Supplementary Materials for
|
|
|
- Εἰλείθυια Καλογιάννης
- 7 χρόνια πριν
- Προβολές:
Transcript
1 Supplementary Materials for PD-1 Marks Dysfunctional Regulatory T cells in Malignant Gliomas Daniel E. Lowther 1,, Brittany A. Goods 2,, Liliana E. Lucca 1,, Benjamin A. Lerner 1, Khadir Raddassi 1, David van Dijk 3, Amanda L. Hernandez 1, Xiangguo Duan 1,*, Murat Gunel 3.4, Vlad Coric 5, Smita Krishnaswamy 3, J. Christopher Love 2,6,, and David A. Hafler 1,6, 1 Departments of Neurology and Immunobiology, Yale School of Medicine, New Haven, Connecticut 06520, 2 Departments of Biological Engineering and Chemical Engineering, Koch Institute for Integrative Cancer Research, Massachusetts Institute of Technology, Cambridge, Massachusetts 02139, USA, 3 Department of Genetics, Yale School of Medicine, New Haven, Connecticut 06520, 4 Department of Neurosurgery, Yale School of Medicine, New Haven, Connecticut 06520, 5 Bristol-Myers Squibb, Wallingford, Connecticut, 06492, 6 The Broad Institute of MIT and Harvard, Cambridge, MA 02142, USA This PDF file includes: Materials and Methods Supplementary Figures 1 to 17 Supplementary Tables 1 to 11 References (64-68)
2
3 Supplementary Figure 1. Gating strategy for the isolation of Programmed cell death protein 1 (PD-1) high and PD-1 - FoxP3 + Tregs. a, Representative plot showing gates for Tregs (CD25 hi CD127 lo ) and Teffs (CD25 lo CD127 + ) among total CD4 + cells, and gates for PD-1 high Tregs and Teffs based on isotype control b, quantification of the frequency of PD-1 high cells among Tregs and Teffs. N = 9, mean +/- S.D., paired Student s t test. c, Representative FoxP3 staining of total Teff (black line), total Tregs (purple line), PD-1- Tregs (blue line) and PD-1 high Tregs (red line). d, Frequency of Foxp3% cells among total, PD-1 high and PD-1 - Treg gates for 8 individuals, mean + S.D, paired Student s t test. 36
4 Supplementary Figure 2. Blocking IFNγ doesn t restore suppression activity in Programmed cell death protein 1 (PD-1) high Tregs. a, Proliferation of T effectors co-cultured with PD-1 - or PD-1 high Tregs (Teff/Treg ratio 1:2) in the presence of a blocking anti-ifnγ antibody (grey line) or an isotype control (black line). Treg and Teff were stimulated for 4 days with acd3/acd28/acd2. b, Percentage of suppression of 4 individuals calculated from the percentage of divided cells, or c, division index. For b, c Mean + S.D, Student s t test
5 Supplementary Figure 3. Programmed cell death protein 1 (PD-1) high Tregs display reduced proliferation in vitro. Proliferation of sorted Tregs stimulated for 4 days with acd3/acd28/acd2 in the presence of IL-2, assessed as dilution of CFSE. CFSE low Tregs were gated among live FoxP3 + cells. Mean + S.D. of 4 individuals. 38
6 Supplementary Figure 4. Blocking the Programmed cell death protein 1 (PD-1)- Programmed cell death ligand 1 (PD-L1) axis does not restore suppression activity in PD- 1 high Tregs. a, Proliferation of T effectors co-cultured with PD-1 high Tregs (Teff/Treg ratio 1:2) in the presence of blocking antibodies for PD-1, PD-L1, or both in combination. Treg and Teff were stimulated for 4 days with acd3/acd28/acd2. b, Division index of T effectors alone (black dots) or co-cultured with PD-1 high Tregs (white dots) of 4 individuals. Mean + S.D., One-way Anova (b) and Two-way ANOVA (c) with Sidak multiple comparison correction. 39
7 Supplementary Figure 5. Quality control metrics of RNA-sequencing libraries. a, Indicated quality metrics for each library are shown, mean and standard deviation are shown. b, Ratio metrics for each library are shown, where healthy control libraries are highlighted in red. Mean and standard deviation are shown. c, Boxplots of select transcripts are shown. Bounds of the boxes: 25 th and 75 th percentile; line within the box: median; top and bottom whiskers: maximum and minimum values
