, W., Wiemann, B., Starnes, C.O., and Pierce, G.F Keratinocyte growth factor is an important endogenous mediator of hair follicle growth, development, and differentiation. Normalization of the nu/nu follicular differentiation defect and amelioration of chemotherapy-induced alopecia. The American journal of pathology 147(1): Eckert, R.L., and Welter, J.F Transcription factor regulation of epidermal keratinocyte gene expression. Mol Biol Rep 23(1): Ehrlund, A., Mejhert, N., Lorente-Cebrián, S., Åström, G., Dahlman, I., Laurencikiene, J., and Rydén, M Characterization of the Wnt inhibitors secreted frizzled-related proteins (SFRPs) in human adipose tissue. The Journal of Clinical Endocrinology & Metabolism 98(3): E503-E508. Ezan, J., Leroux, L., Barandon, L., Dufourcq, P., Jaspard, B., Moreau, C., Allières, C., Daret, D., Couffinhal, T., and Duplàa, C FrzA/sFRP-1, a secreted antagonist of the Wnt-Frizzled pathway, controls vascular cell proliferation in vitro and in vivo. Cardiovasc Res 63(4): Fuchs, E., and Raghavan, S Getting under the skin of epidermal morphogenesis. Nature Reviews Genetics 3(3): Fujimaki, S., Wakabayashi, T., Takemasa, T., Asashima, M., and Kuwabara, T The regulation of stem cell aging by Wnt signaling. Histology and histopathology 30(12): Geng, R., Yuan, C., and Chen, Y Exploring differentially expressed genes by RNA-Seq in cashmere goat (Capra hircus) skin during hair follicle development and cycling. Plos One 8(4): e Harkness, J.E., Turner, P.V., VandeWoude, S., and Wheler, C.L Harkness and 16
18 Page 17 of Wagner's biology and medicine of rabbits and rodents. John Wiley & Sons. Harris, V.M Protein detection by Simple Western analysis. Western Blotting: Methods and Protocols: Hoang, B., Moos, M., Vukicevic, S., and Luyten, F.P Primary structure and tissue distribution of FRZB, a novel protein related to Drosophila frizzled, suggest a role in skeletal morphogenesis. J Biol Chem 271(42): Kim, B.-K., and Yoon, S.K Expression of sfrp2 is increased in catagen of hair follicles and inhibits keratinocyte proliferation. Annals of dermatology 26(1): Ladher, R., Church, V., Allen, S., Robson, L., Abdelfattah, A., Brown, N., Hattersley, G., Rosen, V., Luyten, F., and Dale, L Cloning and expression of the Wnt antagonists Sfrp-2 and Frzb during chick development. Dev Biol 218(2): McNitt, J.I., Lukefahr, S.D., Cheeke, P.R., and Patton, N.M Rabbit production. CABI Millar, S.E Molecular mechanisms regulating hair follicle development. Journal of Investigative Dermatology 118(2): Pan, L., Liu, Y., Wei, Q., Xiao, C., Ji, Q., Bao, G., and Wu, X Solexa-Sequencing Based Transcriptome Study of Plaice Skin Phenotype in Rex Rabbits (Oryctolagus cuniculus). Plos One 10. Qin, S., Zhang, Z., Li, J., and Zang, L FRZB knockdown upregulates β-catenin activity and enhances cell aggressiveness in gastric cancer. Oncology reports 31(5): Rizzo, J.M., Romano, R.A., Bard, J., and Sinha, S RNA-seq Studies Reveal New Insights into p63 and the Transcriptomic Landscape of the Mouse Skin. Journal of Investigative Dermatology 135(2): Schmittgen, T.D., and Livak, K.J Analyzing real-time PCR data by the comparative CT method. Nature protocols 3(6): Skah, S., Nadjar, J., Sirakov, M., and Plateroti, M The secreted Frizzled-Related Protein 2 modulates cell fate and the Wnt pathway in the murine intestinal epithelium. Experimental cell research 330(1): Smeyne, R.J., Schilling, K., Robertson, L., Luk, D., Oberdick, J., Curran, T., and 17
19 Page 18 of Morgan, J.I Fos-IacZ transgenic mice: Mapping sites of gene induction in the central nervous system. Neuron 8(1): Stoick-Cooper, C.L., Moon, R.T., and Weidinger, G Advances in signaling in vertebrate regeneration as a prelude to regenerative medicine. Genes & development 21(11): Tao, Y Studies on the quality of rex rabbit fur. World Rabbit Sci 2(1): Trapnell, C., Pachter, L., and Salzberg, S.L TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25(9): Trapnell, C., Williams, B.A., Pertea, G., Mortazavi, A., Kwan, G., van Baren, M.J., Salzberg, S.L., Wold, B.J., and Pachter, L Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nature Biotechnology 28(5): Whelan, J.T., Hollis, S.E., Cha, D.S., Asch, A.S., and Lee, M.H Post transcriptional regulation of the Ras ERK/MAPK signaling pathway. Journal of cellular physiology 227(3): Xu, T., Guo, X., Wang, H., Hao, F., Du, X., Gao, X., and Liu, D Differential gene expression analysis between anagen and telogen of Capra hircus skin based on the de novo assembled transcriptome sequence. Gene 520(1):
20 Page 19 of Figure Legends Figure 1. Mapped region distribution of each sample. Mapped read distribution for each sample, showing the percentage of reads that mapped to each type of genomic region (genes, coding regions, splice sequences, intron sequences, intergenic regions, and non-coding regions (5 UTR and 3 UTR)) in the Ensembl OryCun2.0 database. Figure 2. Volcano plot of differentially expressed genes. Red dots show upregulated genes, green dots show downregulated genes, and two blue lines show a 2-fold change in expression (P=0.05). Figure 3. GO enrichment of genes related to fur development. Within the 404 molecular function category, we identified genes induced by fur development in rex rabbits. GO categories included biological processes, cellular component, and molecular function. Figure 4. Diagram of the interaction network. Thicker lines show stronger interactions of differentially expressed genes and their partner genes. Figure 5. Analysis of differentially expressed genes involved in the regulation of fur development in rabbits. (A) The mrna levels of FRZB, SFRP2, DUSP26, PTP4A3, EN1, CACNA1S, HBB1 and MRPL36 between back and belly groups. The expression level of genes in the back group was normalized to the belly group. (B) The protein levels of SFRP2 between the back and belly group. Each group had three biological replicates. Error bars represent the mean ± S.D. of triplicate experiments. *, P<0.05; **, P<
21 Page 20 of Table 1. Primer sequences used in qrt-pcr for validation of differentially expressed genes. Gene GAPDH Primers Forward primer: 5 -TCACCATCTTCCAGGAGCGA-3 Reverse primer: 5 - CACAATGCCGAAGTGGTCGT-3 FRZB Forward primer: 5 -CATCAAGTACCGCCACTCGT-3 Reverse primer: 5 -GCCCCTCTACAGTTTCCATTGCT-3 SFRP2 Forward primer: 5 -CCAGCCCGACTTCTCCTACAAGC-3 Reverse primer: 5 -TCCAGCACCTCTTTCATGGTCT-3 EN1 Forward primer: 5 -CTCCTGGGGCTTATCCGTCC-3 Reverse primer: 5 -CTCCCAGTTCCAGCCAAGGTC-3 CACNA1S Forward primer: 5 -TCATCCTCAGCGAGATCGACAC-3 Reverse primer: 5 -GATCAGCCTCATGACCCGGAAC-3 DUSP26 Forward primer: 5 -TAACTGGCTCTGGGCATCCAT-3 PTP4A3 Forward primer: 5 - Reverse primer: 5 -CCGCTCCAGCTCGAAGACGTT-3 AGAACATGCGCTTCCTCATCACC-3 20
22 Page 21 of 138 Reverse primer: 5 -TGTCGTAGGTCACTTCGCACAC-3 HBB1 Forward primer: 5 -GCTGCTGGTTGTCTACCCAT-3 Reverse primer: 5 -AGCCAGCACCTTCTTGCCAT-3 MRPL36 Forward primer: 5 -CCCGCGCTGGGCTTCAAGAC-3 Reverse primer: 5 - GGGTTGCTCTCGCAGTACACGAAC
23 Page 22 of Table 2. Summary of RNA-seq data for each sample. Sample ID Raw reads Quality trimmed Adaptor trimmed Clean reads Clean ratio rrna ratio 422 B1 62,459,166 59,686,034 58,308,948 54,544, % 0.30% 423 B2 42,040,878 41,785,111 41,048,067 40,089, % 0.70% 424 B3 57,515,444 56,961,934 56,014,421 54,639, % 0.20% F1 45,589,348 45,273,339 44,450,260 43,352, % 2.60% 425 F2 56,952,408 56,446,129 55,441,949 53,996, % 0.90% 426 F3 49,299,600 48,995,596 48,156,222 47,052, % 0.10% Clean ratio= (Clean reads/raw reads) %
24 Page 23 of Table 3. Mapping statistics for each sample. Sample All reads Mapped Mapped Mapped broken Mapped unique Mapped multiple Mapping ratio ID reads paired paired reads reads reads reads B1 54,391,204 44,245,125 42,943,776 1,301,349 39,786,635 4,458, % B2 39,826,940 33,675,819 32,120,474 1,555,345 30,460,340 3,215, % B3 54,509,484 45,897,437 44,222,232 1,675,205 41,895,083 4,002, % F1 42,242,866 35,960,580 34,407,676 1,552,904 32,706,780 3,253, % F2 53,524,836 45,300,446 43,337,930 1,962,516 40,933,492 4,366, % F3 47,012,530 39,549,833 37,741,242 1,808,591 35,663,951 3,885, % 431 Mapping ratio=mapped reads/all read 23
25 Page 24 of Supporting Information Figure S1. Bar plot of GO enrichment results. GO functional enrichment analysis of the differentially expressed genes, which were all considered statistically significant. Figure S2. Bar plot of KEGG enrichment results. KEGG pathway analysis of the differentially expressed genes, which were all considered statistically significant. Figure S3. Expressed genes annotated in the Wnt signaling pathway. The expressed genes are in the red box. Table S1. Gene expression in the two groups. 441 Table S2. Differentially expressed genes in the two groups Table S3. Significant GO terms of the differentially expressed genes. Table S4. Significant KEGG pathways of the differentially expressed genes. 24
26 Page 25 of 138 Figure 1. Mapped region distribution of each sample. Mapped read distribution for each sample, showing the percentage of reads that mapped to each type of genomic region (genes, coding regions, splice sequences, intron sequences, intergenic regions, and non-coding regions (5 UTR and 3 UTR)) in the Ensembl OryCun2.0 database. 78x53mm (300 x 300 DPI)
27 Page 26 of 138 Figure 2. Volcano plot of differentially expressed genes. Red dots show upregulated genes, green dots show downregulated genes, and two blue lines show a 2-fold change in expression (P=0.05). 190x189mm (300 x 300 DPI)
28 Page 27 of 138 Figure 3. GO enrichment of genes related to fur development. Within the molecular function category, we identified genes induced by fur development in rex rabbits. GO categories included biological processes, cellular component, and molecular function. 219x112mm (300 x 300 DPI)
29 Page 28 of 138 Figure 4. Diagram of the interaction network. Thicker lines show stronger interactions of differentially expressed genes and their partner genes. 169x145mm (300 x 300 DPI)
30 Page 29 of 138 Figure 5. Analysis of differentially expressed genes involved in the regulation of fur development in rabbits. (A) The mrna levels of FRZB, SFRP2, DUSP26, PTP4A3, EN1, CACNA1S, HBB1 and MRPL36 between back and belly groups. The expression level of genes in the back group was normalized to the belly group. (B) The protein levels of SFRP2 between the back and belly group. Each group had three biological replicates. Error bars represent the mean ± S.D. of triplicate experiments. *, P<0.05; **, P< x59mm (300 x 300 DPI)
31 Page 30 of x127mm (300 x 300 DPI)
32 Page 31 of x127mm (300 x 300 DPI)
33 Page 32 of x66mm (300 x 300 DPI)
34 Page 33 of 138 Supplemental Table S1. Gene expression in the two groups Gene_id gene_name Descriptionlocus Group B_FPKM Group F_FPKM ENSOCUG ZNF536 zinc finger GL018722: ENSOCUG NELL2 NEL-like 2 8: ENSOCUG IQCF1 IQ motif co9: ENSOCUG Uncharacte X: ENSOCUG AURKC aurora kinasgl018763: ENSOCUG Uncharacte 4: ENSOCUG FTCD formimidoy14: ENSOCUG CHIC1 cysteine-ricx: ENSOCUG HMGA2 high mobili 4: ENSOCUG SYT15 Uncharacte 18: ENSOCUG CDH24 cadherin 2417: ENSOCUG SMARCC1 GL018812: ENSOCUG FUT4 fucosyltrans1: ENSOCUG FAM222A family with GL019210: ENSOCUG C17orf64 chromosom19: ENSOCUG Uncharacte 17: ENSOCUG Uncharacte AAGW ENSOCUG WDR96 cilia and fla18: ENSOCUG NPTXR neuronal pe4: ENSOCUG RSPH4A radial spoke12: ENSOCUG RARB retinoic acid14: ENSOCUG C7orf61 chromosom6: ENSOCUG NAT8L N-acetyltranGL018796: ENSOCUG WDR76 Oryctolagus17: ENSOCUG CYP2C14 cytochrome18: ENSOCUG EFR3B EFR3 homo2: ENSOCUG BCAS1 breast carci GL018712: ENSOCUG HSD11B2 Oryctolagus5: ENSOCUG DUOXA2 dual oxidas 17: ENSOCUG Uncharacte GL019254: ENSOCUG CEP57L1 centrosoma 12: ENSOCUG GPR141 G protein-c 10: ENSOCUG IL8 Oryctolagus15: ENSOCUG Uncharacte 12: ENSOCUG Uncharacte GL018733: ENSOCUG COL6A5 collagen, ty14: ENSOCUG Uncharacte 12: ENSOCUG NCKAP5 NCK-assoc 7: ENSOCUG ATP6V0D2 ATPase, H+3: ENSOCUG GPR97 G protein-c 5: ENSOCUG CASP1 caspase 1, a1: ENSOCUG TSPYL5 TSPY-like 53: ENSOCUG GRM2 glutamate re9: ENSOCUG NLGN3 neuroligin 3X: ENSOCUG : ENSOCUG RHOH ras homolog2: ENSOCUG CDC25A cell division9: ENSOCUG RSPH9 radial spoke12:
35 Page 34 of 138 ENSOCUG Uncharacte GL019297: ENSOCUG TTPA tocopherol (3: ENSOCUG PFAS phosphoribo19: ENSOCUG SMPDL3B sphingomye13: ENSOCUG IL18R1 Oryctolagus2: ENSOCUG Uncharacte GL018981: ENSOCUG PLXNC1 plexin C1 [S4: ENSOCUG KIAA1958 KIAA1958 1: ENSOCUG CHST11 carbohydrat4: ENSOCUG SKA1 spindle and 9: ENSOCUG INTU inturned pla15: ENSOCUG SLC47A2 solute carriegl018817: ENSOCUG NKG7 natural killegl019218: ENSOCUG GCH1 GTP cycloh17: ENSOCUG CD4 Oryctolagus8: ENSOCUG P2RY6 pyrimidiner1: ENSOCUG TAF4B TAF4b RNA9: ENSOCUG SH2D2A SH2 domain13: ENSOCUG CD226 CD226 mol9: ENSOCUG Uncharacte GL018823: ENSOCUG Uncharacte X: ENSOCUG SNX22 sorting nexi17: ENSOCUG Uncharacte 2: ENSOCUG PLD6 phospholipagl018920: ENSOCUG CLSPN claspin [SouGL018704: ENSOCUG DHRS9 dehydrogen7: ENSOCUG Uncharacte 18: ENSOCUG RAD54B RAD54 hom3: ENSOCUG HIST2H2AAhistone clus13: ENSOCUG GIMAP8 GTPase, IMGL018806: ENSOCUG KL klotho [SouGL018702: ENSOCUG TIFAB TRAF-inter3: ENSOCUG Uncharacte 5: ENSOCUG : ENSOCUG : ENSOCUG FAM64A family with 19: ENSOCUG TPMT OryctolagusGL018711: ENSOCUG PARVG parvin, gamgl019802: ENSOCUG Uncharacte 9: ENSOCUG Uncharacte GL018806: ENSOCUG RAD51AP1 RAD51 ass 8: ENSOCUG IKZF1 IKAROS fagl018745: ENSOCUG FOXM1 forkhead bo8: ENSOCUG SK 7SK RNA [3: ENSOCUG TFF1 trefoil factoaagw ENSOCUG RIC3 RIC3 acetyl1: ENSOCUG ZDHHC23 zinc finger, 14: ENSOCUG ECE2 endothelin c14: ENSOCUG CIDEB cell death-in17: ENSOCUG FANCA Fanconi anegl018965:
36 Page 35 of 138 ENSOCUG Uncharacte GL018888: ENSOCUG CD5 OryctolagusGL018717: ENSOCUG CLDN19 claudin 19 [13: ENSOCUG SKA3 spindle and 8: ENSOCUG MFSD4 major facili 16: ENSOCUG KANK4 KN motif a 13: ENSOCUG PPP1R14D protein pho 17: ENSOCUG GUCY1A2 guanylate c 1: ENSOCUG Uncharacte 15: ENSOCUG CD247 Oryctolagus13: ENSOCUG UBE2T ubiquitin-co16: ENSOCUG TRPV5 Oryctolagus7: ENSOCUG DLX1 distal-less h7: ENSOCUG Uncharacte X: ENSOCUG STIL SCL/TAL1 13: ENSOCUG PLA1A phospholipa14: ENSOCUG DTX1 deltex 1, E321: ENSOCUG SKIDA1 SKI/DACH16: ENSOCUG HEMGN hemogen [S1: ENSOCUG GL018986: ENSOCUG SH3GL3 SH3-domai GL018737: ENSOCUG BCHE butyrylchol 14: ENSOCUG PRIMA1 proline rich 20: ENSOCUG Uncharacte GL018781: ENSOCUG CDH6 cadherin 6, 11: ENSOCUG ADAM33 ADAM met4: ENSOCUG Uncharacte 1: ENSOCUG TBX21 T-box 21 [S19: ENSOCUG Uncharacte 5: ENSOCUG HAVCR2 hepatitis A 3: ENSOCUG NGEF neuronal gugl018736: ENSOCUG SLC4A1 solute carrie19: ENSOCUG Uncharacte GL018758: ENSOCUG Uncharacte GL018760: ENSOCUG SARDH sarcosine degl019710: ENSOCUG SLC44A4 solute carrie12: ENSOCUG U3 Small nucle19: ENSOCUG ARG1 Arginase-1 12: ENSOCUG SUCNR1 succinate re14: ENSOCUG Uncharacte AAGW ENSOCUG ESM1 endothelial 11: ENSOCUG SSC5D GL019887: ENSOCUG Uncharacte GL018707: ENSOCUG X: ENSOCUG ALX3 ALX homeo13: ENSOCUG PROX1 prospero ho16: ENSOCUG GPR17 G protein-c 7: ENSOCUG GINS4 GINS compgl018706: ENSOCUG DHH desert hedg 4: ENSOCUG RASAL3 RAS proteingl018829:
37 Page 36 of 138 ENSOCUG FBXO41 F-box prote2: ENSOCUG KLRD1 killer cell le8: ENSOCUG NT5M 5',3'-nucleo GL018920: ENSOCUG SK 7SK RNA [15: ENSOCUG CREG2 cellular repr2: ENSOCUG GL018747: ENSOCUG TSHZ2 teashirt zincgl018712: ENSOCUG ARHGEF39 Rho guanin 1: ENSOCUG GNAT3 guanine nuc7: ENSOCUG protein disuaagw ENSOCUG SMCO3 single-pass 8: ENSOCUG CTSE Oryctolagus16: ENSOCUG Uncharacte 5: ENSOCUG TCFL5 transcriptio GL019426: ENSOCUG Uncharacte 13: ENSOCUG WSCD2 WSC doma GL018777: ENSOCUG Uncharacte GL019405: ENSOCUG SNORA23 Small nucle1: ENSOCUG Histone H2 12: ENSOCUG AADACL3 arylacetamigl018739: ENSOCUG Uncharacte 17: ENSOCUG Uncharacte GL018792: ENSOCUG FASLG Fas ligand (13: ENSOCUG SGOL1 shugoshin-l14: ENSOCUG HS3ST1 heparan sul 2: ENSOCUG SK 7SK RNA [GL019070: ENSOCUG Uncharacte 14: ENSOCUG GL018886: ENSOCUG HCAR1 hydroxycar GL018824: ENSOCUG RASL10B RAS-like, f 19: ENSOCUG BIK BCL2-inter GL019017: ENSOCUG HTR1B Oryctolagus12: ENSOCUG GCG Glucagon [7: ENSOCUG ZNF775 zinc finger GL018806: ENSOCUG NTRK3 neurotrophigl018934: ENSOCUG RAB39A RAB39A, m1: ENSOCUG CD8B CD8b mole 2: ENSOCUG TEX101 testis expresgl019267: ENSOCUG FGF2 fibroblast g 15: ENSOCUG SH2D1A SH2 domainx: ENSOCUG GOLGA7B golgin A7 f 18: ENSOCUG Histone H2 12: ENSOCUG FGF16 fibroblast g X: ENSOCUG T cell recepgl018758: ENSOCUG CA1 carbonic an 3: ENSOCUG NFE2 nuclear fact4: ENSOCUG AWAT2 acyl-coa wx: ENSOCUG Uncharacte GL019021: ENSOCUG C14orf37 chromosom17: ENSOCUG RPL35A Uncharacte 3:
Cellular Physiology and Biochemistry
 Original Paper 2016 The Author(s). 2016 Published The Author(s) by S. Karger AG, Basel Published online: November 25, 2016 www.karger.com/cpb Published by S. Karger AG, Basel 486 www.karger.com/cpb Accepted:
Original Paper 2016 The Author(s). 2016 Published The Author(s) by S. Karger AG, Basel Published online: November 25, 2016 www.karger.com/cpb Published by S. Karger AG, Basel 486 www.karger.com/cpb Accepted:
Μελέτη της έκφρασης του ογκοκατασταλτικού γονιδίου Cyld στον καρκίνο του μαστού
 Σχολή Θετικών Επιστημών Τμήμα Βιολογίας Πρόγραμμα Μεταπτυχιακών Σπουδών Κατεύθυνση: Εφαρμοσμένη γενετική και βιοτεχνολογία ΜΕΤΑΠΤΥΧΙΑΚΗ ΔΙΠΛΩΜΑΤΙΚΗ ΕΡΓΑΣΙΑ Μελέτη της έκφρασης του ογκοκατασταλτικού γονιδίου
Σχολή Θετικών Επιστημών Τμήμα Βιολογίας Πρόγραμμα Μεταπτυχιακών Σπουδών Κατεύθυνση: Εφαρμοσμένη γενετική και βιοτεχνολογία ΜΕΤΑΠΤΥΧΙΑΚΗ ΔΙΠΛΩΜΑΤΙΚΗ ΕΡΓΑΣΙΑ Μελέτη της έκφρασης του ογκοκατασταλτικού γονιδίου
Optimizing Microwave-assisted Extraction Process for Paprika Red Pigments Using Response Surface Methodology
 2012 34 2 382-387 http / /xuebao. jxau. edu. cn Acta Agriculturae Universitatis Jiangxiensis E - mail ndxb7775@ sina. com 212018 105 W 42 2 min 0. 631 TS202. 3 A 1000-2286 2012 02-0382 - 06 Optimizing
2012 34 2 382-387 http / /xuebao. jxau. edu. cn Acta Agriculturae Universitatis Jiangxiensis E - mail ndxb7775@ sina. com 212018 105 W 42 2 min 0. 631 TS202. 3 A 1000-2286 2012 02-0382 - 06 Optimizing
ΓΕΩΠΟΝΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΑΘΗΝΩΝ ΤΜΗΜΑ ΕΠΙΣΤΗΜΗΣ ΤΡΟΦΙΜΩΝ ΚΑΙ ΔΙΑΤΡΟΦΗΣ ΤΟΥ ΑΝΘΡΩΠΟΥ
 ΓΕΩΠΟΝΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΑΘΗΝΩΝ ΤΜΗΜΑ ΕΠΙΣΤΗΜΗΣ ΤΡΟΦΙΜΩΝ ΚΑΙ ΔΙΑΤΡΟΦΗΣ ΤΟΥ ΑΝΘΡΩΠΟΥ Πρόγραμμα Μεταπτυχιακών Σπουδών «Επιστήμη και Τεχνολογία Τροφίμων και Διατροφή του Ανθρώπου» Κατεύθυνση: «Διατροφή, Δημόσια
ΓΕΩΠΟΝΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΑΘΗΝΩΝ ΤΜΗΜΑ ΕΠΙΣΤΗΜΗΣ ΤΡΟΦΙΜΩΝ ΚΑΙ ΔΙΑΤΡΟΦΗΣ ΤΟΥ ΑΝΘΡΩΠΟΥ Πρόγραμμα Μεταπτυχιακών Σπουδών «Επιστήμη και Τεχνολογία Τροφίμων και Διατροφή του Ανθρώπου» Κατεύθυνση: «Διατροφή, Δημόσια
Identification of Fish Species using DNA Method
 5 * * * Identification of Fish Species using DNA Method Yuumi KAWASHIMA*, Takayuki KATAYAMA* and Yukihiko YAMAZAKI* *Central Customs Laboratory, Ministry of Finance 6-3-5, Kashiwanoha, Kashiwa, Chiba 277-0882,
5 * * * Identification of Fish Species using DNA Method Yuumi KAWASHIMA*, Takayuki KATAYAMA* and Yukihiko YAMAZAKI* *Central Customs Laboratory, Ministry of Finance 6-3-5, Kashiwanoha, Kashiwa, Chiba 277-0882,
HIV HIV HIV HIV AIDS 3 :.1 /-,**1 +332
 ,**1 The Japanese Society for AIDS Research The Journal of AIDS Research +,, +,, +,, + -. / 0 1 +, -. / 0 1 : :,**- +,**. 1..+ - : +** 22 HIV AIDS HIV HIV AIDS : HIV AIDS HIV :HIV AIDS 3 :.1 /-,**1 HIV
,**1 The Japanese Society for AIDS Research The Journal of AIDS Research +,, +,, +,, + -. / 0 1 +, -. / 0 1 : :,**- +,**. 1..+ - : +** 22 HIV AIDS HIV HIV AIDS : HIV AIDS HIV :HIV AIDS 3 :.1 /-,**1 HIV
ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΥΠΡΟΥ ΤΜΗΜΑ ΝΟΣΗΛΕΥΤΙΚΗΣ
 ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΥΠΡΟΥ ΤΜΗΜΑ ΝΟΣΗΛΕΥΤΙΚΗΣ ΠΤΥΧΙΑΚΗ ΕΡΓΑΣΙΑ ΨΥΧΟΛΟΓΙΚΕΣ ΕΠΙΠΤΩΣΕΙΣ ΣΕ ΓΥΝΑΙΚΕΣ ΜΕΤΑ ΑΠΟ ΜΑΣΤΕΚΤΟΜΗ ΓΕΩΡΓΙΑ ΤΡΙΣΟΚΚΑ Λευκωσία 2012 ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΥΠΡΟΥ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ
ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΥΠΡΟΥ ΤΜΗΜΑ ΝΟΣΗΛΕΥΤΙΚΗΣ ΠΤΥΧΙΑΚΗ ΕΡΓΑΣΙΑ ΨΥΧΟΛΟΓΙΚΕΣ ΕΠΙΠΤΩΣΕΙΣ ΣΕ ΓΥΝΑΙΚΕΣ ΜΕΤΑ ΑΠΟ ΜΑΣΤΕΚΤΟΜΗ ΓΕΩΡΓΙΑ ΤΡΙΣΟΚΚΑ Λευκωσία 2012 ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΥΠΡΟΥ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ
«ΑΓΡΟΤΟΥΡΙΣΜΟΣ ΚΑΙ ΤΟΠΙΚΗ ΑΝΑΠΤΥΞΗ: Ο ΡΟΛΟΣ ΤΩΝ ΝΕΩΝ ΤΕΧΝΟΛΟΓΙΩΝ ΣΤΗΝ ΠΡΟΩΘΗΣΗ ΤΩΝ ΓΥΝΑΙΚΕΙΩΝ ΣΥΝΕΤΑΙΡΙΣΜΩΝ»
 I ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΣΧΟΛΗ ΝΟΜΙΚΩΝ ΟΙΚΟΝΟΜΙΚΩΝ ΚΑΙ ΠΟΛΙΤΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΤΜΗΜΑ ΟΙΚΟΝΟΜΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ ΣΤΗΝ «ΔΙΟΙΚΗΣΗ ΚΑΙ ΟΙΚΟΝΟΜΙΑ» ΚΑΤΕΥΘΥΝΣΗ: ΟΙΚΟΝΟΜΙΚΗ
I ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΣΧΟΛΗ ΝΟΜΙΚΩΝ ΟΙΚΟΝΟΜΙΚΩΝ ΚΑΙ ΠΟΛΙΤΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΤΜΗΜΑ ΟΙΚΟΝΟΜΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ ΣΤΗΝ «ΔΙΟΙΚΗΣΗ ΚΑΙ ΟΙΚΟΝΟΜΙΑ» ΚΑΤΕΥΘΥΝΣΗ: ΟΙΚΟΝΟΜΙΚΗ
Longitudinal Changes in Component Processes of Working Memory
 This Accepted Manuscript has not been copyedited and formatted. The final version may differ from this version. Research Article: New Research Cognition and Behavior Longitudinal Changes in Component Processes
This Accepted Manuscript has not been copyedited and formatted. The final version may differ from this version. Research Article: New Research Cognition and Behavior Longitudinal Changes in Component Processes
Introduction to Bioinformatics
 Introduction to Bioinformatics 260.602.01 September 2, 2005 Jonathan Pevsner, Ph.D. pevsner@jhmi.edu bioinformatics medical informatics Tool-users public health informatics databases algorithms Tool-makers
Introduction to Bioinformatics 260.602.01 September 2, 2005 Jonathan Pevsner, Ph.D. pevsner@jhmi.edu bioinformatics medical informatics Tool-users public health informatics databases algorithms Tool-makers
IL - 13 /IL - 18 ELISA PCR RT - PCR. IL - 13 IL - 18 mrna. 13 IL - 18 mrna IL - 13 /IL Th1 /Th2
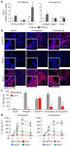 344 IL - 13 /IL - 18 1 2 1 2 1 2 1 2 1 2 3 1 2 13 18 IL - 13 /IL - 18 10% / OVA /AL OH 3 5% 16 ~ 43 d 44 d ELISA BALF IL - 13 IL - 18 PCR RT - PCR IL - 13 IL - 18 mrna IL - 13 mrna 0. 01 IL - 18 mrna 0.
344 IL - 13 /IL - 18 1 2 1 2 1 2 1 2 1 2 3 1 2 13 18 IL - 13 /IL - 18 10% / OVA /AL OH 3 5% 16 ~ 43 d 44 d ELISA BALF IL - 13 IL - 18 PCR RT - PCR IL - 13 IL - 18 mrna IL - 13 mrna 0. 01 IL - 18 mrna 0.
Salmonella produce microrna-like RNA fragment Sal-1 in the infected cells to. facilitate intracellular survival
 Salmonella produce microrna-like RNA fragment Sal-1 in the infected cells to facilitate intracellular survival Hongwei Gu 1,2*, Chihao Zhao 1,2*, Tianfu Zhang 1,2*, Hongwei Liang 1,2, Xiao-Ming Wang 1,
Salmonella produce microrna-like RNA fragment Sal-1 in the infected cells to facilitate intracellular survival Hongwei Gu 1,2*, Chihao Zhao 1,2*, Tianfu Zhang 1,2*, Hongwei Liang 1,2, Xiao-Ming Wang 1,
TABLE OF CONTENTS Page
 TABLE OF CONTENTS Page Acknowledgements... VI Declaration... VII List of abbreviations... VIII 1. INTRODUCTION... 1 2. BASIC PRINCIPLES... 3 2.1. Hereditary Colorectal Cancer... 3 2.1.1. Differential diagnosis...
TABLE OF CONTENTS Page Acknowledgements... VI Declaration... VII List of abbreviations... VIII 1. INTRODUCTION... 1 2. BASIC PRINCIPLES... 3 2.1. Hereditary Colorectal Cancer... 3 2.1.1. Differential diagnosis...
ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΥΠΡΟΥ ΣΧΟΛΗ ΓΕΩΠΟΝΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΒΙΟΤΕΧΝΟΛΟΓΙΑΣ ΚΑΙ ΕΠΙΣΤΗΜΗΣ ΤΡΟΦΙΜΩΝ. Πτυχιακή εργασία
 ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΥΠΡΟΥ ΣΧΟΛΗ ΓΕΩΠΟΝΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΒΙΟΤΕΧΝΟΛΟΓΙΑΣ ΚΑΙ ΕΠΙΣΤΗΜΗΣ ΤΡΟΦΙΜΩΝ Πτυχιακή εργασία ΜΕΛΕΤΗ ΠΟΛΥΦΑΙΝΟΛΩΝ ΚΑΙ ΑΝΤΙΟΞΕΙΔΩΤΙΚΗΣ ΙΚΑΝΟΤΗΤΑΣ ΣΟΚΟΛΑΤΑΣ Αναστασία Σιάντωνα Λεμεσός
ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΥΠΡΟΥ ΣΧΟΛΗ ΓΕΩΠΟΝΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΒΙΟΤΕΧΝΟΛΟΓΙΑΣ ΚΑΙ ΕΠΙΣΤΗΜΗΣ ΤΡΟΦΙΜΩΝ Πτυχιακή εργασία ΜΕΛΕΤΗ ΠΟΛΥΦΑΙΝΟΛΩΝ ΚΑΙ ΑΝΤΙΟΞΕΙΔΩΤΙΚΗΣ ΙΚΑΝΟΤΗΤΑΣ ΣΟΚΟΛΑΤΑΣ Αναστασία Σιάντωνα Λεμεσός
A strategy for the identification of combinatorial bioactive compounds. contributing to the holistic effect of herbal medicines
 1 2 Supplementary information 3 4 A strategy for the identification of combinatorial bioactive compounds contributing to the holistic effect of herbal medicines 5 6 Fang Long 1, Hua Yang 1, Yanmin Xu,
1 2 Supplementary information 3 4 A strategy for the identification of combinatorial bioactive compounds contributing to the holistic effect of herbal medicines 5 6 Fang Long 1, Hua Yang 1, Yanmin Xu,
ΕΘΝΙΚΟ ΜΕΤΣΟΒΙΟ ΠΟΛΥΤΕΧΝΕΙΟ
 ΕΘΝΙΚΟ ΜΕΤΣΟΒΙΟ ΠΟΛΥΤΕΧΝΕΙΟ ΣΧΟΛΗ ΗΛΕΚΤΡΟΛΟΓΩΝ ΜΗΧΑΝΙΚΩΝ ΚΑΙ ΜΗΧΑΝΙΚΩΝ ΥΠΟΛΟΓΙΣΤΩΝ ΤΟΜΕΑΣ ΤΕΧΝΟΛΟΓΙΑΣ ΠΛΗΡΟΦΟΡΙΚΗΣ ΚΑΙ ΥΠΟΛΟΓΙΣΤΩΝ Σύστημα Διαχείρισης Διαχρονικών Δεδομένων για Γονίδια ΔΙΠΛΩΜΑΤΙΚΗ ΕΡΓΑΣΙΑ
ΕΘΝΙΚΟ ΜΕΤΣΟΒΙΟ ΠΟΛΥΤΕΧΝΕΙΟ ΣΧΟΛΗ ΗΛΕΚΤΡΟΛΟΓΩΝ ΜΗΧΑΝΙΚΩΝ ΚΑΙ ΜΗΧΑΝΙΚΩΝ ΥΠΟΛΟΓΙΣΤΩΝ ΤΟΜΕΑΣ ΤΕΧΝΟΛΟΓΙΑΣ ΠΛΗΡΟΦΟΡΙΚΗΣ ΚΑΙ ΥΠΟΛΟΓΙΣΤΩΝ Σύστημα Διαχείρισης Διαχρονικών Δεδομένων για Γονίδια ΔΙΠΛΩΜΑΤΙΚΗ ΕΡΓΑΣΙΑ
Christopher Stephen Inchley, 2015
 Christopher Stephen Inchley, 2015 Series of dissertations submitted to the Faculty of Medicine, University of Oslo No. 2009 ISBN 978-82-8264-990-2 All rights reserved. No part of this publication may be
Christopher Stephen Inchley, 2015 Series of dissertations submitted to the Faculty of Medicine, University of Oslo No. 2009 ISBN 978-82-8264-990-2 All rights reserved. No part of this publication may be
EE512: Error Control Coding
 EE512: Error Control Coding Solution for Assignment on Finite Fields February 16, 2007 1. (a) Addition and Multiplication tables for GF (5) and GF (7) are shown in Tables 1 and 2. + 0 1 2 3 4 0 0 1 2 3
EE512: Error Control Coding Solution for Assignment on Finite Fields February 16, 2007 1. (a) Addition and Multiplication tables for GF (5) and GF (7) are shown in Tables 1 and 2. + 0 1 2 3 4 0 0 1 2 3
Mean bond enthalpy Standard enthalpy of formation Bond N H N N N N H O O O
 Q1. (a) Explain the meaning of the terms mean bond enthalpy and standard enthalpy of formation. Mean bond enthalpy... Standard enthalpy of formation... (5) (b) Some mean bond enthalpies are given below.
Q1. (a) Explain the meaning of the terms mean bond enthalpy and standard enthalpy of formation. Mean bond enthalpy... Standard enthalpy of formation... (5) (b) Some mean bond enthalpies are given below.
Abstract... I. Zusammenfassung... II. 1 Aim of the work Introduction Short overview of Chinese hamster ovary cell lines...
 III Contents I II III Abstract... I Zusammenfassung... II Contents... III 1 Aim of the work... 1 2 Introduction... 2 2.1 Short overview of Chinese hamster ovary cell lines... 2 2.2 Unspecific mutations:
III Contents I II III Abstract... I Zusammenfassung... II Contents... III 1 Aim of the work... 1 2 Introduction... 2 2.1 Short overview of Chinese hamster ovary cell lines... 2 2.2 Unspecific mutations:
Αξιολόγηση των Φασματικού Διαχωρισμού στην Διάκριση Διαφορετικών Τύπων Εδάφους ΔΙΠΛΩΜΑΤΙΚΗ ΕΡΓΑΣΙΑ. Σπίγγος Γεώργιος
 ΕΘΝΙΚΟ ΜΕΤΣΟΒΙΟ ΠΟΛΥΤΕΧΝΕΙΟ ΤΜΗΜΑ ΑΓΡΟΝΟΜΩΝ ΤΟΠΟΓΡΑΦΩΝ ΜΗΧΑΝΙΚΩΝ ΤΟΜΕΑΣ ΤΟΠΟΓΡΑΦΙΑΣ-ΕΡΓΑΣΤΗΡΙΟ ΤΗΛΕΠΙΣΚΟΠΗΣΗΣ Αξιολόγηση των Φασματικού Διαχωρισμού στην Διάκριση Διαφορετικών Τύπων Εδάφους ΔΙΠΛΩΜΑΤΙΚΗ
ΕΘΝΙΚΟ ΜΕΤΣΟΒΙΟ ΠΟΛΥΤΕΧΝΕΙΟ ΤΜΗΜΑ ΑΓΡΟΝΟΜΩΝ ΤΟΠΟΓΡΑΦΩΝ ΜΗΧΑΝΙΚΩΝ ΤΟΜΕΑΣ ΤΟΠΟΓΡΑΦΙΑΣ-ΕΡΓΑΣΤΗΡΙΟ ΤΗΛΕΠΙΣΚΟΠΗΣΗΣ Αξιολόγηση των Φασματικού Διαχωρισμού στην Διάκριση Διαφορετικών Τύπων Εδάφους ΔΙΠΛΩΜΑΤΙΚΗ
2 Composition. Invertible Mappings
 Arkansas Tech University MATH 4033: Elementary Modern Algebra Dr. Marcel B. Finan Composition. Invertible Mappings In this section we discuss two procedures for creating new mappings from old ones, namely,
Arkansas Tech University MATH 4033: Elementary Modern Algebra Dr. Marcel B. Finan Composition. Invertible Mappings In this section we discuss two procedures for creating new mappings from old ones, namely,
ΕΦΑΡΜΟΓΗ ΕΥΤΕΡΟΒΑΘΜΙΑ ΕΠΕΞΕΡΓΑΣΜΕΝΩΝ ΥΓΡΩΝ ΑΠΟΒΛΗΤΩΝ ΣΕ ΦΥΣΙΚΑ ΣΥΣΤΗΜΑΤΑ ΚΛΙΝΗΣ ΚΑΛΑΜΙΩΝ
 ΤΕΧΝΟΛΟΓΙΚΟ ΕΚΠΑΙ ΕΥΤΙΚΟ Ι ΡΥΜΑ ΚΡΗΤΗΣ ΤΜΗΜΑ ΦΥΣΙΚΩΝ ΠΟΡΩΝ ΚΑΙ ΠΕΡΙΒΑΛΛΟΝΤΟΣ ΕΦΑΡΜΟΓΗ ΕΥΤΕΡΟΒΑΘΜΙΑ ΕΠΕΞΕΡΓΑΣΜΕΝΩΝ ΥΓΡΩΝ ΑΠΟΒΛΗΤΩΝ ΣΕ ΦΥΣΙΚΑ ΣΥΣΤΗΜΑΤΑ ΚΛΙΝΗΣ ΚΑΛΑΜΙΩΝ ΕΠΙΜΕΛΕΙΑ: ΑΡΜΕΝΑΚΑΣ ΜΑΡΙΝΟΣ ΧΑΝΙΑ
ΤΕΧΝΟΛΟΓΙΚΟ ΕΚΠΑΙ ΕΥΤΙΚΟ Ι ΡΥΜΑ ΚΡΗΤΗΣ ΤΜΗΜΑ ΦΥΣΙΚΩΝ ΠΟΡΩΝ ΚΑΙ ΠΕΡΙΒΑΛΛΟΝΤΟΣ ΕΦΑΡΜΟΓΗ ΕΥΤΕΡΟΒΑΘΜΙΑ ΕΠΕΞΕΡΓΑΣΜΕΝΩΝ ΥΓΡΩΝ ΑΠΟΒΛΗΤΩΝ ΣΕ ΦΥΣΙΚΑ ΣΥΣΤΗΜΑΤΑ ΚΛΙΝΗΣ ΚΑΛΑΜΙΩΝ ΕΠΙΜΕΛΕΙΑ: ΑΡΜΕΝΑΚΑΣ ΜΑΡΙΝΟΣ ΧΑΝΙΑ
MSM Men who have Sex with Men HIV -
 ,**, The Japanese Society for AIDS Research The Journal of AIDS Research HIV,0 + + + + +,,, +, : HIV : +322,*** HIV,0,, :., n,0,,. + 2 2, CD. +3-ml n,, AIDS 3 ARC 3 +* 1. A, MSM Men who have Sex with Men
,**, The Japanese Society for AIDS Research The Journal of AIDS Research HIV,0 + + + + +,,, +, : HIV : +322,*** HIV,0,, :., n,0,,. + 2 2, CD. +3-ml n,, AIDS 3 ARC 3 +* 1. A, MSM Men who have Sex with Men
ΕΠΙΔΡΑΣΗ ΤΗΣ ΧΡΗΣΗΣ ΕΛΑΙΟΠΛΑΚΟΥΝΤΑ ΣΤΗΝ ΔΙΑΤΡΟΦΗ ΤΩΝ ΑΙΓΩΝ ΔΑΜΑΣΚΟΥ ΩΣ ΠΡΟΣ ΤΗΝ ΠΟΣΟΤΗΤΑ ΚΑΙ ΠΟΙΟΤΗΤΑ ΤΟΥ ΠΑΡΑΓΟΜΕΝΟΥ ΓΑΛΑΚΤΟΣ
 Σχολή Γεωπονικών Επιστημών και Διαχείρισης Περιβάλλοντος Πτυχιακή εργασία ΕΠΙΔΡΑΣΗ ΤΗΣ ΧΡΗΣΗΣ ΕΛΑΙΟΠΛΑΚΟΥΝΤΑ ΣΤΗΝ ΔΙΑΤΡΟΦΗ ΤΩΝ ΑΙΓΩΝ ΔΑΜΑΣΚΟΥ ΩΣ ΠΡΟΣ ΤΗΝ ΠΟΣΟΤΗΤΑ ΚΑΙ ΠΟΙΟΤΗΤΑ ΤΟΥ ΠΑΡΑΓΟΜΕΝΟΥ ΓΑΛΑΚΤΟΣ
Σχολή Γεωπονικών Επιστημών και Διαχείρισης Περιβάλλοντος Πτυχιακή εργασία ΕΠΙΔΡΑΣΗ ΤΗΣ ΧΡΗΣΗΣ ΕΛΑΙΟΠΛΑΚΟΥΝΤΑ ΣΤΗΝ ΔΙΑΤΡΟΦΗ ΤΩΝ ΑΙΓΩΝ ΔΑΜΑΣΚΟΥ ΩΣ ΠΡΟΣ ΤΗΝ ΠΟΣΟΤΗΤΑ ΚΑΙ ΠΟΙΟΤΗΤΑ ΤΟΥ ΠΑΡΑΓΟΜΕΝΟΥ ΓΑΛΑΚΤΟΣ
ΠΤΥΧΙΑΚΗ ΕΡΓΑΣΙΑ "ΠΟΛΥΚΡΙΤΗΡΙΑ ΣΥΣΤΗΜΑΤΑ ΛΗΨΗΣ ΑΠΟΦΑΣΕΩΝ. Η ΠΕΡΙΠΤΩΣΗ ΤΗΣ ΕΠΙΛΟΓΗΣ ΑΣΦΑΛΙΣΤΗΡΙΟΥ ΣΥΜΒΟΛΑΙΟΥ ΥΓΕΙΑΣ "
 ΤΕΧΝΟΛΟΓΙΚΟ ΕΚΠΑΙΔΕΥΤΙΚΟ ΙΔΡΥΜΑ ΚΑΛΑΜΑΤΑΣ ΣΧΟΛΗ ΔΙΟΙΚΗΣΗΣ ΟΙΚΟΝΟΜΙΑΣ ΤΜΗΜΑ ΜΟΝΑΔΩΝ ΥΓΕΙΑΣ ΚΑΙ ΠΡΟΝΟΙΑΣ ΠΤΥΧΙΑΚΗ ΕΡΓΑΣΙΑ "ΠΟΛΥΚΡΙΤΗΡΙΑ ΣΥΣΤΗΜΑΤΑ ΛΗΨΗΣ ΑΠΟΦΑΣΕΩΝ. Η ΠΕΡΙΠΤΩΣΗ ΤΗΣ ΕΠΙΛΟΓΗΣ ΑΣΦΑΛΙΣΤΗΡΙΟΥ ΣΥΜΒΟΛΑΙΟΥ
ΤΕΧΝΟΛΟΓΙΚΟ ΕΚΠΑΙΔΕΥΤΙΚΟ ΙΔΡΥΜΑ ΚΑΛΑΜΑΤΑΣ ΣΧΟΛΗ ΔΙΟΙΚΗΣΗΣ ΟΙΚΟΝΟΜΙΑΣ ΤΜΗΜΑ ΜΟΝΑΔΩΝ ΥΓΕΙΑΣ ΚΑΙ ΠΡΟΝΟΙΑΣ ΠΤΥΧΙΑΚΗ ΕΡΓΑΣΙΑ "ΠΟΛΥΚΡΙΤΗΡΙΑ ΣΥΣΤΗΜΑΤΑ ΛΗΨΗΣ ΑΠΟΦΑΣΕΩΝ. Η ΠΕΡΙΠΤΩΣΗ ΤΗΣ ΕΠΙΛΟΓΗΣ ΑΣΦΑΛΙΣΤΗΡΙΟΥ ΣΥΜΒΟΛΑΙΟΥ
NATIONAL AND KAPODISTRIAN UNIVERSITY OF ATHENS SCHOOL OF SCIENCE FACULTY OF INFORMATICS AND TELECOMMUNICATIONS
 NATIONAL AND KAPODISTRIAN UNIVERSITY OF ATHENS SCHOOL OF SCIENCE FACULTY OF INFORMATICS AND TELECOMMUNICATIONS INTERDICIPLINARY POSTGRADUATE PROGRAMME "INFORMATION TECHNOLOGIES IN MEDICINE AND BIOLOGY"
NATIONAL AND KAPODISTRIAN UNIVERSITY OF ATHENS SCHOOL OF SCIENCE FACULTY OF INFORMATICS AND TELECOMMUNICATIONS INTERDICIPLINARY POSTGRADUATE PROGRAMME "INFORMATION TECHNOLOGIES IN MEDICINE AND BIOLOGY"
ΠΑΝΔΠΗΣΖΜΗΟ ΠΑΣΡΩΝ ΣΜΖΜΑ ΖΛΔΚΣΡΟΛΟΓΩΝ ΜΖΥΑΝΗΚΩΝ ΚΑΗ ΣΔΥΝΟΛΟΓΗΑ ΤΠΟΛΟΓΗΣΩΝ ΣΟΜΔΑ ΤΣΖΜΑΣΩΝ ΖΛΔΚΣΡΗΚΖ ΔΝΔΡΓΔΗΑ
 ΠΑΝΔΠΗΣΖΜΗΟ ΠΑΣΡΩΝ ΣΜΖΜΑ ΖΛΔΚΣΡΟΛΟΓΩΝ ΜΖΥΑΝΗΚΩΝ ΚΑΗ ΣΔΥΝΟΛΟΓΗΑ ΤΠΟΛΟΓΗΣΩΝ ΣΟΜΔΑ ΤΣΖΜΑΣΩΝ ΖΛΔΚΣΡΗΚΖ ΔΝΔΡΓΔΗΑ Γηπισκαηηθή Δξγαζία ηνπ Φνηηεηή ηνπ ηκήκαηνο Ζιεθηξνιόγσλ Μεραληθώλ θαη Σερλνινγίαο Ζιεθηξνληθώλ
ΠΑΝΔΠΗΣΖΜΗΟ ΠΑΣΡΩΝ ΣΜΖΜΑ ΖΛΔΚΣΡΟΛΟΓΩΝ ΜΖΥΑΝΗΚΩΝ ΚΑΗ ΣΔΥΝΟΛΟΓΗΑ ΤΠΟΛΟΓΗΣΩΝ ΣΟΜΔΑ ΤΣΖΜΑΣΩΝ ΖΛΔΚΣΡΗΚΖ ΔΝΔΡΓΔΗΑ Γηπισκαηηθή Δξγαζία ηνπ Φνηηεηή ηνπ ηκήκαηνο Ζιεθηξνιόγσλ Μεραληθώλ θαη Σερλνινγίαο Ζιεθηξνληθώλ
Αξιοποίηση Φυσικών Αντιοξειδωτικών στην Εκτροφή των Αγροτικών