8 Supplementary Figure 6. Fully annotated heatmap of functional cluster shown in Figure 2B
9 Supplementary Figure 7. Characterization of Programmed cell death protein 1 (PD-1) high and PD-1 CD4 Treg. Expression of selected cell markers within PD-1 high and PD-1 Tregs (CD4 + CD25 + CD127 Foxp3 + ). Bounds of the boxes: 25 th and 75 th percentile; line within the box: median; top and bottom whiskers: maximum and minimum values
10 Supplementary Figure 8. Example gene sets enrichment analysis results for healthy control Programmed cell death protein 1 (PD-1) high RNAseq libraries with select genes in leading edge highlighted. a, Apoptosis pathway, b, Mitotic G1 G1 S Phase, and c, Cell cycle checkpoints. Leading edge is highlighted with a yellow bar, PD-1 high with a grey bar, and PD-1 with a black bar
11 1 2 3 Supplementary Figure 9. Representative staining for Programmed cell death protein 1 (PD- 1) and T-cell immunoglobulin and mucin domain-containing 3 (TIM-3) in the blood and 44
12 tumor infiltrate of patients with different WHO grade gliomas (grade II = G15, grade III = G11 grade IV = G3). a, Plots are gated on live CD4 + CD25 hi CD127 FoxP3 +. b, Representative FoxP3 staining from the same samples. Histograms are gated on live CD4 + CD25 hi CD127 45
13 Supplementary Figure 10. Treg (CD4 + CD25 + CD127 ) populations from peripheral blood of glioblastoma multiforme (GBM) patients co-cultured with CD4 effectors (CD4 + CD25 CD127 + ). Treg and Teff were cultured at a 1:2 ratio (n=4; G37-40). Statistical analyses were performed using paired student s t-test
14 Supplementary Figure 11. Scatter plot matrix of log10(counts + 1) and Pearson correlation 22 coefficient for each indicated tumor-derived sample. a, BT229 PD-1, b, BT213 PD-1, c, 23 BT215 PD-1, d, BT219 PD-1, e, BT220 PD-1, f, BT223 PD-1, g, BT228 PD-1, h, BT PD-1high, i, BT213 PD-1high, j, BT215 PD-1high, k, BT219 PD-1high, l, BT220 PD-1high, m, BT PD-1high, n, BT228 PD-1high
15 Supplementary Figure 12. Differential expression of Programmed cell death protein 1 (PD- 1) high or PD-1 - Tregs within tumor or blood. a, Venn diagram comparison of differential expression of PD-1high or PD-1 Tregs from tumor as compared to blood. Gene sets were filtered based on p-value for top differentially expressed genes and lists were compared. b, Heatmap of 41 common genes identified. Data was hierarchically clustered and k means clustered using GeneE. 48
16 Supplementary Figure 13. Differential expression of Programmed cell death protein 1 (PD- 1) high or PD-1 - Tregs within tumor or blood, unique gene sets are shown as indicated from Supplementary Figure 10a. Data was hierarchically clustered and k means clustered using GeneE. 49
17 Supplementary Figure 14. Representative gating strategy for Tregs in cytometry by time of 6 flight (CyTOF) data. Representative of four patients
18 Supplementary Figure 15. Multi-dimensional single-cell Treg data measured by cytometry by time of flight (CyTOF). Tregs were sorted in silico as CD3 + CD45 + CD8 - CD4 + CD127 + Foxp3 +. Single-cells organized in a nested structure by patient (shown on left 51
19 1 2 3 colorbar) and louvain clusters (shown on right colorbar). Data for week 1 blood, week 7 blood, and tumors are shown. Transformed data represents arcsin transformed raw values. 52
20 1 2 Supplementary 3 representation of cytometry by time of flight (CyTOF) data. a. Multidimentional tsne plot 4 colored according to week 1 blood, week 7 blood, or tumor. b. Same tsne plots as in a, but 5 colored according to protein. Figure 16. T-distributed stochastic neighbor embedding (tsne)