 Ζώων για Παραγωγή Προϊόντων Ποιότητας» 1 Αξιοποίηση Φυσικών Αντιοξειδωτικών στην Εκτροφή των Αγροτικών Ζώων για Παραγωγή Προϊόντων Ποιότητας Γεωπονικό Πανεπιστήμιο Αθηνών Εργαστήριο Ζωοτεχνίας MIS 380231
Ζώων για Παραγωγή Προϊόντων Ποιότητας» 1 Αξιοποίηση Φυσικών Αντιοξειδωτικών στην Εκτροφή των Αγροτικών Ζώων για Παραγωγή Προϊόντων Ποιότητας Γεωπονικό Πανεπιστήμιο Αθηνών Εργαστήριο Ζωοτεχνίας MIS 380231
Congruence Classes of Invertible Matrices of Order 3 over F 2
 International Journal of Algebra, Vol. 8, 24, no. 5, 239-246 HIKARI Ltd, www.m-hikari.com http://dx.doi.org/.2988/ija.24.422 Congruence Classes of Invertible Matrices of Order 3 over F 2 Ligong An and
International Journal of Algebra, Vol. 8, 24, no. 5, 239-246 HIKARI Ltd, www.m-hikari.com http://dx.doi.org/.2988/ija.24.422 Congruence Classes of Invertible Matrices of Order 3 over F 2 Ligong An and
HOMEWORK 4 = G. In order to plot the stress versus the stretch we define a normalized stretch:
 HOMEWORK 4 Problem a For the fast loading case, we want to derive the relationship between P zz and λ z. We know that the nominal stress is expressed as: P zz = ψ λ z where λ z = λ λ z. Therefore, applying
HOMEWORK 4 Problem a For the fast loading case, we want to derive the relationship between P zz and λ z. We know that the nominal stress is expressed as: P zz = ψ λ z where λ z = λ λ z. Therefore, applying
ΠΕΡΙΛΗΨΗ. Εισαγωγή. Σκοπός
 ΠΕΡΙΛΗΨΗ Εισαγωγή Η παιδική παχυσαρκία έχει φτάσει σε επίπεδα επιδημίας στις μέρες μας. Μαστίζει παιδιά από μικρές ηλικίες μέχρι και σε εφήβους. Συντείνουν αρκετοί παράγοντες που ένα παιδί γίνεται παχύσαρκο
ΠΕΡΙΛΗΨΗ Εισαγωγή Η παιδική παχυσαρκία έχει φτάσει σε επίπεδα επιδημίας στις μέρες μας. Μαστίζει παιδιά από μικρές ηλικίες μέχρι και σε εφήβους. Συντείνουν αρκετοί παράγοντες που ένα παιδί γίνεται παχύσαρκο
Μελέτη των μεταβολών των χρήσεων γης στο Ζαγόρι Ιωαννίνων 0
 Μελέτη των μεταβολών των χρήσεων γης στο Ζαγόρι Ιωαννίνων 0 ΕΘΝΙΚΟ ΜΕΤΣΟΒΙΟ ΠΟΛΥΤΕΧΝΕΙΟ ΔΙΕΠΙΣΤΗΜΟΝΙΚΟ - ΔΙΑΤΜΗΜΑΤΙΚΟ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ (Δ.Π.Μ.Σ.) "ΠΕΡΙΒΑΛΛΟΝ ΚΑΙ ΑΝΑΠΤΥΞΗ" 2 η ΚΑΤΕΥΘΥΝΣΗ
Μελέτη των μεταβολών των χρήσεων γης στο Ζαγόρι Ιωαννίνων 0 ΕΘΝΙΚΟ ΜΕΤΣΟΒΙΟ ΠΟΛΥΤΕΧΝΕΙΟ ΔΙΕΠΙΣΤΗΜΟΝΙΚΟ - ΔΙΑΤΜΗΜΑΤΙΚΟ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ (Δ.Π.Μ.Σ.) "ΠΕΡΙΒΑΛΛΟΝ ΚΑΙ ΑΝΑΠΤΥΞΗ" 2 η ΚΑΤΕΥΘΥΝΣΗ
ΘΕΩΡΗΤΙΚΗ ΚΑΙ ΠΕΙΡΑΜΑΤΙΚΗ ΙΕΡΕΥΝΗΣΗ ΤΗΣ ΙΕΡΓΑΣΙΑΣ ΣΚΛΗΡΥΝΣΗΣ ΙΑ ΛΕΙΑΝΣΕΩΣ
 ΠΑΝΕΠΙΣΤΗΜΙΟ ΠΑΤΡΩΝ ΠΟΛΥΤΕΧΝΙΚΗ ΣΧΟΛΗ ΤΜΗΜΑ ΜΗΧΑΝΟΛΟΓΩΝ ΚΑΙ ΑΕΡΟΝΑΥΠΗΓΩΝ ΜΗΧΑΝΙΚΩΝ ΕΡΓΑΣΤΗΡΙΟ ΣΥΣΤΗΜΑΤΩΝ ΠΑΡΑΓΩΓΗΣ ΚΑΙ ΑΥΤΟΜΑΤΙΣΜΟΥ / ΥΝΑΜΙΚΗΣ & ΘΕΩΡΙΑΣ ΜΗΧΑΝΩΝ ΙΕΥΘΥΝΤΗΣ: Καθηγητής Γ. ΧΡΥΣΟΛΟΥΡΗΣ Ι ΑΚΤΟΡΙΚΗ
ΠΑΝΕΠΙΣΤΗΜΙΟ ΠΑΤΡΩΝ ΠΟΛΥΤΕΧΝΙΚΗ ΣΧΟΛΗ ΤΜΗΜΑ ΜΗΧΑΝΟΛΟΓΩΝ ΚΑΙ ΑΕΡΟΝΑΥΠΗΓΩΝ ΜΗΧΑΝΙΚΩΝ ΕΡΓΑΣΤΗΡΙΟ ΣΥΣΤΗΜΑΤΩΝ ΠΑΡΑΓΩΓΗΣ ΚΑΙ ΑΥΤΟΜΑΤΙΣΜΟΥ / ΥΝΑΜΙΚΗΣ & ΘΕΩΡΙΑΣ ΜΗΧΑΝΩΝ ΙΕΥΘΥΝΤΗΣ: Καθηγητής Γ. ΧΡΥΣΟΛΟΥΡΗΣ Ι ΑΚΤΟΡΙΚΗ
ΠΟΛΥΤΕΧΝΕΙΟ ΚΡΗΤΗΣ ΣΧΟΛΗ ΜΗΧΑΝΙΚΩΝ ΠΕΡΙΒΑΛΛΟΝΤΟΣ
 ΠΟΛΥΤΕΧΝΕΙΟ ΚΡΗΤΗΣ ΣΧΟΛΗ ΜΗΧΑΝΙΚΩΝ ΠΕΡΙΒΑΛΛΟΝΤΟΣ Τομέας Περιβαλλοντικής Υδραυλικής και Γεωπεριβαλλοντικής Μηχανικής (III) Εργαστήριο Γεωπεριβαλλοντικής Μηχανικής TECHNICAL UNIVERSITY OF CRETE SCHOOL of
ΠΟΛΥΤΕΧΝΕΙΟ ΚΡΗΤΗΣ ΣΧΟΛΗ ΜΗΧΑΝΙΚΩΝ ΠΕΡΙΒΑΛΛΟΝΤΟΣ Τομέας Περιβαλλοντικής Υδραυλικής και Γεωπεριβαλλοντικής Μηχανικής (III) Εργαστήριο Γεωπεριβαλλοντικής Μηχανικής TECHNICAL UNIVERSITY OF CRETE SCHOOL of
ΓΕΩΠΟΝΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΑΘΗΝΩΝ ΤΜΗΜΑ ΑΓΡΟΤΙΚΗΣ ΟΙΚΟΝΟΜΙΑΣ & ΑΝΑΠΤΥΞΗΣ
 ΓΕΩΠΟΝΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΑΘΗΝΩΝ ΤΜΗΜΑ ΑΓΡΟΤΙΚΗΣ ΟΙΚΟΝΟΜΙΑΣ & ΑΝΑΠΤΥΞΗΣ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ «ΟΛΟΚΛΗΡΩΜΕΝΗ ΑΝΑΠΤΥΞΗ & ΔΙΑΧΕΙΡΙΣΗ ΤΟΥ ΑΓΡΟΤΙΚΟΥ ΧΩΡΟΥ» ΜΕΤΑΠΤΥΧΙΑΚΗ ΕΡΓΑΣΙΑ «Οικονομετρική διερεύνηση
ΓΕΩΠΟΝΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΑΘΗΝΩΝ ΤΜΗΜΑ ΑΓΡΟΤΙΚΗΣ ΟΙΚΟΝΟΜΙΑΣ & ΑΝΑΠΤΥΞΗΣ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ «ΟΛΟΚΛΗΡΩΜΕΝΗ ΑΝΑΠΤΥΞΗ & ΔΙΑΧΕΙΡΙΣΗ ΤΟΥ ΑΓΡΟΤΙΚΟΥ ΧΩΡΟΥ» ΜΕΤΑΠΤΥΧΙΑΚΗ ΕΡΓΑΣΙΑ «Οικονομετρική διερεύνηση
Research on Economics and Management
 36 5 2015 5 Research on Economics and Management Vol. 36 No. 5 May 2015 490 490 F323. 9 A DOI:10.13502/j.cnki.issn1000-7636.2015.05.007 1000-7636 2015 05-0052 - 10 2008 836 70% 1. 2 2010 1 2 3 2015-03
36 5 2015 5 Research on Economics and Management Vol. 36 No. 5 May 2015 490 490 F323. 9 A DOI:10.13502/j.cnki.issn1000-7636.2015.05.007 1000-7636 2015 05-0052 - 10 2008 836 70% 1. 2 2010 1 2 3 2015-03
Inferring regulatory subnetworks through the analysis of genome-wide expression profiles
 NATIONAL AND KAPODISTRIAN UNIVERSITY OF ATHENS SCHOOL OF SCIENCE DEPARTMENT OF INFORMATICS AND TELECOMMUNICATIONS POSTGRADUATE PROGRAM "INFORMATION TECHNOLOGIES IN MEDICINE AND BIOLOGY" MASTER S THESIS
NATIONAL AND KAPODISTRIAN UNIVERSITY OF ATHENS SCHOOL OF SCIENCE DEPARTMENT OF INFORMATICS AND TELECOMMUNICATIONS POSTGRADUATE PROGRAM "INFORMATION TECHNOLOGIES IN MEDICINE AND BIOLOGY" MASTER S THESIS
Πτυχιακή Εργασία. Παραδοσιακά Προϊόντα Διατροφική Αξία και η Πιστοποίηση τους
 ΑΛΕΞΑΝΔΡΕΙΟ ΤΕΧΝΟΛΟΓΙΚΟ ΕΚΠΑΙΔΕΥΤΙΚΟ ΙΔΡΥΜΑ ΣΧΟΛΗ ΤΕΧΝΟΛΟΓΙΑΣ ΤΡΟΦΙΜΩΝ ΚΑΙ ΔΙΑΤΡΟΦΗΣ ΤΜΗΜΑ ΔΙΑΤΡΟΦΗΣ ΚΑΙ ΔΙΑΙΤΟΛΟΓΙΑΣ Πτυχιακή Εργασία Παραδοσιακά Προϊόντα Διατροφική Αξία και η Πιστοποίηση τους Εκπόνηση:
ΑΛΕΞΑΝΔΡΕΙΟ ΤΕΧΝΟΛΟΓΙΚΟ ΕΚΠΑΙΔΕΥΤΙΚΟ ΙΔΡΥΜΑ ΣΧΟΛΗ ΤΕΧΝΟΛΟΓΙΑΣ ΤΡΟΦΙΜΩΝ ΚΑΙ ΔΙΑΤΡΟΦΗΣ ΤΜΗΜΑ ΔΙΑΤΡΟΦΗΣ ΚΑΙ ΔΙΑΙΤΟΛΟΓΙΑΣ Πτυχιακή Εργασία Παραδοσιακά Προϊόντα Διατροφική Αξία και η Πιστοποίηση τους Εκπόνηση:
Οι επιδόσεις Ελλήνων στο Mini Mental State Examination με βάση την ηλικία και τη νοητική κατάσταση από την παιδική στην τρίτη ηλικία.
 CLINICAL TRIALS ΚΛΙΝΙΚΕΣ ΜΕΛΕΤΕΣ 25 Οι επιδόσεις Ελλήνων στο Mini Mental State Examination με βάση την ηλικία και τη νοητική κατάσταση από την παιδική στην τρίτη ηλικία. Τσάνταλη, Ε. 1,2,3, Οικονομίδης,
CLINICAL TRIALS ΚΛΙΝΙΚΕΣ ΜΕΛΕΤΕΣ 25 Οι επιδόσεις Ελλήνων στο Mini Mental State Examination με βάση την ηλικία και τη νοητική κατάσταση από την παιδική στην τρίτη ηλικία. Τσάνταλη, Ε. 1,2,3, Οικονομίδης,
ΜΟΡΙΑΚΕΣ ΜΕΘΟΔΟΙ ΚΡΙΤΗΡΙΑ ΕΠΙΛΟΓΗΣ ΑΞΙΟΛΟΓΗΣΗ
 - - ό ό : Ω ί ό ί ώ.. ά ή ί ός ή ς ύ ί ί ή έ ώ ό. ή ς ά ς ής ό ός ά ς ή έ ός ή ς ί ής ί ς. ό έ ώ - ώ ή ής ή ς- ί ά ά ς ί ς. ά ί ί έ έ ά ύ ή, ό ί ά, ό ό ά έ ά ής ί ύ George Wald ή ί έ ς ί ύ ό ς ί ς ά έ
- - ό ό : Ω ί ό ί ώ.. ά ή ί ός ή ς ύ ί ί ή έ ώ ό. ή ς ά ς ής ό ός ά ς ή έ ός ή ς ί ής ί ς. ό έ ώ - ώ ή ής ή ς- ί ά ά ς ί ς. ά ί ί έ έ ά ύ ή, ό ί ά, ό ό ά έ ά ής ί ύ George Wald ή ί έ ς ί ύ ό ς ί ς ά έ
ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΤΜΗΜΑ ΟΔΟΝΤΙΑΤΡΙΚΗΣ ΕΡΓΑΣΤΗΡΙΟ ΟΔΟΝΤΙΚΗΣ ΚΑΙ ΑΝΩΤΕΡΑΣ ΠΡΟΣΘΕΤΙΚΗΣ
 ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΤΜΗΜΑ ΟΔΟΝΤΙΑΤΡΙΚΗΣ ΕΡΓΑΣΤΗΡΙΟ ΟΔΟΝΤΙΚΗΣ ΚΑΙ ΑΝΩΤΕΡΑΣ ΠΡΟΣΘΕΤΙΚΗΣ ΣΥΓΚΡΙΤΙΚΗ ΜΕΛΕΤΗ ΤΗΣ ΣΥΓΚΡΑΤΗΤΙΚΗΣ ΙΚΑΝΟΤΗΤΑΣ ΟΡΙΣΜΕΝΩΝ ΠΡΟΚΑΤΑΣΚΕΥΑΣΜΕΝΩΝ ΣΥΝΔΕΣΜΩΝ ΑΚΡΙΒΕΙΑΣ
ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΤΜΗΜΑ ΟΔΟΝΤΙΑΤΡΙΚΗΣ ΕΡΓΑΣΤΗΡΙΟ ΟΔΟΝΤΙΚΗΣ ΚΑΙ ΑΝΩΤΕΡΑΣ ΠΡΟΣΘΕΤΙΚΗΣ ΣΥΓΚΡΙΤΙΚΗ ΜΕΛΕΤΗ ΤΗΣ ΣΥΓΚΡΑΤΗΤΙΚΗΣ ΙΚΑΝΟΤΗΤΑΣ ΟΡΙΣΜΕΝΩΝ ΠΡΟΚΑΤΑΣΚΕΥΑΣΜΕΝΩΝ ΣΥΝΔΕΣΜΩΝ ΑΚΡΙΒΕΙΑΣ
ΠΕΡΙΕΧΟΜΕΝΑ. Κεφάλαιο 1: Κεφάλαιο 2: Κεφάλαιο 3:
 4 Πρόλογος Η παρούσα διπλωµατική εργασία µε τίτλο «ιερεύνηση χωρικής κατανοµής µετεωρολογικών µεταβλητών. Εφαρµογή στον ελληνικό χώρο», ανατέθηκε από το ιεπιστηµονικό ιατµηµατικό Πρόγραµµα Μεταπτυχιακών
4 Πρόλογος Η παρούσα διπλωµατική εργασία µε τίτλο «ιερεύνηση χωρικής κατανοµής µετεωρολογικών µεταβλητών. Εφαρµογή στον ελληνικό χώρο», ανατέθηκε από το ιεπιστηµονικό ιατµηµατικό Πρόγραµµα Μεταπτυχιακών
Supplemental Table S1. Tumor specific networks are enriched with somatically mutated genes (taken from the database COSMIC)
 Additional file 1 Supplemental Table S1. Tumor specific networks are enriched with somatically mutated genes (taken from the database COSMIC) COSMIC genes in tumor network COSMIC genes not in tumor network
Additional file 1 Supplemental Table S1. Tumor specific networks are enriched with somatically mutated genes (taken from the database COSMIC) COSMIC genes in tumor network COSMIC genes not in tumor network
ΤΕΧΝΟΛΟΓΙΚΟ ΕΚΠΑΙ ΕΥΤΙΚΟ Ι ΡΥΜΑ ΚΡΗΤΗΣ ΣΧΟΛΗ ΙΟΙΚΗΣΗΣ ΚΑΙ ΟΙΚΟΝΟΜΙΑΣ ΤΜΗΜΑ ΙΟΙΚΗΣΗΣ ΕΠΙΧΕΙΡΗΣΕΩΝ ΠΤΥΧΙΑΚΗ ΕΡΓΑΣΙΑ
 ΤΕΧΝΟΛΟΓΙΚΟ ΕΚΠΑΙ ΕΥΤΙΚΟ Ι ΡΥΜΑ ΚΡΗΤΗΣ ΣΧΟΛΗ ΙΟΙΚΗΣΗΣ ΚΑΙ ΟΙΚΟΝΟΜΙΑΣ ΤΜΗΜΑ ΙΟΙΚΗΣΗΣ ΕΠΙΧΕΙΡΗΣΕΩΝ ΠΤΥΧΙΑΚΗ ΕΡΓΑΣΙΑ Το franchising ( δικαιόχρηση ) ως µέθοδος ανάπτυξης των επιχειρήσεων λιανικού εµπορίου
ΤΕΧΝΟΛΟΓΙΚΟ ΕΚΠΑΙ ΕΥΤΙΚΟ Ι ΡΥΜΑ ΚΡΗΤΗΣ ΣΧΟΛΗ ΙΟΙΚΗΣΗΣ ΚΑΙ ΟΙΚΟΝΟΜΙΑΣ ΤΜΗΜΑ ΙΟΙΚΗΣΗΣ ΕΠΙΧΕΙΡΗΣΕΩΝ ΠΤΥΧΙΑΚΗ ΕΡΓΑΣΙΑ Το franchising ( δικαιόχρηση ) ως µέθοδος ανάπτυξης των επιχειρήσεων λιανικού εµπορίου
ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΥΠΡΟΥ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΥΓΕΙΑΣ. Πτυχιακή εργασία ΑΓΧΟΣ ΚΑΙ ΚΑΤΑΘΛΙΨΗ ΣΕ ΓΥΝΑΙΚΕΣ ΜΕ ΚΑΡΚΙΝΟΥ ΤΟΥ ΜΑΣΤΟΥ ΜΕΤΑ ΑΠΟ ΜΑΣΤΕΚΤΟΜΗ
 ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΥΠΡΟΥ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΥΓΕΙΑΣ Πτυχιακή εργασία ΑΓΧΟΣ ΚΑΙ ΚΑΤΑΘΛΙΨΗ ΣΕ ΓΥΝΑΙΚΕΣ ΜΕ ΚΑΡΚΙΝΟΥ ΤΟΥ ΜΑΣΤΟΥ ΜΕΤΑ ΑΠΟ ΜΑΣΤΕΚΤΟΜΗ ΧΡΥΣΟΒΑΛΑΝΤΗΣ ΒΑΣΙΛΕΙΟΥ ΛΕΜΕΣΟΣ 2014 ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ
ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΥΠΡΟΥ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΥΓΕΙΑΣ Πτυχιακή εργασία ΑΓΧΟΣ ΚΑΙ ΚΑΤΑΘΛΙΨΗ ΣΕ ΓΥΝΑΙΚΕΣ ΜΕ ΚΑΡΚΙΝΟΥ ΤΟΥ ΜΑΣΤΟΥ ΜΕΤΑ ΑΠΟ ΜΑΣΤΕΚΤΟΜΗ ΧΡΥΣΟΒΑΛΑΝΤΗΣ ΒΑΣΙΛΕΙΟΥ ΛΕΜΕΣΟΣ 2014 ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ
Study of urban housing development projects: The general planning of Alexandria City
 Paper published at Alexandria Engineering Journal, vol, No, July, Study of urban housing development projects: The general planning of Alexandria City Hisham El Shimy Architecture Department, Faculty of
Paper published at Alexandria Engineering Journal, vol, No, July, Study of urban housing development projects: The general planning of Alexandria City Hisham El Shimy Architecture Department, Faculty of
ΠΑΝΕΠΙΣΤΗΜΙΟ ΠΑΤΡΩΝ ΣΧΟΛΗ ΑΝΘΡΩΠΙΣΤΙΚΩΝ ΚΑΙ ΚΟΙΝΩΝΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΤΜΗΜΑ ΦΙΛΟΛΟΓΙΑΣ
 ΠΑΝΕΠΙΣΤΗΜΙΟ ΠΑΤΡΩΝ ΣΧΟΛΗ ΑΝΘΡΩΠΙΣΤΙΚΩΝ ΚΑΙ ΚΟΙΝΩΝΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΤΜΗΜΑ ΦΙΛΟΛΟΓΙΑΣ Π.Μ.Σ: «Σύγχρονες Προσεγγίσεις στη γλώσσα και στα κείμενα» ΚΑΤΕΥΘΥΝΣΗ ΓΛΩΣΣΟΛΟΓΙΑΣ ΜΕΤΑΠΤΥΧΙΑΚΗ ΔΙΑΤΡΙΒΗ Το φωνηεντικό
ΠΑΝΕΠΙΣΤΗΜΙΟ ΠΑΤΡΩΝ ΣΧΟΛΗ ΑΝΘΡΩΠΙΣΤΙΚΩΝ ΚΑΙ ΚΟΙΝΩΝΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΤΜΗΜΑ ΦΙΛΟΛΟΓΙΑΣ Π.Μ.Σ: «Σύγχρονες Προσεγγίσεις στη γλώσσα και στα κείμενα» ΚΑΤΕΥΘΥΝΣΗ ΓΛΩΣΣΟΛΟΓΙΑΣ ΜΕΤΑΠΤΥΧΙΑΚΗ ΔΙΑΤΡΙΒΗ Το φωνηεντικό
ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΓΕΩΠΟΝΙΚΗ ΣΧΟΛΗ ΤΟΜΕΑΣ ΕΠΙΣΤΗΜΗΣ ΚΑΙ ΤΕΧΝΟΛΟΓΙΑΣ ΤΡΟΦΙΜΩΝ ΜΑΡΙΑΣ ΦΩΤΙΟΥ ΠΤΥΧΙΟΥΧΟΥ ΓΕΩΠΟΝΟΥ
 ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΓΕΩΠΟΝΙΚΗ ΣΧΟΛΗ ΤΟΜΕΑΣ ΕΠΙΣΤΗΜΗΣ ΚΑΙ ΤΕΧΝΟΛΟΓΙΑΣ ΤΡΟΦΙΜΩΝ ΜΑΡΙΑΣ ΦΩΤΙΟΥ ΠΤΥΧΙΟΥΧΟΥ ΓΕΩΠΟΝΟΥ Συγκέντρωση των ελεύθερων αµινοξέων στο αµνιακό υγρό σε σχέση µε την εβδοµάδα
ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΓΕΩΠΟΝΙΚΗ ΣΧΟΛΗ ΤΟΜΕΑΣ ΕΠΙΣΤΗΜΗΣ ΚΑΙ ΤΕΧΝΟΛΟΓΙΑΣ ΤΡΟΦΙΜΩΝ ΜΑΡΙΑΣ ΦΩΤΙΟΥ ΠΤΥΧΙΟΥΧΟΥ ΓΕΩΠΟΝΟΥ Συγκέντρωση των ελεύθερων αµινοξέων στο αµνιακό υγρό σε σχέση µε την εβδοµάδα
CHAPTER 25 SOLVING EQUATIONS BY ITERATIVE METHODS
 CHAPTER 5 SOLVING EQUATIONS BY ITERATIVE METHODS EXERCISE 104 Page 8 1. Find the positive root of the equation x + 3x 5 = 0, correct to 3 significant figures, using the method of bisection. Let f(x) =
CHAPTER 5 SOLVING EQUATIONS BY ITERATIVE METHODS EXERCISE 104 Page 8 1. Find the positive root of the equation x + 3x 5 = 0, correct to 3 significant figures, using the method of bisection. Let f(x) =
ΠΕΡΙΕΧΟΜΕΝΑ. Μάρκετινγκ Αθλητικών Τουριστικών Προορισμών 1
 ΠΑΝΕΠΙΣΤΗΜΙΟ ΑΙΓΑΙΟΥ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΤΗΣ ΔΙΟΙΚΗΣΗΣ ΤΜΗΜΑ ΔΙΟΙΚΗΣΗΣ ΕΠΙΧΕΙΡΗΣΕΩΝ ΔΙΑΤΜΗΜΑΤΙΚΟ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ «Σχεδιασμός, Διοίκηση και Πολιτική του Τουρισμού» ΜΑΡΚΕΤΙΝΓΚ ΑΘΛΗΤΙΚΩΝ ΤΟΥΡΙΣΤΙΚΩΝ
ΠΑΝΕΠΙΣΤΗΜΙΟ ΑΙΓΑΙΟΥ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΤΗΣ ΔΙΟΙΚΗΣΗΣ ΤΜΗΜΑ ΔΙΟΙΚΗΣΗΣ ΕΠΙΧΕΙΡΗΣΕΩΝ ΔΙΑΤΜΗΜΑΤΙΚΟ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ «Σχεδιασμός, Διοίκηση και Πολιτική του Τουρισμού» ΜΑΡΚΕΤΙΝΓΚ ΑΘΛΗΤΙΚΩΝ ΤΟΥΡΙΣΤΙΚΩΝ
ΙΩΑΝΝΗ ΑΘ. ΠΑΠΑΪΩΑΝΝΟΥ
 ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΣΧΟΛΗ ΓΕΩΠΟΝΙΑΣ, ΔΑΣΟΛΟΓΙΑΣ ΚΑΙ ΦΥΣΙΚΟΥ ΠΕΡΙΒΑΛΛΟΝΤΟΣ ΤΜΗΜΑ ΓΕΩΠΟΝΙΑΣ ΤΟΜΕΑΣ ΑΓΡΟΤΙΚΗΣ ΟΙΚΟΝΟΜΙΑΣ ΙΩΑΝΝΗ ΑΘ. ΠΑΠΑΪΩΑΝΝΟΥ Πτυχιούχου Γεωπόνου Κατόχου Μεταπτυχιακού
ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΣΧΟΛΗ ΓΕΩΠΟΝΙΑΣ, ΔΑΣΟΛΟΓΙΑΣ ΚΑΙ ΦΥΣΙΚΟΥ ΠΕΡΙΒΑΛΛΟΝΤΟΣ ΤΜΗΜΑ ΓΕΩΠΟΝΙΑΣ ΤΟΜΕΑΣ ΑΓΡΟΤΙΚΗΣ ΟΙΚΟΝΟΜΙΑΣ ΙΩΑΝΝΗ ΑΘ. ΠΑΠΑΪΩΑΝΝΟΥ Πτυχιούχου Γεωπόνου Κατόχου Μεταπτυχιακού
Τ.Ε.Ι. ΔΥΤΙΚΗΣ ΜΑΚΕΔΟΝΙΑΣ ΠΑΡΑΡΤΗΜΑ ΚΑΣΤΟΡΙΑΣ ΤΜΗΜΑ ΔΗΜΟΣΙΩΝ ΣΧΕΣΕΩΝ & ΕΠΙΚΟΙΝΩΝΙΑΣ
 Τ.Ε.Ι. ΔΥΤΙΚΗΣ ΜΑΚΕΔΟΝΙΑΣ ΠΑΡΑΡΤΗΜΑ ΚΑΣΤΟΡΙΑΣ ΤΜΗΜΑ ΔΗΜΟΣΙΩΝ ΣΧΕΣΕΩΝ & ΕΠΙΚΟΙΝΩΝΙΑΣ ΠΤΥΧΙΑΚΗ ΕΡΓΑΣΙΑ Η προβολή επιστημονικών θεμάτων από τα ελληνικά ΜΜΕ : Η κάλυψή τους στον ελληνικό ημερήσιο τύπο Σαραλιώτου
Τ.Ε.Ι. ΔΥΤΙΚΗΣ ΜΑΚΕΔΟΝΙΑΣ ΠΑΡΑΡΤΗΜΑ ΚΑΣΤΟΡΙΑΣ ΤΜΗΜΑ ΔΗΜΟΣΙΩΝ ΣΧΕΣΕΩΝ & ΕΠΙΚΟΙΝΩΝΙΑΣ ΠΤΥΧΙΑΚΗ ΕΡΓΑΣΙΑ Η προβολή επιστημονικών θεμάτων από τα ελληνικά ΜΜΕ : Η κάλυψή τους στον ελληνικό ημερήσιο τύπο Σαραλιώτου
Διδακτορική Διατριβή
 ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΥΠΡΟΥ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΥΓΕΙΑΣ Διδακτορική Διατριβή Η ΕΠΙΔΡΑΣΗ ΕΚΠΑΙΔΕΥΤΙΚΗΣ ΠΑΡΕΜΒΑΣΗΣ ΥΠΟΒΟΗΘΟΥΜΕΝΗΣ ΑΠΟ ΥΠΟΛΟΓΙΣΤΗ ΣΤΗΝ ΠΟΙΟΤΗΤΑ ΖΩΗΣ ΚΑΙ ΣΤΗΝ ΑΥΤΟΦΡΟΝΤΙΔΑ ΑΣΘΕΝΩΝ ΜΕ ΚΑΡΔΙΑΚΗ
ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΥΠΡΟΥ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΥΓΕΙΑΣ Διδακτορική Διατριβή Η ΕΠΙΔΡΑΣΗ ΕΚΠΑΙΔΕΥΤΙΚΗΣ ΠΑΡΕΜΒΑΣΗΣ ΥΠΟΒΟΗΘΟΥΜΕΝΗΣ ΑΠΟ ΥΠΟΛΟΓΙΣΤΗ ΣΤΗΝ ΠΟΙΟΤΗΤΑ ΖΩΗΣ ΚΑΙ ΣΤΗΝ ΑΥΤΟΦΡΟΝΤΙΔΑ ΑΣΘΕΝΩΝ ΜΕ ΚΑΡΔΙΑΚΗ
ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΥΠΡΟΥ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΥΓΕΙΑΣ ΤΜΗΜΑ ΝΟΣΗΛΕΥΤΙΚΗΣ ΠΤΥΧΙΑΚΗ ΕΡΓΑΣΙΑ ΕΠΗΡΕΑΖΕΙ ΤΗΝ ΠΡΟΛΗΨΗ ΚΑΡΚΙΝΟΥ ΤΟΥ ΜΑΣΤΟΥ
 ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΥΠΡΟΥ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΥΓΕΙΑΣ ΤΜΗΜΑ ΝΟΣΗΛΕΥΤΙΚΗΣ ΠΤΥΧΙΑΚΗ ΕΡΓΑΣΙΑ ΠΩΣ Η ΚΑΤΑΝΑΛΩΣΗ ΦΡΟΥΤΩΝ ΚΑΙ ΛΑΧΑΝΙΚΩΝ ΕΠΗΡΕΑΖΕΙ ΤΗΝ ΠΡΟΛΗΨΗ ΚΑΡΚΙΝΟΥ ΤΟΥ ΜΑΣΤΟΥ Όνομα φοιτήτριας ΚΑΛΑΠΟΔΑ ΜΑΡΚΕΛΛΑ
ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΥΠΡΟΥ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΥΓΕΙΑΣ ΤΜΗΜΑ ΝΟΣΗΛΕΥΤΙΚΗΣ ΠΤΥΧΙΑΚΗ ΕΡΓΑΣΙΑ ΠΩΣ Η ΚΑΤΑΝΑΛΩΣΗ ΦΡΟΥΤΩΝ ΚΑΙ ΛΑΧΑΝΙΚΩΝ ΕΠΗΡΕΑΖΕΙ ΤΗΝ ΠΡΟΛΗΨΗ ΚΑΡΚΙΝΟΥ ΤΟΥ ΜΑΣΤΟΥ Όνομα φοιτήτριας ΚΑΛΑΠΟΔΑ ΜΑΡΚΕΛΛΑ
Single-site association results for 136 SCARB1 genotyped variants with HDL-C.
 Table S9. Single-site association results for 136 SCARB1 genotyped variants with HDL-C. SNP Name a SNP ID b Chr12 Position c Location Amino Acid Change RegDB Score d MA, MAF Genotype Genotype Count Adjusted
Table S9. Single-site association results for 136 SCARB1 genotyped variants with HDL-C. SNP Name a SNP ID b Chr12 Position c Location Amino Acid Change RegDB Score d MA, MAF Genotype Genotype Count Adjusted
A Systematic Review of Procalcitonin for Early Detection of Septicemia of Newborn
 56 [ ] 167-3619(009)03-056-05 Journal of Tropical Medicine Vol.9 No.3 Mar.009 * ( 510515) (PCT) CNKI VIP PCT QUADAS Metadisc1.4 DOR (SROC) SROC (AUC) Review Manager5 Meta 8 ; 7 8 (χ =3.8P=0.858I =0.0%)
56 [ ] 167-3619(009)03-056-05 Journal of Tropical Medicine Vol.9 No.3 Mar.009 * ( 510515) (PCT) CNKI VIP PCT QUADAS Metadisc1.4 DOR (SROC) SROC (AUC) Review Manager5 Meta 8 ; 7 8 (χ =3.8P=0.858I =0.0%)
1 h, , CaCl 2. pelamis) 58.1%, (Headspace solid -phase microextraction and gas chromatography -mass spectrometry,hs -SPME - Vol. 15 No.