21 Supplementary Figure 17. Robustness analysis of conditional-density rescaled estimate of mutual information (DREMI). DREMI was computed on measured proteins against Programmed cell death protein 1 (PD-1) in blood from week 1, blood from week 7 and in the tumor at recurrence, in order to score and compare relationship strength. Here we compute the robustness of the DREMI scores by computing the standard deviation in DREMI scores in 100 trials where 10% of cells were randomly discarded. 54
22 1 Supplementary Table 1, RNA-sequencing quality control information for all samples. Full BT and Subset Sample ID Number of Cells Number of reads Mapped (%) 5'/3' 213BloodTregPD1pos E % BloodTregPD1neg E % TumorTregPD1pos E % TumorTregPD1neg E % BloodTregPD1pos E % TumorTregPD1pos E % TumorTregPD1neg E % BloodTregPD1neg E % TumorTregPD1pos E % TumorTregPD1neg E % BloodTregPD1pos E % BloodTregPD1neg E % TumorTregPD1pos E % TumorTregPD1neg E % BloodTregPD1pos E % BloodTregPD1neg E % TumorTregPD1pos E % TumorTregPD1neg E % BloodTregPD1pos E % BloodTregPD1neg E % BloodTregPD1pos E % BloodTregPD1neg E % TumorTregPD1pos E % TumorTregPD1neg E % TumorTregPD1pos E % TumorTregPD1neg E % BloodTregPD1pos E % BloodTregPD1neg E % BloodTregPD1neg E % BloodTregtotal E % BloodTregPD1pos E % BloodTregtotal E % BloodTregPD1pos E % BloodTregPD1neg E %
23 BloodTregPD1pos E % BloodTregtotal E % BloodTregPD1neg E %
24 1 2 1) high and PD-1 - Tregs from healthy donors. Supplementary Table 2, Differentially expressed genes in Programmed cell death protein 1 (PD- BloodPD1 BloodPD1ne negtreg_ BloodPD1negT BloodPD1neg gtreg_hcvs HCvsBloo reg_hcvsblood Treg_HCvsBl Transcript Gene Symbol BloodPD1pos dpd1post PD1posTreg_H oodpd1postr Treg_HC_- reg_hc_l C_pvalue eg_hc_fdr log10pval og2fc uc001qqs.3,uc001qqt.4,uc001qqu.3 LAG E E-19 uc003jsh.3,uc011cqm.2,uc011cqn.2 DEPDC1B E E-14 uc003jpm.3 GZMA E E-14 uc002wcq.4,uc010fzs.3,uc010fzt.3 PDCD E E-13 uc003cnx.4,uc003cny.1,uc010hiq.3,uc010hir.3 KIF E E-13 uc002tgc.3,uc002tgd.2,uc010fkb.3,uc010yxh.2 BUB E E-12 uc001ycb.2,uc001ycc.2,uc001ycd.3,uc001yce.1 ASB E E-12 uc002qza.3 ID E E-12 uc002ffm.3,uc002ffn.3 MAF E E-12 uc010unf.1,uc010ung.1,uc021srk.1 CTSH E E-11 uc001dhm.2,uc001dhn.3,uc001dho.3 AK E E-11 uc003tff.3,uc003tfg.3,uc010kxe.3,uc011kaz.2 ANLN E E-10 uc002bul.3,uc010bon.3,uc010boo.1,uc010urq.2,uc010 urr.1,uc021sxi.1 IGF1R E E-10 uc002wwp.1,uc010gdv.1 TPX E E-10 uc004eqd.3,uc004eqe.3,uc010nqf.3,uc010nqg.3,uc011 mtf.2,uc011mtg.2,uc011mth.2,uc011mti.2,uc011mtj.2, uc011mtl.2 PLS E E-10 uc001gtu.3,uc001gtv.3,uc001gtw.4 ASPM E E-09 uc004eun.4,uc004eup.4,uc011muk.1,uc011mul.1 SMARCA E E-09 uc003jvm.3,uc010ixb.3,uc031sjm.1 CCNB E E-09 uc001unt.3,uc001unu.3,uc001unv.3 SKA E E-09 uc010dka.1,uc010dkb.1,uc010dkc.1 TYMS E E-09 uc002oay.3,uc002oaz.3 ZBTB E E-09 uc002iup.2 NOG E E-09 uc002hrg.2,uc002hrh.2,uc002hri.2,uc002hrj.1,uc002hr k.1,uc010cvr.1 PLXDC E E-09 uc010sgt.2 KLRB E E-09 uc002teg.3,uc002teh.3,uc002tei.3,uc002tej.3,uc002tek. 4,uc002tel.3 LIMS E E-08 uc002hus.3 IGFBP E E-08 uc001uvd.3,uc010abi.3,uc010tee.1,uc010tef.2,uc010te g.1,uc021ric.1,uc021rid.1 NBEA E E-08 uc003iut.2,uc010irs.3 NEIL E E-08 uc001hiz.4,uc009xdc.3,uc010ptb.2 DTL E E-08 uc002jor.1,uc002jos.4,uc010dgi.1 MYO15B E E-08 57