 2 0 1 5 7 Journal of Chinese Institute of Food Science and Technology Vol. 15 No. 7 Jul. 2 0 1 5 1,2 1 1 1 1* ( 1 315211 2 315502) : ( ) : (E-Nose), - : GC-MS, 20.98% 2.5%, 53.15% 11.2%, 0. 51% 71.86%
2 0 1 5 7 Journal of Chinese Institute of Food Science and Technology Vol. 15 No. 7 Jul. 2 0 1 5 1,2 1 1 1 1* ( 1 315211 2 315502) : ( ) : (E-Nose), - : GC-MS, 20.98% 2.5%, 53.15% 11.2%, 0. 51% 71.86%
High mobility group 1 HMG1
 Vol. 29, pp.705 ~ 711, 2001 High mobility group 1 HMG1 13 12 20 anti-neutrophil cytoplasmic antibodies, ANCA ANCA 1982 Davies 1980 1 high mobility group HMG1 HMG2 30 kd high mobility group HMGHMG HMG1
Vol. 29, pp.705 ~ 711, 2001 High mobility group 1 HMG1 13 12 20 anti-neutrophil cytoplasmic antibodies, ANCA ANCA 1982 Davies 1980 1 high mobility group HMG1 HMG2 30 kd high mobility group HMGHMG HMG1
5.4 The Poisson Distribution.
 The worst thing you can do about a situation is nothing. Sr. O Shea Jackson 5.4 The Poisson Distribution. Description of the Poisson Distribution Discrete probability distribution. The random variable
The worst thing you can do about a situation is nothing. Sr. O Shea Jackson 5.4 The Poisson Distribution. Description of the Poisson Distribution Discrete probability distribution. The random variable
Code Breaker. TEACHER s NOTES
 TEACHER s NOTES Time: 50 minutes Learning Outcomes: To relate the genetic code to the assembly of proteins To summarize factors that lead to different types of mutations To distinguish among positive,
TEACHER s NOTES Time: 50 minutes Learning Outcomes: To relate the genetic code to the assembly of proteins To summarize factors that lead to different types of mutations To distinguish among positive,
AΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΠΟΛΥΤΕΧΝΙΚΗ ΣΧΟΛΗ ΤΜΗΜΑ ΠΟΛΙΤΙΚΩΝ ΜΗΧΑΝΙΚΩΝ
 AΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΠΟΛΥΤΕΧΝΙΚΗ ΣΧΟΛΗ ΤΜΗΜΑ ΠΟΛΙΤΙΚΩΝ ΜΗΧΑΝΙΚΩΝ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ ΕΙΔΙΚΕΥΣΗΣ ΠΡΟΣΤΑΣΙΑ ΠΕΡΙΒΑΛΛΟΝΤΟΣ ΚΑΙ ΒΙΩΣΙΜΗ ΑΝΑΠΤΥΞΗ ΔΙΕΡΕΥΝΗΣΗ ΤΩΝ ΠΙΕΣΕΩΝ ΣΤΟ ΠΕΡΙΒΑΛΛΟΝ
AΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΠΟΛΥΤΕΧΝΙΚΗ ΣΧΟΛΗ ΤΜΗΜΑ ΠΟΛΙΤΙΚΩΝ ΜΗΧΑΝΙΚΩΝ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ ΕΙΔΙΚΕΥΣΗΣ ΠΡΟΣΤΑΣΙΑ ΠΕΡΙΒΑΛΛΟΝΤΟΣ ΚΑΙ ΒΙΩΣΙΜΗ ΑΝΑΠΤΥΞΗ ΔΙΕΡΕΥΝΗΣΗ ΤΩΝ ΠΙΕΣΕΩΝ ΣΤΟ ΠΕΡΙΒΑΛΛΟΝ
ΜΕΤΑΠΤΥΧΙΑΚΗ ΕΡΕΥΝΗΤΙΚΗ ΔΙΑΤΡΙΒΗ
 ΓΕΩΠΟΝΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΑΘΗΝΩΝ ΤΜΗΜΑ ΕΠΙΣΤΗΜΗΣ & ΤΕΧΝΟΛΟΓΙΑΣ ΤΡΟΦΙΜΩΝ ΕΡΓΑΣΤΗΡΙΟ ΠΟΙΟΤΙΚΟΥ ΕΛΕΓΧΟΥ & ΥΓΙΕΙΝΗΣ ΤΡΟΦΙΜΩΝ ΚΑΙ ΠΟΤΩΝ Π.Μ.Σ. «ΕΠΙΣΤΗΜΗ & ΤΕΧΝΟΛΟΓΙΑ ΤΡΟΦΙΜΩΝ & ΔΙΑΤΡΟΦΗ ΤΟΥ ΑΝΘΡΩΠΟΥ» Η ΕΠΙΔΡΑΣΗ
ΓΕΩΠΟΝΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΑΘΗΝΩΝ ΤΜΗΜΑ ΕΠΙΣΤΗΜΗΣ & ΤΕΧΝΟΛΟΓΙΑΣ ΤΡΟΦΙΜΩΝ ΕΡΓΑΣΤΗΡΙΟ ΠΟΙΟΤΙΚΟΥ ΕΛΕΓΧΟΥ & ΥΓΙΕΙΝΗΣ ΤΡΟΦΙΜΩΝ ΚΑΙ ΠΟΤΩΝ Π.Μ.Σ. «ΕΠΙΣΤΗΜΗ & ΤΕΧΝΟΛΟΓΙΑ ΤΡΟΦΙΜΩΝ & ΔΙΑΤΡΟΦΗ ΤΟΥ ΑΝΘΡΩΠΟΥ» Η ΕΠΙΔΡΑΣΗ
ΠΑΝΕΠΙΣΤΗΜΙΟ ΠΑΤΡΩΝ ΤΜΗΜΑ ΗΛΕΚΤΡΟΛΟΓΩΝ ΜΗΧΑΝΙΚΩΝ ΚΑΙ ΤΕΧΝΟΛΟΓΙΑΣ ΥΠΟΛΟΓΙΣΤΩΝ ΤΟΜΕΑΣ ΣΥΣΤΗΜΑΤΩΝ ΗΛΕΚΤΡΙΚΗΣ ΕΝΕΡΓΕΙΑΣ
 ΠΑΝΕΠΙΣΤΗΜΙΟ ΠΑΤΡΩΝ ΤΜΗΜΑ ΗΛΕΚΤΡΟΛΟΓΩΝ ΜΗΧΑΝΙΚΩΝ ΚΑΙ ΤΕΧΝΟΛΟΓΙΑΣ ΥΠΟΛΟΓΙΣΤΩΝ ΤΟΜΕΑΣ ΣΥΣΤΗΜΑΤΩΝ ΗΛΕΚΤΡΙΚΗΣ ΕΝΕΡΓΕΙΑΣ ιπλωµατική Εργασία του φοιτητή του τµήµατος Ηλεκτρολόγων Μηχανικών και Τεχνολογίας Ηλεκτρονικών
ΠΑΝΕΠΙΣΤΗΜΙΟ ΠΑΤΡΩΝ ΤΜΗΜΑ ΗΛΕΚΤΡΟΛΟΓΩΝ ΜΗΧΑΝΙΚΩΝ ΚΑΙ ΤΕΧΝΟΛΟΓΙΑΣ ΥΠΟΛΟΓΙΣΤΩΝ ΤΟΜΕΑΣ ΣΥΣΤΗΜΑΤΩΝ ΗΛΕΚΤΡΙΚΗΣ ΕΝΕΡΓΕΙΑΣ ιπλωµατική Εργασία του φοιτητή του τµήµατος Ηλεκτρολόγων Μηχανικών και Τεχνολογίας Ηλεκτρονικών
Strain gauge and rosettes
 Strain gauge and rosettes Introduction A strain gauge is a device which is used to measure strain (deformation) on an object subjected to forces. Strain can be measured using various types of devices classified
Strain gauge and rosettes Introduction A strain gauge is a device which is used to measure strain (deformation) on an object subjected to forces. Strain can be measured using various types of devices classified
«Συντήρηση αχλαδιών σε νερό. υπό την παρουσία σπόρων σιναπιού (Sinapis arvensis).»
 ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΓΕΩΠΟΝΙΚΗ ΣΧΟΛΗ ΤΟΜΕΑΣ ΕΠΙΣΤΗΜΗΣ & ΤΕΧΝΟΛΟΓΙΑΣ ΤΡΟΦΙΜΩΝ ΕΛΕΝΗ Π. ΠΑΠΑΤΣΑΡΟΥΧΑ Πτυχιούχος Τεχνολόγος Τροφίμων της Γεωπονικής Σχολής (Α.Π.Θ.) «Συντήρηση αχλαδιών σε
ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΓΕΩΠΟΝΙΚΗ ΣΧΟΛΗ ΤΟΜΕΑΣ ΕΠΙΣΤΗΜΗΣ & ΤΕΧΝΟΛΟΓΙΑΣ ΤΡΟΦΙΜΩΝ ΕΛΕΝΗ Π. ΠΑΠΑΤΣΑΡΟΥΧΑ Πτυχιούχος Τεχνολόγος Τροφίμων της Γεωπονικής Σχολής (Α.Π.Θ.) «Συντήρηση αχλαδιών σε
Contents Part I Psychoneuroimmunology and Systems Biology Mechanisms 1 From Psychoneuroimmunology to Personalized, Systems, and Dynamical Medicine
 Contents Part I Psychoneuroimmunology and Systems Biology Mechanisms 1 From Psychoneuroimmunology to Personalized, Systems, and Dynamical Medicine... 3 1.1 Psychoneuroimmunology (PNI) and Systems Biology...
Contents Part I Psychoneuroimmunology and Systems Biology Mechanisms 1 From Psychoneuroimmunology to Personalized, Systems, and Dynamical Medicine... 3 1.1 Psychoneuroimmunology (PNI) and Systems Biology...
ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΥΠΡΟΥ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΥΓΕΙΑΣ ΤΜΗΜΑ ΝΟΣΗΛΕΥΤΙΚΗΣ
 ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΥΠΡΟΥ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΥΓΕΙΑΣ ΤΜΗΜΑ ΝΟΣΗΛΕΥΤΙΚΗΣ ΚΑΠΝΙΣΤΙΚΕΣ ΣΥΝΗΘΕΙΕΣ ΓΟΝΕΩΝ ΚΑΙ ΕΠΙΡΡΟΗ ΤΟΥΣ ΣΤΗΝ ΕΝΑΡΞΗ ΤΟΥ ΚΑΠΝΙΣΜΑΤΟΣ ΣΤΟΥΣ ΕΦΗΒΟΥΣ Ονοματεπώνυμο Φοιτήτριας: Χριστοφόρου Έλενα
ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΥΠΡΟΥ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΥΓΕΙΑΣ ΤΜΗΜΑ ΝΟΣΗΛΕΥΤΙΚΗΣ ΚΑΠΝΙΣΤΙΚΕΣ ΣΥΝΗΘΕΙΕΣ ΓΟΝΕΩΝ ΚΑΙ ΕΠΙΡΡΟΗ ΤΟΥΣ ΣΤΗΝ ΕΝΑΡΞΗ ΤΟΥ ΚΑΠΝΙΣΜΑΤΟΣ ΣΤΟΥΣ ΕΦΗΒΟΥΣ Ονοματεπώνυμο Φοιτήτριας: Χριστοφόρου Έλενα
Χρηματοοικονομική Ανάπτυξη, Θεσμοί και
 ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΣΧΟΛΗ ΝΟΜΙΚΩΝ, ΟΙΚΟΝΟΜΙΚΩΝ ΚΑΙ ΠΟΛΙΤΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΤΜΗΜΑ ΟΙΚΟΝΟΜΙΚΩΝ ΕΠΙΣΤΗΜΩΝ Τομέας Ανάπτυξης και Προγραμματισμού Χρηματοοικονομική Ανάπτυξη, Θεσμοί και Οικονομική
ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΣΧΟΛΗ ΝΟΜΙΚΩΝ, ΟΙΚΟΝΟΜΙΚΩΝ ΚΑΙ ΠΟΛΙΤΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΤΜΗΜΑ ΟΙΚΟΝΟΜΙΚΩΝ ΕΠΙΣΤΗΜΩΝ Τομέας Ανάπτυξης και Προγραμματισμού Χρηματοοικονομική Ανάπτυξη, Θεσμοί και Οικονομική
C.S. 430 Assignment 6, Sample Solutions
 C.S. 430 Assignment 6, Sample Solutions Paul Liu November 15, 2007 Note that these are sample solutions only; in many cases there were many acceptable answers. 1 Reynolds Problem 10.1 1.1 Normal-order
C.S. 430 Assignment 6, Sample Solutions Paul Liu November 15, 2007 Note that these are sample solutions only; in many cases there were many acceptable answers. 1 Reynolds Problem 10.1 1.1 Normal-order
Comparison of Evapotranspiration between Indigenous Vegetation and Invading Vegetation in a Bog
 J. Jpn. Soc. Soil Phys. No. +*-, p.-3.1,**0 ** * *** Comparison of Evapotranspiration between Indigenous Vegetation and Invading Vegetation in a Bog Toshiki FUJIMOTO*, Ippei IIYAMA*, Mai SAKAI*, Osamu
J. Jpn. Soc. Soil Phys. No. +*-, p.-3.1,**0 ** * *** Comparison of Evapotranspiration between Indigenous Vegetation and Invading Vegetation in a Bog Toshiki FUJIMOTO*, Ippei IIYAMA*, Mai SAKAI*, Osamu
ΕΘΝΙΚΟ ΜΕΤΣΟΒΙΟ ΠΟΛΥΤΕΧΝΕΙΟ ΣΧΟΛΗ ΠΟΛΙΤΙΚΩΝ ΜΗΧΑΝΙΚΩΝ. «Θεσμικό Πλαίσιο Φωτοβολταïκών Συστημάτων- Βέλτιστη Απόδοση Μέσω Τρόπων Στήριξης»
 ΕΘΝΙΚΟ ΜΕΤΣΟΒΙΟ ΠΟΛΥΤΕΧΝΕΙΟ ΣΧΟΛΗ ΠΟΛΙΤΙΚΩΝ ΜΗΧΑΝΙΚΩΝ ΤΟΜΕΑΣ ΑΝΘΡΩΠΙΣΤΙΚΩΝ & ΚΟΙΝΩΝΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΚΑΙ ΔΙΚΑΙΟΥ «Θεσμικό Πλαίσιο Φωτοβολταïκών Συστημάτων- Βέλτιστη Απόδοση Μέσω Τρόπων Στήριξης» Διπλωματική
ΕΘΝΙΚΟ ΜΕΤΣΟΒΙΟ ΠΟΛΥΤΕΧΝΕΙΟ ΣΧΟΛΗ ΠΟΛΙΤΙΚΩΝ ΜΗΧΑΝΙΚΩΝ ΤΟΜΕΑΣ ΑΝΘΡΩΠΙΣΤΙΚΩΝ & ΚΟΙΝΩΝΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΚΑΙ ΔΙΚΑΙΟΥ «Θεσμικό Πλαίσιο Φωτοβολταïκών Συστημάτων- Βέλτιστη Απόδοση Μέσω Τρόπων Στήριξης» Διπλωματική
ΤΜΗΜΑ ΦΥΣΙΚΩΝ ΠΟΡΩΝ & ΠΕΡΙΒΑΛΛΟΝΤΟΣ
 ΤΕΧΝΟΛΟΓΙΚΟ ΕΚΠΑΙ ΕΥΤΙΚΟ Ι ΡΥΜΑ ΚΡΗΤΗΣ ΠΑΡΑΡΤΗΜΑ ΧΑΝΙΩΝ ΤΜΗΜΑ ΦΥΣΙΚΩΝ ΠΟΡΩΝ & ΠΕΡΙΒΑΛΛΟΝΤΟΣ ΤΟΜΕΑΣ ΠΕΡΙΒΑΛΛΟΝΤΙΚΗΣ ΤΕΧΝΟΛΟΓΙΑΣ ΕΡΓΑΣΤΗΡΙΟ ΠΕΡΙΒΑΛΛΟΝΤΙΚΗΣ ΧΗΜΕΙΑΣ & ΒΙΟΧΗΜΙΚΩΝ ΙΕΡΓΑΣΙΩΝ ΠΤΥΧΙΑΚΗ ΕΡΓΑΣΙΑ
ΤΕΧΝΟΛΟΓΙΚΟ ΕΚΠΑΙ ΕΥΤΙΚΟ Ι ΡΥΜΑ ΚΡΗΤΗΣ ΠΑΡΑΡΤΗΜΑ ΧΑΝΙΩΝ ΤΜΗΜΑ ΦΥΣΙΚΩΝ ΠΟΡΩΝ & ΠΕΡΙΒΑΛΛΟΝΤΟΣ ΤΟΜΕΑΣ ΠΕΡΙΒΑΛΛΟΝΤΙΚΗΣ ΤΕΧΝΟΛΟΓΙΑΣ ΕΡΓΑΣΤΗΡΙΟ ΠΕΡΙΒΑΛΛΟΝΤΙΚΗΣ ΧΗΜΕΙΑΣ & ΒΙΟΧΗΜΙΚΩΝ ΙΕΡΓΑΣΙΩΝ ΠΤΥΧΙΑΚΗ ΕΡΓΑΣΙΑ
Correction Table for an Alcoholometer Calibrated at 20 o C
 An alcoholometer is a device that measures the concentration of ethanol in a water-ethanol mixture (often in units of %abv percent alcohol by volume). The depth to which an alcoholometer sinks in a water-ethanol
An alcoholometer is a device that measures the concentration of ethanol in a water-ethanol mixture (often in units of %abv percent alcohol by volume). The depth to which an alcoholometer sinks in a water-ethanol
ΔΗΜΟΚΡΙΤΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΡΑΚΗΣ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΑΓΩΓΗΣ
 ΔΗΜΟΚΡΙΤΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΡΑΚΗΣ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΑΓΩΓΗΣ ΤΜΗΜΑ ΕΠΙΣΤΗΜΩΝ ΕΚΠΑΙΔΕΥΣΗΣ ΣΤΗΝ ΠΡΟΣΧΟΛΙΚΗ ΗΛΙΚΙΑ ΠΤΥΧΙΑΚΗ ΕΡΓΑΣΙΑ Διαπολιτισμική Εκπαίδευση και Θρησκευτική Ετερότητα: εθνικές και θρησκευτικές
ΔΗΜΟΚΡΙΤΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΡΑΚΗΣ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΑΓΩΓΗΣ ΤΜΗΜΑ ΕΠΙΣΤΗΜΩΝ ΕΚΠΑΙΔΕΥΣΗΣ ΣΤΗΝ ΠΡΟΣΧΟΛΙΚΗ ΗΛΙΚΙΑ ΠΤΥΧΙΑΚΗ ΕΡΓΑΣΙΑ Διαπολιτισμική Εκπαίδευση και Θρησκευτική Ετερότητα: εθνικές και θρησκευτικές
Μαρία Κατσιφοδήμου. Ο ρόλος της έκκρισης HLA-G από τα ανθρώπινα έμβρυα στην επιτυχία της εξωσωματικής γονιμοποίησης. Μεταπτυχιακή Διπλωματική Εργασία
 ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΣΧΟΛΗ ΘΕΤΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΤΜΗΜΑ ΒΙΟΛΟΓΙΑΣ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ ΚΑΤΕΥΘΥΝΣΗ: ΕΦΑΡΜΟΣΜΕΝΗ ΓΕΝΕΤΙΚΗ ΚΑΙ ΒΙΟΤΕΧΝΟΛΟΓΙΑ Ο ρόλος της έκκρισης HLA-G από τα ανθρώπινα
ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΣΧΟΛΗ ΘΕΤΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΤΜΗΜΑ ΒΙΟΛΟΓΙΑΣ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ ΚΑΤΕΥΘΥΝΣΗ: ΕΦΑΡΜΟΣΜΕΝΗ ΓΕΝΕΤΙΚΗ ΚΑΙ ΒΙΟΤΕΧΝΟΛΟΓΙΑ Ο ρόλος της έκκρισης HLA-G από τα ανθρώπινα
ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΥΠΡΟΥ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΥΓΕΙΑΣ. Πτυχιακή διατριβή. Ονοματεπώνυμο: Αργυρώ Ιωάννου. Επιβλέπων καθηγητής: Δρ. Αντρέας Χαραλάμπους
 ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΥΠΡΟΥ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΥΓΕΙΑΣ Πτυχιακή διατριβή Διερεύνηση της αποτελεσματικότητας εναλλακτικών και συμπληρωματικών τεχνικών στη βελτίωση της ποιότητας της ζωής σε άτομα με καρκίνο
ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΥΠΡΟΥ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΥΓΕΙΑΣ Πτυχιακή διατριβή Διερεύνηση της αποτελεσματικότητας εναλλακτικών και συμπληρωματικών τεχνικών στη βελτίωση της ποιότητας της ζωής σε άτομα με καρκίνο
ΜΗΤΡΙΚΟΣ ΘΗΛΑΣΜΟΣ ΚΑΙ ΓΝΩΣΤΙΚΗ ΑΝΑΠΤΥΞΗ ΜΕΧΡΙ ΚΑΙ 10 ΧΡΟΝΩΝ
 ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΥΠΡΟΥ ΤΜΗΜΑ ΝΟΣΗΛΕΥΤΙΚΗΣ ΜΗΤΡΙΚΟΣ ΘΗΛΑΣΜΟΣ ΚΑΙ ΓΝΩΣΤΙΚΗ ΑΝΑΠΤΥΞΗ ΜΕΧΡΙ ΚΑΙ 10 ΧΡΟΝΩΝ ΠΤΥΧΙΑΚΗ ΕΡΓΑΣΙΑ Ονοματεπώνυμο Κεντούλλα Πέτρου Αριθμός Φοιτητικής Ταυτότητας 2008761539 Κύπρος
ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΥΠΡΟΥ ΤΜΗΜΑ ΝΟΣΗΛΕΥΤΙΚΗΣ ΜΗΤΡΙΚΟΣ ΘΗΛΑΣΜΟΣ ΚΑΙ ΓΝΩΣΤΙΚΗ ΑΝΑΠΤΥΞΗ ΜΕΧΡΙ ΚΑΙ 10 ΧΡΟΝΩΝ ΠΤΥΧΙΑΚΗ ΕΡΓΑΣΙΑ Ονοματεπώνυμο Κεντούλλα Πέτρου Αριθμός Φοιτητικής Ταυτότητας 2008761539 Κύπρος
Δυσκολίες που συναντούν οι μαθητές της Στ Δημοτικού στην κατανόηση της λειτουργίας του Συγκεντρωτικού Φακού
 ΜΟΥΡΑΤΙΔΗΣ ΧΑΡΑΛΑΜΠΟΣ Δυσκολίες που συναντούν οι μαθητές της Στ Δημοτικού στην κατανόηση της λειτουργίας του Συγκεντρωτικού Φακού Μεταπτυχιακή Εργασία Ειδίκευσης που υποβλήθηκε στο πλαίσιο του Προγράμματος
ΜΟΥΡΑΤΙΔΗΣ ΧΑΡΑΛΑΜΠΟΣ Δυσκολίες που συναντούν οι μαθητές της Στ Δημοτικού στην κατανόηση της λειτουργίας του Συγκεντρωτικού Φακού Μεταπτυχιακή Εργασία Ειδίκευσης που υποβλήθηκε στο πλαίσιο του Προγράμματος
Other Test Constructions: Likelihood Ratio & Bayes Tests
 Other Test Constructions: Likelihood Ratio & Bayes Tests Side-Note: So far we have seen a few approaches for creating tests such as Neyman-Pearson Lemma ( most powerful tests of H 0 : θ = θ 0 vs H 1 :
Other Test Constructions: Likelihood Ratio & Bayes Tests Side-Note: So far we have seen a few approaches for creating tests such as Neyman-Pearson Lemma ( most powerful tests of H 0 : θ = θ 0 vs H 1 :
Main source: "Discrete-time systems and computer control" by Α. ΣΚΟΔΡΑΣ ΨΗΦΙΑΚΟΣ ΕΛΕΓΧΟΣ ΔΙΑΛΕΞΗ 4 ΔΙΑΦΑΝΕΙΑ 1
 Main source: "Discrete-time systems and computer control" by Α. ΣΚΟΔΡΑΣ ΨΗΦΙΑΚΟΣ ΕΛΕΓΧΟΣ ΔΙΑΛΕΞΗ 4 ΔΙΑΦΑΝΕΙΑ 1 A Brief History of Sampling Research 1915 - Edmund Taylor Whittaker (1873-1956) devised a
Main source: "Discrete-time systems and computer control" by Α. ΣΚΟΔΡΑΣ ΨΗΦΙΑΚΟΣ ΕΛΕΓΧΟΣ ΔΙΑΛΕΞΗ 4 ΔΙΑΦΑΝΕΙΑ 1 A Brief History of Sampling Research 1915 - Edmund Taylor Whittaker (1873-1956) devised a
«Χρήσεις γης, αξίες γης και κυκλοφοριακές ρυθμίσεις στο Δήμο Χαλκιδέων. Η μεταξύ τους σχέση και εξέλιξη.»
 ΕΘΝΙΚΟ ΜΕΤΣΟΒΙΟ ΠΟΛΥΤΕΧΝΕΙΟ ΣΧΟΛΗ ΑΓΡΟΝΟΜΩΝ ΚΑΙ ΤΟΠΟΓΡΑΦΩΝ ΜΗΧΑΝΙΚΩΝ ΤΟΜΕΑΣ ΓΕΩΓΡΑΦΙΑΣ ΚΑΙ ΠΕΡΙΦΕΡΕΙΑΚΟΥ ΣΧΕΔΙΑΣΜΟΥ ΔΙΠΛΩΜΑΤΙΚΗ ΕΡΓΑΣΙΑ: «Χρήσεις γης, αξίες γης και κυκλοφοριακές ρυθμίσεις στο Δήμο Χαλκιδέων.
ΕΘΝΙΚΟ ΜΕΤΣΟΒΙΟ ΠΟΛΥΤΕΧΝΕΙΟ ΣΧΟΛΗ ΑΓΡΟΝΟΜΩΝ ΚΑΙ ΤΟΠΟΓΡΑΦΩΝ ΜΗΧΑΝΙΚΩΝ ΤΟΜΕΑΣ ΓΕΩΓΡΑΦΙΑΣ ΚΑΙ ΠΕΡΙΦΕΡΕΙΑΚΟΥ ΣΧΕΔΙΑΣΜΟΥ ΔΙΠΛΩΜΑΤΙΚΗ ΕΡΓΑΣΙΑ: «Χρήσεις γης, αξίες γης και κυκλοφοριακές ρυθμίσεις στο Δήμο Χαλκιδέων.
«ΑΝΑΠΣΤΞΖ ΓΠ ΚΑΗ ΥΩΡΗΚΖ ΑΝΑΛΤΖ ΜΔΣΔΩΡΟΛΟΓΗΚΩΝ ΓΔΓΟΜΔΝΩΝ ΣΟΝ ΔΛΛΑΓΗΚΟ ΥΩΡΟ»
 ΓΔΩΠΟΝΗΚΟ ΠΑΝΔΠΗΣΖΜΗΟ ΑΘΖΝΩΝ ΣΜΗΜΑ ΑΞΙΟΠΟΙΗΗ ΦΤΙΚΩΝ ΠΟΡΩΝ & ΓΕΩΡΓΙΚΗ ΜΗΥΑΝΙΚΗ ΣΟΜΕΑ ΕΔΑΦΟΛΟΓΙΑ ΚΑΙ ΓΕΩΡΓΙΚΗ ΥΗΜΕΙΑ ΕΙΔΙΚΕΤΗ: ΕΦΑΡΜΟΓΕ ΣΗ ΓΕΩΠΛΗΡΟΦΟΡΙΚΗ ΣΟΤ ΦΤΙΚΟΤ ΠΟΡΟΤ «ΑΝΑΠΣΤΞΖ ΓΠ ΚΑΗ ΥΩΡΗΚΖ ΑΝΑΛΤΖ ΜΔΣΔΩΡΟΛΟΓΗΚΩΝ
ΓΔΩΠΟΝΗΚΟ ΠΑΝΔΠΗΣΖΜΗΟ ΑΘΖΝΩΝ ΣΜΗΜΑ ΑΞΙΟΠΟΙΗΗ ΦΤΙΚΩΝ ΠΟΡΩΝ & ΓΕΩΡΓΙΚΗ ΜΗΥΑΝΙΚΗ ΣΟΜΕΑ ΕΔΑΦΟΛΟΓΙΑ ΚΑΙ ΓΕΩΡΓΙΚΗ ΥΗΜΕΙΑ ΕΙΔΙΚΕΤΗ: ΕΦΑΡΜΟΓΕ ΣΗ ΓΕΩΠΛΗΡΟΦΟΡΙΚΗ ΣΟΤ ΦΤΙΚΟΤ ΠΟΡΟΤ «ΑΝΑΠΣΤΞΖ ΓΠ ΚΑΗ ΥΩΡΗΚΖ ΑΝΑΛΤΖ ΜΔΣΔΩΡΟΛΟΓΗΚΩΝ
ΠΑΝΕΠΙΣΤΗΜΙΟ ΠΑΤΡΩΝ ΤΜΗΜΑ ΕΠΙΣΤΗΜΩΝ ΤΗΣ ΕΚΠΑΙΔΕΥΣΗΣ ΚΑΙ ΤΗΣ ΑΓΩΓΗΣ ΣΤΗΝ ΠΡΟΣΧΟΛΙΚΗ ΗΛΙΚΙΑ ΜΕΤΑΠΤΥΧΙΑΚΟΣ ΚΥΚΛΟΣ ΣΠΟΥΔΩΝ 2011-2013
 ΠΑΝΕΠΙΣΤΗΜΙΟ ΠΑΤΡΩΝ ΤΜΗΜΑ ΕΠΙΣΤΗΜΩΝ ΤΗΣ ΕΚΠΑΙΔΕΥΣΗΣ ΚΑΙ ΤΗΣ ΑΓΩΓΗΣ ΣΤΗΝ ΠΡΟΣΧΟΛΙΚΗ ΗΛΙΚΙΑ ΜΕΤΑΠΤΥΧΙΑΚΟΣ ΚΥΚΛΟΣ ΣΠΟΥΔΩΝ 2011-2013 ΜΕΤΑΠΤΥΧΙΑΚΗ ΔΙΠΛΩΜΑΤΙΚΗ ΕΡΓΑΣΙΑ «Αποτίμηση αφηγηματικών ικανοτήτων παιδιών
ΠΑΝΕΠΙΣΤΗΜΙΟ ΠΑΤΡΩΝ ΤΜΗΜΑ ΕΠΙΣΤΗΜΩΝ ΤΗΣ ΕΚΠΑΙΔΕΥΣΗΣ ΚΑΙ ΤΗΣ ΑΓΩΓΗΣ ΣΤΗΝ ΠΡΟΣΧΟΛΙΚΗ ΗΛΙΚΙΑ ΜΕΤΑΠΤΥΧΙΑΚΟΣ ΚΥΚΛΟΣ ΣΠΟΥΔΩΝ 2011-2013 ΜΕΤΑΠΤΥΧΙΑΚΗ ΔΙΠΛΩΜΑΤΙΚΗ ΕΡΓΑΣΙΑ «Αποτίμηση αφηγηματικών ικανοτήτων παιδιών
ΠΑΡΑΜΕΤΡΟΙ ΕΠΗΡΕΑΣΜΟΥ ΤΗΣ ΑΝΑΓΝΩΣΗΣ- ΑΠΟΚΩΔΙΚΟΠΟΙΗΣΗΣ ΤΗΣ BRAILLE ΑΠΟ ΑΤΟΜΑ ΜΕ ΤΥΦΛΩΣΗ
 ΠΑΝΕΠΙΣΤΗΜΙΟ ΜΑΚΕΔΟΝΙΑΣ ΟΙΚΟΝΟΜΙΚΩΝ ΚΑΙ ΚΟΙΝΩΝΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΤΜΗΜΑ ΕΚΠΑΙΔΕΥΤΙΚΗΣ ΚΑΙ ΚΟΙΝΩΝΙΚΗΣ ΠΟΛΙΤΙΚΗΣ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ ΠΑΡΑΜΕΤΡΟΙ ΕΠΗΡΕΑΣΜΟΥ ΤΗΣ ΑΝΑΓΝΩΣΗΣ- ΑΠΟΚΩΔΙΚΟΠΟΙΗΣΗΣ ΤΗΣ BRAILLE
ΠΑΝΕΠΙΣΤΗΜΙΟ ΜΑΚΕΔΟΝΙΑΣ ΟΙΚΟΝΟΜΙΚΩΝ ΚΑΙ ΚΟΙΝΩΝΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΤΜΗΜΑ ΕΚΠΑΙΔΕΥΤΙΚΗΣ ΚΑΙ ΚΟΙΝΩΝΙΚΗΣ ΠΟΛΙΤΙΚΗΣ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ ΠΑΡΑΜΕΤΡΟΙ ΕΠΗΡΕΑΣΜΟΥ ΤΗΣ ΑΝΑΓΝΩΣΗΣ- ΑΠΟΚΩΔΙΚΟΠΟΙΗΣΗΣ ΤΗΣ BRAILLE
Comparison of carbon-sulfur and carbon-amine bond in therapeutic drug: -S-aromatic heterocyclic podophyllum derivatives display antitumor activity
 Comparison of carbon-sulfur and carbon-amine bond in therapeutic drug: -S-aromatic heterocyclic podophyllum derivatives display antitumor activity Jian-Long Li 1,a, Wei Zhao 1,a, Chen Zhou 1,a, Ya-Xuan
Comparison of carbon-sulfur and carbon-amine bond in therapeutic drug: -S-aromatic heterocyclic podophyllum derivatives display antitumor activity Jian-Long Li 1,a, Wei Zhao 1,a, Chen Zhou 1,a, Ya-Xuan
Reaction of a Platinum Electrode for the Measurement of Redox Potential of Paddy Soil
 J. Jpn. Soc. Soil Phys. No. +*0, p.- +*,**1 Eh * ** Reaction of a Platinum Electrode for the Measurement of Redox Potential of Paddy Soil Daisuke MURAKAMI* and Tatsuaki KASUBUCHI** * The United Graduate
J. Jpn. Soc. Soil Phys. No. +*0, p.- +*,**1 Eh * ** Reaction of a Platinum Electrode for the Measurement of Redox Potential of Paddy Soil Daisuke MURAKAMI* and Tatsuaki KASUBUCHI** * The United Graduate
Resurvey of Possible Seismic Fissures in the Old-Edo River in Tokyo
 Bull. Earthq. Res. Inst. Univ. Tokyo Vol. 2.,**3 pp.,,3,.* * +, -. +, -. Resurvey of Possible Seismic Fissures in the Old-Edo River in Tokyo Kunihiko Shimazaki *, Tsuyoshi Haraguchi, Takeo Ishibe +, -.