25 uc001tnj.4,uc009zva.4,uc010sxf.3,uc031qjk.1 CORO1C E E-08 uc001dxx.4,uc021org.1 AMIGO E E-08 uc003sua.1,uc003sub.4,uc003sud.2,uc003sue.3,uc003s uf.2,uc003suh.3,uc003sui.3,uc003suj.3,uc003suk.3,uc0 10kud.2,uc010kue.1,uc011jya.2,uc011jyb.2,uc011jyc.2,uc011jyd.2,uc011jye.2,uc011jyf.2 HDAC E E-08 uc001tjl.3 PMCH E E-08 uc003qbw.4,uc003qbx.4,uc003qby.4,uc003qbz.1,uc01 0kfg.2,uc031spp.1 SAMD E E-07 uc002huq.3,uc002hur.1 TOP2A E E-07 uc001rvs.2,uc001rvt.2,uc001rvu.2,uc009zlm.1 RACGAP E E-07 uc003tfa.3 EEPD E E-07 uc003int.3,uc003inu.1 RNF E E-07 uc002csm.3,uc002csn.3,uc002cso.3,uc002csq.3,uc010 bsy.1,uc010uwn.2 PKMYT E E-07 uc003lzf.3,uc003lzg.3,uc003lzh.3,uc011dem.2 HMMR E E-07 uc001lke.3,uc001lkf.3,uc009yav.1,uc009yaw.1 MKI E E-07 uc002agk.3,uc002agl.3,uc002agm.3,uc002agn.3,uc010 bgj.3,uc010uhd.2,uc021snc.1 ANXA E E-07 uc003hxb.1,uc003hxc.1,uc003hxd.1 CENPE E E-07 uc002msy.1,uc002msz.1 ZNF E E-07 uc002kvs.4,uc002kvt.4,uc002kvu.4 TAF4B E E-07 uc001jld.3,uc001jle.3,uc001jlg.3,uc010qii.2,uc021prh. 1,uc031pvg.1 CDK E E-07 uc002bsl.4,uc010boe.3 FAM174B E E-07 uc003cbh.3,uc003cbi.3,uc003cbj.3 SATB E E-07 uc002teq.4,uc010fjn.3,uc010yws.2 EDAR E E-07 uc003yrx.3,uc010mdn.3,uc011lis.2,uc022bay.1 TRIB E E-07 uc001gvn.2 GPR E E-07 uc002bnm.1,uc002bnn.1,uc002bno.3,uc002bnp.1,uc00 2bnq.1,uc010bnp.1 FANCI E E-07 uc003tir.3,uc010kxv.3 BLVRA E E-07 uc003ken.4 F2R E E-07 uc001lfg.4,uc001lfk.1,uc001lfn.5,uc001lfo.1,uc010qtl. 2,uc010qtm.2,uc010qtn.2,uc010qto.2,uc010qtp.2,uc01 0qtq.2,uc021pzv.1,uc021pzw.1,uc021pzx.1,uc021pzy.1,uc021pzz.1,uc021qaa.1,uc021qab.1,uc021qac.1,uc031 pxk.1 FGFR E E-07 uc001dku.4,uc009wcm.3 SYDE E E-06 uc003wda.4 EPHA1-AS E E-06 uc002fid.3 COTL E E-06 uc002iou.4,uc002iov.4,uc002iow.3 ZNF E E-06 uc001mhk.3,uc001mhl.3,uc010rbw.2,uc010rbx.2 DENND5A E E-06 uc001zck.3,uc001zcl.3,uc001zcm.1,uc010azj.2,uc010u as.1,uc010uat.2 APBA E E-06 58
26 uc002kli.3 NDC E E-06 uc004eji.4,uc004ejj.4,uc010nod.3,uc022cbc.1,uc022cb d.1 GPRASP E E-06 uc010ywt.1 SH3RF E E-06 uc010gce.1 ISM E E-06 uc002mbo.3,uc002mbp.3,uc010duf.3,uc010xik.2 UHRF E E-06 uc003alz.3,uc003ama.3,uc010gwm.3 YWHAH E E-06 uc003cpo.4,uc010hjd.3,uc021wxb.1 CCR E E-06 uc003cdx.4,uc003cdy.4,uc010hfn.3,uc011axc.2 EOMES E E-06 uc003wcz.3 EPHA E E-06 uc001tlh.4,uc031qji.1 C12orf E E-06 uc003xep.1,uc003xeq.1,uc003xer.1,uc011lae.1 CDCA E E-06 uc001ocp.2 CDCA E E-06 uc003cpc.1,uc010hix.1,uc021www.1 CXCR E E-06 uc003pnw.3,uc010kch.3,uc011eab.2 BACH E E-06 uc001wrq.3,uc010aml.3 AKAP E E-06 uc021uec.1 EPR E E-06 uc003xsq.4,uc003xsr.4,uc010lyi.3,uc010lyj.3,uc022aur.1 PLAG E E-06 uc001scj.2,uc001sck.2,uc010soe.1 ESPL E E-06 uc010ppj.1,uc010ppk.1 KIF E E-06 uc002ktw.3,uc002ktx.1,uc002kty.3,uc002ktz.3,uc002k ua.3,uc010xap.2,uc021uid.1 RBBP E E-06 uc001zoa.3,uc001zob.3,uc010bcg.2,uc010ucx.1 LTK E E-06 uc003zzn.4,uc010mll.4,uc010mlm.4,uc011lpm.3,uc01 1lpn.3,uc011lpo.3,uc011lpp.3,uc011lpq.3,uc011lpr.3,u c011lps.3,uc022bgq.2 MELK E E-06 uc003gzm.4,uc003gzn.3,uc011bzr.2,uc011bzs.2 USP E E-06 uc001ezg.3,uc001ezh.3,uc010pdo.2,uc010pdp.2 RORC E E-06 uc001rdb.3,uc009zif.3,uc009zig.3,uc010shv.2 EPS E E-06 uc002jmz.1,uc002jna.1,uc002jnb.1 HN E E-06 uc002jef.2 PECAM E E-06 uc003idu.4,uc003idv.4 ANXA E E-06 uc003ybd.3 ZC2HC1A E E-05 uc003dly.2 PSMD6-AS E E-05 uc002ebq.3,uc002ebr.3,uc002ebs.1,uc010cam.1,uc010 can.3,uc010vfj.1 ITGAM E E-05 uc001egm.3,uc001egn.3,uc001ego.1,uc001egp.4,uc010 owy.2 CD E E-05 uc004cyb.3,uc004cyc.3,uc004cyd.3,uc004cye.3 SCML E E-05 uc001myc.3,uc001myd.3 CD E E-05 uc002mkg.3,uc010xkf.2 MYO1F E E-05 uc002asb.3,uc002asc.3,uc010bih.2,uc010bii.3,uc010uk c.2 KIF E E-05 uc003jqp.3,uc003jqq.3,uc003jqr.3,uc010iwb.3,uc010i IL6ST E E-05 59
27 wd.3,uc010iwe.1,uc010iwf.1,uc011cqk.2 uc002qvn.1,uc010evy.1 FAM72D E E-05 uc001bhe.2 E2F E E-05 uc001joz.3,uc001jpa.1,uc021prx.1 SRGN E E-05 uc003ccg.2,uc010hfc.2 UBE2E E E-05 uc003ont.3 CCDC E E-05 uc002ggi.4,uc002ggj.4,uc010cmf.3 TNK E E-05 uc002orh.1,uc010xwd.1 CEACAM E E-05 uc003ink.2,uc021xtj.1,uc021xtk.1 MND E E-05 uc001iks.1,uc001ikt.3,uc001iku.3 USP6NL E E-05 uc002lkm.4,uc010dqo.3,uc021uli.1 CD E E-05 uc002uhq.1,uc002uhs.3,uc010zdz.1,uc010zea.1,uc010z eb.2 PDK E E-05 uc004bgf.1,uc011lwx.1 KIAA E E-05 uc003ikc.3,uc003ikd.3,uc003ikf.3,uc010iov.3,uc011cic.2 SMAD E E-05 uc002fdk.3,uc002fdl.1,uc010cgr.1,uc010vmz.1 ZNRF E E-05 uc003xqk.2,uc003xql.2,uc010lxw.2,uc011ldi.2 MCM E E-05 uc001fmg.3,uc010pgq.2 SYT E E-05 uc003cen.3,uc003ceo.3,uc021wut.1 TGFBR E E-05 uc022cew.1,uc022cex.1 LINC E E-05 uc003xgg.3,uc003xgh.3,uc010luy.1 ESCO E E-05 uc001fyz.1,uc001fza.1 FCER1G E E-05 uc001dqz.4,uc010otv.2,uc010otx.2 CNN E E-05 uc001uvm.3,uc001uvn.3,uc001uvo.3,uc001uvp.2,uc00 1uvq.3,uc010ten.2 SPG E E-05 uc002taf.3,uc002tag.3,uc002tah.1,uc010fiq.1,uc010fir. 1,uc010yvr.1 AFF E E-05 uc002wlp.3,uc002wlq.3,uc010zqs.1 PCNA E E-05 uc003gpp.4,uc011bxj.3 NCAPG E E-05 uc003gol.1,uc021xmk.1 CD E E-05 uc003dyw.3,uc003dyx.3,uc003dyy.3,uc003dyz.3,uc01 0hqd.1 CD E E-05 uc002qys.3,uc002qyu.3 RNF144A E E-05 uc004ajf.1,uc004ajg.1,uc031tee.1 ANXA E E-05 uc001nnl.3,uc001nnm.3,uc010rko.2,uc021qjn.1 FAM111B E E-05 uc003vog.3,uc003voh.3,uc003voi.3,uc003voj.4,uc003v ok.2,uc010llr.1,uc010lls.1,uc010llt.3,uc010llu.1,uc010l lv.1,uc010llw.1,uc011kot.1,uc011kou.1 IRF E E-05 uc003ejd.2,uc011bkl.2 TXNRD E E-05 uc002xlb.1,uc010zwj.1 MYBL E E-05 uc003xyx.1 MSC E E-05 uc003gij.3,uc003gik.3,uc011bwb.2 EVC E E-05 uc001qez.3,uc001qfb.3,uc009zco.3,uc009zcp.3,uc009z cq.2 ARHGAP E E-05 60
28 uc001quy.3,uc009zgg.2 RIMKLB E E-05 uc003bhl.1,uc003bhm.1,uc003bhn.3,uc011aqy.2,uc011 aqz.2 GTSE E E-05 uc003kgy.1,uc010jap.2,uc011ctl.2,uc011ctm.2 DHFR E E-05 uc003lww.3,uc003lwy.3,uc003lwz.4,uc011ddr.1,uc021 ygq.1 ADAM E E-05 uc003opa.3,uc003opb.3,uc003opd.3,uc003ope.3 MOCS E E-05 uc003fli.1,uc003flj.1,uc003flp.1,uc011bqr.1,uc011bqs. 1 MCF2L E E-05 uc021vdr.1 RRM E E-05 uc003fhs.3,uc003fht.3,uc003fhu.4,uc003fhv.1 PLD E E-05 uc002aco.3,uc002acp.3,uc002acq.3,uc002acr.3 RAB27A E E-05 uc003awr.3 APOBEC3C E uc002qrh.2,uc002qri.2,uc002qrj.3,uc002qrk.1,uc002qrl.2,uc002qrm.2,uc010yhs.1,uc010yht.1 ZSCAN E uc003xtw.1 TOX E uc003upz.3 ARPC1B E uc002urx.4,uc002ury.4,uc010fsb.3 INPP E uc001mpm.3,uc001mpn.5,uc001mpo.2,uc009yhv.3 E2F E uc002fnp.3,uc002fnq.3 CPNE E uc002icp.4,uc002icq.3,uc002ict.3,uc002icu.3,uc002idc.1,uc002idd.3,uc002ide.1,uc010cyx.3,uc010cyy.1,uc01 0cyz.2,uc010cza.2,uc010czc.3,uc010whl.2,uc010whm. 2,uc010whn.2,uc010who.3,uc010whp.2,uc010whq.1,u c010whr.1,uc010whs.1,uc010wht.1 BRCA E uc002tvo.3,uc002tvp.3,uc002tvr.3,uc002tvs.3,uc002tvt.2,uc010fnn.3,uc010fno.3,uc021vqf.1,uc021vqg.1 GTDC E uc002tff.3,uc021vlz.1 LIMS E uc001hzh.3,uc009xgq.3,uc021plj.1,uc021plk.1 EXO E uc001stw.1 IFNG E uc002xle.4,uc002xlf.4,uc010ggo.3,uc010ggp.3,uc010z wk.2,uc021wdz.2,uc031rtm.1 TOX E uc001vtv.3,uc001vtw.3,uc001vty.4,uc010agx.3,uc010t kd.2,uc010tke.2 TFDP E uc003srm.3,uc003srn.5,uc003sro.5,uc003srq.4,uc003sr r.4,uc003srs.2,uc010ktr.3,uc010kts.4 ICA E uc003wnv.1,uc003wnw.1,uc003wnx.1,uc010lqu.1,uc0 11kwc.1,uc011kwd.1,uc011kwe.1 NCAPG E uc002unu.3,uc002unv.3,uc010frj.1,uc010zfl.1 ITGA E uc001pwf.3,uc001pwg.1 MCAM E uc003vez.3,uc003vfa.3,uc003vfc.3,uc003vfd.3,uc003v fe.3,uc011kmj.2,uc011kmk.2,uc022aka.1 NRCAM E uc004dyd.1,uc004dye.1 PDZD E uc003pjb.4,uc003pjc.3 TTK E uc001tyj.3,uc001tyl.3,uc001tym.3 DYNLL E
29 uc003xqh.1 CEBPD E uc004dan.3,uc004dao.3,uc004dap.3,uc004dar.3,uc004 dat.1,uc011mjt.2 ACOT E uc001crc.1,uc001crd.1,uc001cre.1,uc001crf.1,uc001cr g.1,uc010omn.1,uc010omo.1 STIL E uc001tay.3,uc001taz.3 DUSP E uc001kxa.3,uc001kxb.3,uc001kxc.3,uc001kxd.1 CALHM E uc003qvb.1,uc003qvc.1,uc003qvd.1,uc010kkk.1,uc010 kkl.1,uc011ego.1 RPS6KA E uc004aqh.3 CKS E uc001mwo.4,uc001mwp.3,uc009ykk.3,uc010rfc.2 PRR5L E uc002qot.3,uc002qou.3,uc010ygy.2,uc010ygz.2,uc010 yha.2 ZNF E uc001oht.3,uc001ohu.3,uc001ohv.3,uc009yrf.3 SLC29A E uc001zci.1,uc010azf.3,uc010azg.1,uc010azh.1,uc010az i.1,uc010uap.2 WHAMMP E uc003oef.4,uc011drf.2 KIFC E uc003giu.4,uc003giv.4,uc010idb.1,uc010idc.1,uc010id d.1,uc010ide.3,uc011bwc.2 JAKMIP E uc001mus.4,uc001mut.4,uc001muu.4,uc001muv.4,uc0 