Bull. Earthq. Res. Inst. Univ. Tokyo Vol. 2.,**3 pp.,,3,.* * +, -. +, -. Resurvey of Possible Seismic Fissures in the Old-Edo River in Tokyo Kunihiko Shimazaki *, Tsuyoshi Haraguchi, Takeo Ishibe +, -.
[1] P Q. Fig. 3.1
![[1] P Q. Fig. 3.1 [1] P Q. Fig. 3.1](/thumbs/79/80362156.jpg) 1 (a) Define resistance....... [1] (b) The smallest conductor within a computer processing chip can be represented as a rectangular block that is one atom high, four atoms wide and twenty atoms long. One
1 (a) Define resistance....... [1] (b) The smallest conductor within a computer processing chip can be represented as a rectangular block that is one atom high, four atoms wide and twenty atoms long. One
Φυτοεξυγίανση εδάφους από Cd και Pb με τα αλόφυτα: Halimione portulacoides(l.) Aellen, Tamarix parviflora (DC) και Limoniastrum monopetalum (L.
 ΠΟΛΥΤΕΧΝΕΙΟ ΚΡΗΤΗΣ ΤΜΗΜΑ ΜΗΧΑΝΙΚΩΝ ΠΕΡΙΒΑΛΛΟΝΤΟΣ Π.Μ.Σ.: Περιβαλλοντική και Υγειονομική Μηχανική Μεταπτυχιακή Διατριβή Φυτοεξυγίανση εδάφους από Cd και Pb με τα αλόφυτα: Halimione portulacoides(l.) Aellen,
ΠΟΛΥΤΕΧΝΕΙΟ ΚΡΗΤΗΣ ΤΜΗΜΑ ΜΗΧΑΝΙΚΩΝ ΠΕΡΙΒΑΛΛΟΝΤΟΣ Π.Μ.Σ.: Περιβαλλοντική και Υγειονομική Μηχανική Μεταπτυχιακή Διατριβή Φυτοεξυγίανση εδάφους από Cd και Pb με τα αλόφυτα: Halimione portulacoides(l.) Aellen,
CPT. Tsuchiya. beta. quantitative RT PCR QIAGEN IGFBP. Fect Transfection Reagent sirna. RT PCR RNA Affymetrix GeneChip Expression Array
 7 PTEN p Tsuchiya DNA Hepatocyte nuclear factor beta HNF beta HNFbeta IGFBP GLUT G Pase MODY maturity onset diabetes of the young Tsuchiya HNFbeta HNFbeta HNFbeta sirna CPT HNFbeta HNFbeta TOVG KOC ESMCAS
7 PTEN p Tsuchiya DNA Hepatocyte nuclear factor beta HNF beta HNFbeta IGFBP GLUT G Pase MODY maturity onset diabetes of the young Tsuchiya HNFbeta HNFbeta HNFbeta sirna CPT HNFbeta HNFbeta TOVG KOC ESMCAS
Study on the Strengthen Method of Masonry Structure by Steel Truss for Collapse Prevention
 33 2 2011 4 Vol. 33 No. 2 Apr. 2011 1002-8412 2011 02-0096-08 1 1 1 2 3 1. 361005 3. 361004 361005 2. 30 TU746. 3 A Study on the Strengthen Method of Masonry Structure by Steel Truss for Collapse Prevention
33 2 2011 4 Vol. 33 No. 2 Apr. 2011 1002-8412 2011 02-0096-08 1 1 1 2 3 1. 361005 3. 361004 361005 2. 30 TU746. 3 A Study on the Strengthen Method of Masonry Structure by Steel Truss for Collapse Prevention
ΔΙΕΡΕΥΝΗΣΗ ΤΗΣ ΣΕΞΟΥΑΛΙΚΗΣ ΔΡΑΣΤΗΡΙΟΤΗΤΑΣ ΤΩΝ ΓΥΝΑΙΚΩΝ ΚΑΤΑ ΤΗ ΔΙΑΡΚΕΙΑ ΤΗΣ ΕΓΚΥΜΟΣΥΝΗΣ ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΥΠΡΟΥ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΥΓΕΙΑΣ
 ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΥΠΡΟΥ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΥΓΕΙΑΣ Πτυχιακή Εργασία ΔΙΕΡΕΥΝΗΣΗ ΤΗΣ ΣΕΞΟΥΑΛΙΚΗΣ ΔΡΑΣΤΗΡΙΟΤΗΤΑΣ ΤΩΝ ΓΥΝΑΙΚΩΝ ΚΑΤΑ ΤΗ ΔΙΑΡΚΕΙΑ ΤΗΣ ΕΓΚΥΜΟΣΥΝΗΣ ΑΝΔΡΕΟΥ ΣΤΕΦΑΝΙΑ Λεμεσός 2012 i ii ΤΕΧΝΟΛΟΓΙΚΟ
ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΥΠΡΟΥ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΥΓΕΙΑΣ Πτυχιακή Εργασία ΔΙΕΡΕΥΝΗΣΗ ΤΗΣ ΣΕΞΟΥΑΛΙΚΗΣ ΔΡΑΣΤΗΡΙΟΤΗΤΑΣ ΤΩΝ ΓΥΝΑΙΚΩΝ ΚΑΤΑ ΤΗ ΔΙΑΡΚΕΙΑ ΤΗΣ ΕΓΚΥΜΟΣΥΝΗΣ ΑΝΔΡΕΟΥ ΣΤΕΦΑΝΙΑ Λεμεσός 2012 i ii ΤΕΧΝΟΛΟΓΙΚΟ
ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ ΣΤΙΣ «ΚΛΙΝΙΚΕΣ ΚΑΙ ΚΛΙΝΙΚΟΕΡΓΑΣΤΗΡΙΑΚΕΣ ΙΑΤΡΙΚΕΣ ΕΙΔΙΚΟΤΗΤΕΣ»
 ΠΑΝΕΠΙΣΤΗΜΙΟ ΠΑΤΡΩΝ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΥΓΕΙΑΣ ΤΜΗΜΑ ΙΑΤΡΙΚΗΣ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ ΣΤΙΣ «ΚΛΙΝΙΚΕΣ ΚΑΙ ΚΛΙΝΙΚΟΕΡΓΑΣΤΗΡΙΑΚΕΣ ΙΑΤΡΙΚΕΣ ΕΙΔΙΚΟΤΗΤΕΣ» ΑΡΝΗΤΙΚΗ ΡΥΘΜΙΣΗ ΤΗΣ ΜΕΤΑΒΙΒΑΣΗΣ ΤΟΥ ΣΗΜΑΤΟΣ ΤΗΣ
ΠΑΝΕΠΙΣΤΗΜΙΟ ΠΑΤΡΩΝ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΥΓΕΙΑΣ ΤΜΗΜΑ ΙΑΤΡΙΚΗΣ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ ΣΤΙΣ «ΚΛΙΝΙΚΕΣ ΚΑΙ ΚΛΙΝΙΚΟΕΡΓΑΣΤΗΡΙΑΚΕΣ ΙΑΤΡΙΚΕΣ ΕΙΔΙΚΟΤΗΤΕΣ» ΑΡΝΗΤΙΚΗ ΡΥΘΜΙΣΗ ΤΗΣ ΜΕΤΑΒΙΒΑΣΗΣ ΤΟΥ ΣΗΜΑΤΟΣ ΤΗΣ
ΠΣΤΥΙΑΚΗ ΔΡΓΑΙΑ. Μειέηε Υξόλνπ Απνζηείξσζεο Κνλζέξβαο κε Τπνινγηζηηθή Ρεπζηνδπλακηθή. Αζαλαζηάδνπ Βαξβάξα
 ΣΔΥΝΟΛΟΓΙΚΟ ΔΚΠΑΙΓΔΤΣΙΚΟ ΙΓΡΤΜΑ ΘΔΑΛΟΝΙΚΗ ΥΟΛΗ ΣΔΥΝΟΛΟΓΙΑ ΣΡΟΦΙΜΩΝ & ΓΙΑΣΡΟΦΗ ΣΜΗΜΑ ΣΔΥΝΟΛΟΓΙΑ ΣΡΟΦΙΜΩΝ ΠΣΤΥΙΑΚΗ ΔΡΓΑΙΑ Μειέηε Υξόλνπ Απνζηείξσζεο Κνλζέξβαο κε Τπνινγηζηηθή Ρεπζηνδπλακηθή Αζαλαζηάδνπ Βαξβάξα
ΣΔΥΝΟΛΟΓΙΚΟ ΔΚΠΑΙΓΔΤΣΙΚΟ ΙΓΡΤΜΑ ΘΔΑΛΟΝΙΚΗ ΥΟΛΗ ΣΔΥΝΟΛΟΓΙΑ ΣΡΟΦΙΜΩΝ & ΓΙΑΣΡΟΦΗ ΣΜΗΜΑ ΣΔΥΝΟΛΟΓΙΑ ΣΡΟΦΙΜΩΝ ΠΣΤΥΙΑΚΗ ΔΡΓΑΙΑ Μειέηε Υξόλνπ Απνζηείξσζεο Κνλζέξβαο κε Τπνινγηζηηθή Ρεπζηνδπλακηθή Αζαλαζηάδνπ Βαξβάξα
Η ΨΥΧΙΑΤΡΙΚΗ - ΨΥΧΟΛΟΓΙΚΗ ΠΡΑΓΜΑΤΟΓΝΩΜΟΣΥΝΗ ΣΤΗΝ ΠΟΙΝΙΚΗ ΔΙΚΗ
 ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΝΟΜΙΚΗ ΣΧΟΛΗ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ ΤΟΜΕΑΣ ΙΣΤΟΡΙΑΣ ΦΙΛΟΣΟΦΙΑΣ ΚΑΙ ΚΟΙΝΩΝΙΟΛΟΓΙΑΣ ΤΟΥ ΔΙΚΑΙΟΥ Διπλωματική εργασία στο μάθημα «ΚΟΙΝΩΝΙΟΛΟΓΙΑ ΤΟΥ ΔΙΚΑΙΟΥ»
ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΝΟΜΙΚΗ ΣΧΟΛΗ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ ΤΟΜΕΑΣ ΙΣΤΟΡΙΑΣ ΦΙΛΟΣΟΦΙΑΣ ΚΑΙ ΚΟΙΝΩΝΙΟΛΟΓΙΑΣ ΤΟΥ ΔΙΚΑΙΟΥ Διπλωματική εργασία στο μάθημα «ΚΟΙΝΩΝΙΟΛΟΓΙΑ ΤΟΥ ΔΙΚΑΙΟΥ»
ΙΔΡΥΜΑ. Θεσσαλονίκη, ύλα
 ΑΛΕΞΑΝΔΡΕΙΟ ΤΕΧΝΟΛΟΓΙΚΟ ΕΚΠΑΙΔΕΥΤΙΚΟ ΙΔΡΥΜΑ ΘΕΣΣΑΛΟΝΙΚΗΣ ΣΧΟΛΗ ΤΕΧΝΟΛΟΓΙΑΣ ΓΕΩΠΟΝΙΑΣ, ΤΡΟΦΙΜΩΝ & ΔΙΑΤΡΟΦΗΔ ΗΣ ΤΜΗΜΑ ΔΙΑΤΡΟΦΗΣ & ΔΙΑΙΤΟΛΟΓΙΑΣ ΠΑΡΕΜΒΑΤΙΚΟ ΠΡΟΓΡΑΜΜΑ ΔΙΑΤΡΟΦΙΚΗΣ ΑΓΩΓΗΣ ΣΤΟΝ ΔΗΜΟ ΚΑΒΑΛΑΣ Πτυχιακή
ΑΛΕΞΑΝΔΡΕΙΟ ΤΕΧΝΟΛΟΓΙΚΟ ΕΚΠΑΙΔΕΥΤΙΚΟ ΙΔΡΥΜΑ ΘΕΣΣΑΛΟΝΙΚΗΣ ΣΧΟΛΗ ΤΕΧΝΟΛΟΓΙΑΣ ΓΕΩΠΟΝΙΑΣ, ΤΡΟΦΙΜΩΝ & ΔΙΑΤΡΟΦΗΔ ΗΣ ΤΜΗΜΑ ΔΙΑΤΡΟΦΗΣ & ΔΙΑΙΤΟΛΟΓΙΑΣ ΠΑΡΕΜΒΑΤΙΚΟ ΠΡΟΓΡΑΜΜΑ ΔΙΑΤΡΟΦΙΚΗΣ ΑΓΩΓΗΣ ΣΤΟΝ ΔΗΜΟ ΚΑΒΑΛΑΣ Πτυχιακή
ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΥΠΡΟΥ ΣΧΟΛΗ ΓΕΩΤΕΧΝΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΚΑΙ ΔΙΑΧΕΙΡΙΣΗΣ ΠΕΡΙΒΑΛΛΟΝΤΟΣ. Πτυχιακή εργασία
 ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΥΠΡΟΥ ΣΧΟΛΗ ΓΕΩΤΕΧΝΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΚΑΙ ΔΙΑΧΕΙΡΙΣΗΣ ΠΕΡΙΒΑΛΛΟΝΤΟΣ Πτυχιακή εργασία ΥΔΡΟΠΟΝΙΚΗ ΚΑΛΛΙΕΡΓΕΙΑ ΔΥΟΣΜΟΥ ΣΕ ΔΙΑΦΟΡΕΤΙΚΑ ΘΡΕΠΤΙΚΑ ΔΙΑΛΥΜΑΤΑ ΕΡΑΤΩ ΝΙΚΟΛΑΪΔΟΥ Λεμεσός 2014
ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΥΠΡΟΥ ΣΧΟΛΗ ΓΕΩΤΕΧΝΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΚΑΙ ΔΙΑΧΕΙΡΙΣΗΣ ΠΕΡΙΒΑΛΛΟΝΤΟΣ Πτυχιακή εργασία ΥΔΡΟΠΟΝΙΚΗ ΚΑΛΛΙΕΡΓΕΙΑ ΔΥΟΣΜΟΥ ΣΕ ΔΙΑΦΟΡΕΤΙΚΑ ΘΡΕΠΤΙΚΑ ΔΙΑΛΥΜΑΤΑ ΕΡΑΤΩ ΝΙΚΟΛΑΪΔΟΥ Λεμεσός 2014
«ΙΕΡΕΥΝΗΣΗ ΤΩΝ ΠΑΡΑΓΟΝΤΩΝ ΠΟΥ ΕΠΙ ΡΟΥΝ ΣΤΗΝ ΑΦΟΣΙΩΣΗ ΤΟΥ ΠΕΛΑΤΗ ΣΕ ΕΠΩΝΥΜΑ ΠΡΟΪΟΝΤΑ ΤΡΟΦΙΜΩΝ. Η ΠΕΡΙΠΤΩΣΗ ΤΩΝ ΕΠΩΝΥΜΩΝ ΓΑΛΑΚΤΟΚΟΜΙΚΩΝ ΠΡΟΪΟΝΤΩΝ»
 ΓΕΩΠΟΝΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΑΘΗΝΩΝ ΤΜΗΜΑ ΑΓΡΟΤΙΚΗΣ ΟΙΚΟΝΟΜΙΑΣ ΚΑΙ ΑΝΑΠΤΥΞΗΣ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥ ΩΝ ΟΡΓΑΝΩΣΗ ΚΑΙ ΙΟΙΚΗΣΗ ΕΠΙΧΕΙΡΗΣΕΩΝ ΤΡΟΦΙΜΩΝ & ΓΕΩΡΓΙΑΣ ΣΥΝΕΡΓΑΖΟΜΕΝΟ ΤΜΗΜΑ: ΕΠΙΣΤΗΜΗΣ & ΤΕΧΝΟΛΟΓΙΑΣ
ΓΕΩΠΟΝΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΑΘΗΝΩΝ ΤΜΗΜΑ ΑΓΡΟΤΙΚΗΣ ΟΙΚΟΝΟΜΙΑΣ ΚΑΙ ΑΝΑΠΤΥΞΗΣ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥ ΩΝ ΟΡΓΑΝΩΣΗ ΚΑΙ ΙΟΙΚΗΣΗ ΕΠΙΧΕΙΡΗΣΕΩΝ ΤΡΟΦΙΜΩΝ & ΓΕΩΡΓΙΑΣ ΣΥΝΕΡΓΑΖΟΜΕΝΟ ΤΜΗΜΑ: ΕΠΙΣΤΗΜΗΣ & ΤΕΧΝΟΛΟΓΙΑΣ
Instruction Execution Times
 1 C Execution Times InThisAppendix... Introduction DL330 Execution Times DL330P Execution Times DL340 Execution Times C-2 Execution Times Introduction Data Registers This appendix contains several tables
1 C Execution Times InThisAppendix... Introduction DL330 Execution Times DL330P Execution Times DL340 Execution Times C-2 Execution Times Introduction Data Registers This appendix contains several tables