09yjx.3,uc009yjy.3,uc009yjz.3,uc009yka.3 CD E uc001hkm.3 CENPF E uc003fgi.3,uc010hwq.3,uc021xhe.1 GPR E uc001mlm.2,uc001mln.3,uc010rcr.2 PDE3B E uc002ewa.3,uc010cfe.3,uc010vlh.2 SMPD E uc001lyw.4,uc009yeh.1,uc009yei.3,uc009yej.3,uc010q yc.2,uc010qyd.2 RRM E uc001kfr.3,uc001kfs.3,uc001kft.3,uc001kfw.3,uc009xt p.3,uc010qna.2,uc010qnb.2,uc010qnc.2,uc010qnd.2,uc 010qne.2,uc031pwn.1 FAS E uc002ffw.4,uc002ffx.2,uc002ffy.4,uc010vnl.1,uc010vn m.1 CENPN E uc001vud.3,uc001vuf.3,uc001vug.3 GAS E uc001wjc.2,uc031qns.1 PPP1R3E E uc004edh.3,uc011mqr.2 ITM2A E uc003wul.3,uc003wum.3,uc003wun.3,uc003wuo.3,uc0 03wup.3,uc003wuq.3,uc003wuu.3,uc010lsc.3,uc011kx l.2 CTSB E uc003ype.3,uc010mdg.3 SNTB E uc011jwz.2 ZNF E uc001aub.3,uc001auc.3,uc001aud.4,uc001aue.1,uc009 vnm.3 DHRS E uc002yjx.2,uc021whl.1,uc031ruy.1,uc031ruz.1 NRIP E uc001qqw.1,uc001qqx.1,uc001qqy.1,uc001qra.1,uc001 qrb.1,uc009zfd.1,uc010sfn.1 GPR E
30 uc011dkv.2 ZNF204P E uc003idx.1 TMEM E uc001noq.3,uc001not.3,uc009ymv.3,uc010rla.2,uc010r lb.2 MS4A6A E uc001cgj.3 CITED E uc003cjl.3,uc021wwa.1,uc021wwb.1,uc021wwc.1,uc0 21wwd.1 CX3CR E uc001cix.3,uc001ciy.3 CDC E uc002ndi.2,uc002ndj.2,uc002ndk.1,uc002ndl.1 TPM E uc001slf.2,uc001slg.2 TIMELESS E uc001elb.4 AK E uc001xdw.3,uc001xdx.3,uc010trv.2,uc010trw.2 DACT E uc001vjt.1,uc001vju.1,uc001vjv.3,uc001vjw.1,uc001vj x.1,uc010thv.2,uc010thw.2,uc021rkq.1 LMO E uc003xlp.3,uc003xlu.3,uc003xlv.3,uc003xlw.1,uc010l wf.3,uc010lwk.3,uc011lbr.2,uc011lbs.2,uc011lbt.1,uc0 11lbu.2,uc011lbv.2,uc011lbw.2,uc011lbx.1,uc022aua.1,uc022aub.1,uc022auc.1,uc022aud.1 FGFR E uc002fjz.1,uc002fka.1,uc002fkb.3 ZCCHC E uc003hyt.2,uc003hyu.2,uc003hyv.2,uc003hyw.1,uc010 imb.2,uc011cfj.1,uc011cfk.2 LEF E uc003gxx.4,uc003gxy.1,uc010igj.3,uc011bzj.2 TXK E uc001xbs.3,uc001xbt.3 DLGAP E uc002rhr.3,uc002rhs.3,uc002rht.3 CENPA E uc001fwf.4,uc001fwg.4,uc001fwh.4,uc001fwi.4,uc001 fwj.3,uc001fwk.3,uc009wtn.3 CD E uc001kic.3 KIF E uc001ima.3,uc001imb.3,uc001imc.3 MCM E uc003icr.4,uc003ics.4,uc003ict.3,uc003icu.4 USP E uc003gsv.4,uc011bxy.3,uc021xne.2 DTHD E uc002tfw.3 LIMS E uc001qbc.3,uc001qbd.2,uc001qbe.3,uc001qbf.1,uc010 saq.2,uc010sar.2 ROBO E uc004azc.3,uc004azd.3,uc004aze.3,uc011lvc.2 TGFBR E uc002ckm.1 LMF E uc001qmu.3,uc001qmw.3,uc009zeg.3,uc010sep.2,uc01 0seq.2 RAD51AP E uc004dau.3,uc004dav.3,uc010nfv.3,uc022bty.1,uc031t ha.1 SAT E uc002the.2,uc002thf.4,uc010yxl.1 LOC E uc002niy.3,uc002niz.3,uc010ebp.3 SSBP E uc003yca.2 FABP E uc001tqq.3 ARPC E uc003kdl.4,uc003kdm.4,uc010izm.3 FAM169A E uc002jvc.3,uc002jvd.2,uc002jve.1,uc002jvf.3,uc002jv BIRC E
31 g.3,uc002jvh.3,uc002jvi.3,uc010dhk.1,uc010dhl.1 uc003nem.3,uc003nen.3,uc021ymn.1,uc021ymo.1 GMNN E uc002juw.2 TK E uc001knh.3,uc010qov.2 PGAM E uc002dhv.3,uc002dhx.3,uc002dhy.4 LOC E uc003zoe.2,uc003zof.3,uc011lne.1,uc011lnf.1 MLLT E uc003eed.3,uc003eee.4 POLQ E uc004erh.4 SLC25A E uc003wfb.2,uc003wfc.2,uc003wfd.2,uc011kug.2,uc01 1kuh.2,uc011kui.2,uc011kuj.2,uc022aov.1 EZH E uc001rxy.3,uc001rxz.3,uc001rya.3 POU6F E uc002pwj.3,uc002pwk.3 NKG E uc003fid.3,uc003fie.3,uc010hwu.2,uc021xhk.1 TNFSF E uc001rtv.4,uc001rtw.4,uc001rtx.4,uc009zlh.3,uc031qh d.1,uc031qhe.1 TROAP E uc001twr.2,uc009zws.1,uc010sza.1,uc010szb.1 WSB E uc002kag.3,uc002kak.2,uc002kal.2,uc021ufb.1 ACTG E uc003zyu.4,uc003zyv.3,uc003zyw.3,uc003zyx.3,uc010 mle.1,uc031tdr.1 RECK E uc001mok.3,uc001mol.3,uc009yho.2,uc010rdc.1,uc01 0rdd.2,uc021qep.1 LDHA E uc002ugy.4,uc002ugz.4,uc031rqa.1 CYBRD E uc001vog.3,uc021rma.1 GPR E uc003fay.1 PA2G4P E uc004bdo.1,uc004bdp.1,uc010mts.1 CTNNAL E uc002qbu.2,uc010eqq.1 ZNF E uc001bzi.3,uc009vux.3 CLSPN E uc001pav.3,uc001paw.3,uc001pax.3,uc001pay.3,uc001 paz.3,uc001pbb.3,uc001pbc.3,uc001pbd.3,uc001pbf.4, uc009yvj.3,uc010rte.2,uc010rtf.2,uc010rtg.2,uc010rth. 2,uc010rti.2,uc010rtj.2 SYTL E uc003fbc.3 LINC E uc001jlv.3,uc001jlw.3,uc001jlx.2,uc009xpf.1 RTKN E uc001zyv.3 SPPL2A E uc002uic.1,uc002uid.1,uc010zej.1,uc010zek.1 CDCA E uc001msc.2,uc001msd.3 KIF18A E uc003ldh.3,uc003ldi.3,uc003ldj.3,uc003ldl.3,uc011cyx.2,uc011cyy.2 CTNNA E uc003oer.3,uc003oes.3,uc003oet.3,uc003oeu.4,uc010jv b.3 BAK E uc002klp.3,uc002klq.3 MYOM E uc002ofq.3,uc002ofr.1 ZNF585B E uc003oon.3 KCNK E uc004ejk.3,uc004ejl.3,uc004ejm.3,uc022cay.1,uc022cb e.1,uc022cbf.1,uc022cbg.1,uc022cbh.1 GPRASP E
32 uc003vop.2,uc003voq.2 TSPAN E uc002dca.4,uc002dcb.4 CPPED E uc002rwo.4,uc002rwp.2,uc002rws.2,uc010fbm.3,uc01 0yol.2,uc010yom.2,uc021vhf.1 STON E uc003awv.3,uc003aww.3,uc003awx.3,uc003awy.3,uc0 11aog.1,uc021wpr.1 APOBEC3F E uc003fgn.1 AY E uc002xbc.2,uc002xbd.2,uc002xbe.2,uc010gey.2,uc010 zum.1 ACSS E uc002spl.2,uc002spm.2,uc010fgi.2,uc010fgj.1 CAPG E uc003qvl.3,uc003qvm.4,uc003qvn.4,uc010kkm.3 CCR E uc001ozg.3,uc001ozh.3 GAB E uc002hhn.3,uc010ctc.3,uc010wca.1,uc021tux.1 CDK5R E uc001xhn.1,uc001xho.1,uc001xhp.2,uc001xhq.1,uc010 aqh.1 PLEKHG E uc002erk.1 FBXL E uc003mib.1,uc003mic.3,uc010jkp.1,uc021yix.1 FAM153A E uc001ezl.3 S100A E uc010efz.2,uc010ega.2,uc021utz.1,uc021uua.1,uc021u ub.1,uc021uuc.1,uc021uud.1,uc021uue.1,uc021uuf.1 RASGRP E uc001udg.1,uc001udh.3,uc001udi.1,uc001udj.1,uc001u dk.1 HIP1R E uc001usu.3 LINC E uc002jlc.2,uc002jld.2,uc002jlk.3,uc010dfu.3,uc010wrb.2,uc010wrc.2,uc010wre.2 RAB E uc002ftx.3,uc031qxv.1 OVCA E uc001crw.4,uc001crx.4 BEND E uc001jzi.3,uc001jzj.3,uc001jzk.3,uc001jzl.4,uc009xru. 1 DLG E uc004ajv.4,uc004ajx.4 OSTF E uc002afz.3,uc031qsi.1 CCNB E uc001bkm.2,uc009vry.1 MAN1C E uc002rqv.3,uc021vgb.1,uc021vgc.1 HNRPLL E uc002iko.4,uc002iks.2,uc010wkd.1 ARL17A E uc003iwp.3,uc003iwq.3,uc003iwr.1 MLF1IP E uc002dbf.3,uc021tda.1 SNN E uc003ita.3,uc003itb.2,uc011ckc.1 HMGB E uc002fgt.3,uc010chg.1 PLCG E uc002jfh.3,uc002jfi.3,uc002jfj.1,uc010den.1 AXIN E uc004blh.4,uc004bli.4,uc011lyk.3,uc031tew.1,uc031te x.1 STOM E uc002tzk.4,uc002tzl.4,uc010foe.3,uc010fof.3 ACVR1C E uc003tmg.2,uc003tmh.2,uc003tmi.1,uc003tmj.2,uc010 kym.2,uc022acj.1 MYO1G E uc003fdf.1,uc003fdg.1,uc003fdh.3,uc003fdi.3,uc003fdj SMC E
33 .3,uc003fdl.3,uc010hwc.1,uc010hwd.3 uc003bdk.3 BIK E uc001ffq.3 PMVK E uc002ank.3,uc002anl.3 KIAA E uc003hvn.1,uc003hvo.1 H2AFZ E uc002ete.1,uc010vjn.1,uc010vjo.1 ATP6V0D E uc004cks.2,uc004ckt.2 NPDC E uc001efv.1,uc009wgy.1,uc021ose.1 VANGL E uc004ckj.1 CLIC E uc003kha.2,uc003khb.1,uc011ctn.2 RASGRF E uc002rqy.3 GALM E uc003cek.3,uc003cel.3,uc003cem.3,uc010hfq.3,uc010h fr.3 RBMS E uc002lbu.3,uc002lbv.3 HAUS E uc003tvv.4,uc003tvw.4,uc003tvx.4,uc011keg.2 AUTS E uc003wfr.4,uc003wft.4,uc003wfu.3 LOC E uc001yvg.3,uc010ayc.3,uc010ayd.3,uc010aye.1 WHAMMP E uc001gen.4,uc001geo.3,uc001gep.3,uc001geq.3,uc009 wvh.3,uc031prf.1 MPZL E uc004eaf.3,uc011mpx.2,uc022bys.1 CXCR E uc001mhs.3,uc001mht.3,uc001mhu.3 WEE E uc001fga.3,uc001fgb.3 CKS1B E uc001rfs.3,uc001rft.3,uc009ziz.3 ETNK E uc002eec.4 SHCBP E uc002jal.4,uc002jam.1,uc002jan.1,uc002jao.4,uc010w pe.2 TANC E uc001gcq.1,uc001gcr.1 NUF E uc002xmj.3 ADA E uc003owu.1,uc003owv.1,uc003oww.1,uc003owx.2,uc 003owy.2,uc003owz.1,uc011dvp.1,uc011dvq.2,uc021y zw.1,uc021yzx.1 SLC29A E uc003prd.2,uc003pre.3 PRDM E uc003mll.1,uc003mlm.1 RNF E uc001znr.4,uc001zns.4,uc001znt.4,uc001znu.4,uc001z nv.4,uc001znw.4,uc010ucw.2 NUSAP E uc003vvh.3,uc003vvi.3,uc003vvj.3,uc010lne.3,uc011k qu.2,uc011kqv.2,uc011kqw.2,uc011kqx.1 TBXAS E uc003kor.1 AK E uc003iec.4 CCNA E uc001hce.3,uc009xbj.1 NUAK E uc003kdn.3 GCNT E uc003pyt.4,uc003pyu.2,uc003pyv.2 HSF E uc002jdp.1,uc002jdq.1,uc002jdr.1 CD79B E uc004cqm.3,uc004cqn.4,uc010nda.3 CD E uc002rmr.4,uc002rms.1,uc010ezl.3,uc010ymm.2 FAM179A E
34 uc002ldw.4 ACAA E uc002hez.2,uc010wbn.2 BLMH E uc001ayw.4,uc001ayz.1,uc001aza.5,uc001azb.1,uc001 azc.2,uc009vot.1,uc010oce.1 NBPF E uc002dod.4,uc002doe.2,uc010vcl.2 ZKSCAN E uc002xwf.1,uc002xwg.1,uc010gih.1 ATP9A E uc002ilv.1 TBX E uc002sfr.4,uc002sfs.4,uc010yqn.1,uc010yqo.2 ANXA E uc003wgn.4,uc011kur.2 ATP6V0E2- AS E uc003taz.3,uc003tba.3,uc003tbb.3,uc003tbc.3,uc022ab e.1,uc022abf.1 GGCT E uc002vvg.3,uc010znd.2,uc010zne.2 HJURP E uc002ypn.2 URB E uc003iwg.3,uc003iwh.3,uc003iwi.3 CASP E uc001xjf.3 ATP6V1D E uc001zgw.3,uc001zgy.1,uc010ubw.1,uc010ubx.1 ARHGAP11 A E uc003ukt.1,uc003uku.1,uc003ukv.1,uc003ukx.1,uc003 uky.2,uc003ukz.1,uc010les.1,uc011khl.1 CDK E uc001ade.3,uc001adf.3 TNFRSF E uc002kky.3,uc002kkz.3 YES E uc002efk.2,uc021tht.1 LOC E uc021yfx.1,uc021yfy.1,uc021yfz.1 ZNF E uc001bqa.2,uc001bqb.2,uc001bqc.2,uc001bqe.2,uc001 bqf.2,uc001bqg.2 RCC E uc002znb.3 DQ E uc002sdj.2,uc002sdk.4,uc002sdl.4,uc010yqb.1,uc010y qc.2,uc010yqd.1 CEP E uc002evt.2,uc002evu.2,uc002evv.2,uc010cfb.2,uc010c fc.1 SLC7A uc003kfy.4,uc010jab.4,uc010jac.4,uc010jad.4 HOMER uc003keo.3 F2RL uc002vsy.2 TIGD uc002tuw.4 MCM uc003xuw.3 GGH uc002wia.1,uc002wib.1 LZTS uc004dyf.2,uc004dyg.3,uc010nkw.3 KIF4A uc001tei.3,uc009ztf.3,uc009ztg.3 NTN uc001hpw.3,uc010pvl.2,uc021pjv.1 H3F3A uc002rly.3 LOC uc001jrv.3,uc001jrw.4,uc001jrx.4,uc001jry.3,uc001jrz. 3,uc001jsc.1,uc001jsg.4,uc001jsh.4,uc001jsi.4,uc001jsj CDH
35 .4,uc009xql.3,uc010qjr.2,uc021psl.1 uc001xap.3,uc001xar.3,uc010aoi.1,uc010aoj.2 CDKN uc003lcp.1,uc003lcq.1,uc003lcr.1,uc003lcs.1,uc003lct. 1,uc003lcu.1,uc010jet.1,uc011cyp.1,uc031slf.1 CDC25C uc001oma.4,uc009yry.4,uc031qbz.1 CDK2AP uc003yia.3,uc010mbg.3 LAPTM4B uc011bds.2 DOCK uc001dhg.4,uc001dhh.2,uc010orh.1 ST6GALNA C uc003jpl.1 GZMK uc002vry.4 NMUR uc002kyl.3,uc002kym.3,uc002kyn.1,uc002kyo.1 ZSCAN uc003geu.1,uc003gev.1,uc021xko.1,uc021xkp.1 MXD uc002aga.4 MYO1E uc001tuz.3,uc001tva.3,uc001tvb.3,uc001tvc.3,uc001tv d.1 SLC24A uc001zht.3,uc001zhu.3,uc001zhv.3,uc001zhw.3,uc001 zhx.3,uc001zhy.3,uc001zhz.3,uc001zia.3,uc001zib.3,u c001zic.3,uc001zid.3,uc010bau.3 SLC12A uc003zxj.3,uc010mkx.2,uc011lox.2,uc011loy.2,uc022b gm.1 CCDC uc003bmb.5,uc003bmc.5,uc003bmd.5,uc003bme.5,uc0 10hbd.4,uc011arz.1 TYMP uc002jgk.3,uc002jgl.3 KPNA uc002vpl.2,uc002vpm.2 CCL uc003xbv.3,uc003xbw.4,uc011kzk.1 SORBS uc003ehe.3,uc003ehf.1,uc003ehg.3,uc003ehh.1,uc003e hi.3,uc003ehj.2,uc003ehk.3,uc010hru.2,uc010hrv.1,uc 010hrw.2,uc011bjy.1,uc011bjz.2 KALRN uc001zvu.4,uc001zvv.4 SQRDL uc003pum.3 GTF3C uc003utt.3,uc003utu.3,uc022aii.1 GAL3ST uc003wir.2,uc003wis.2,uc022apy.1 CDK uc002tfc.3,uc002tfe.1,uc021vly.1 RGPD uc002qzx.3 YWHAQ uc002jfd.3,uc002jfe.3,uc002jfg.3,uc010dem.3,uc021ub w.1 RGS uc004ebr.1,uc010nlq.1 FTX uc010fhg.4 ANKRD36B P uc002ohb.2,uc002ohc.2 ZNF uc001nfh.2 PSMC uc004erq.3,uc004err.3,uc022cdk.1 NKRF uc001zmi.4,uc001zml.4,uc001zmm.1,uc001zmn.1,uc0 10bbw.3,uc010bbx.3 RAD
36 uc001ywk.3 NDN uc001lvu.3,uc021qcf.1 ASCL uc003xtz.1,uc003xua.1,uc003xub.3 CA uc003xue.3,uc003xuf.3,uc022aux.1 CHD uc003hub.3,uc011cdz.2 TSPAN uc001vft.1,uc010tgq.1 LINC uc002owp.3,uc010eiu.1 ETHE uc003fsv.3,uc003fsw.3 CCDC uc001gpa.2,uc001gpb.2,uc001gpc.2,uc001gpd.2,uc010 pnt.2,uc021pfv.1,uc021pfw.1 GLUL uc001dyt.2,uc001dyu.2,uc001dyw.4,uc021orj.2 CSF uc002lmd.3,uc021ulp.1 ZNF uc001xgh.4,uc001xgi.4,uc021ruf.1,uc031qox.1 WDR uc003nwr.3 CLIC uc003slw.3 SNX uc003kbk.3,uc010iyn.3,uc010jmw.1,uc011crv.1,uc011 crw.1,uc011crx.1 GUSBP uc001uey.1 RILPL uc001iig.2,uc001iih.2,uc001iii.2,uc009xic.2 ASB uc001bmp.4 HMGN uc003yov.3 DSCC uc003hpi.1,uc003hpj.3,uc003hpk.3,uc003hpl.3,uc003h pm.3,uc010ikf.3 ARHGAP uc003yad.3,uc022awb.1 LY uc011bgy.3 GPR uc003lwk.2,uc003lwl.3 HAVCR uc003soq.4,uc003sor.4,uc003sot.4,uc011jwi.1 ACTB uc001izb.4,uc001izc.4,uc001izd.2,uc009xmc.2,uc010q eu.2,uc031pue.1,uc031pug.1 ZNF uc001alm.1,uc001aln.3 AJAP uc001urx.3,uc001ury.3,uc001urz.3,uc010tdo.1 PAN uc002ffu.3 CMC uc009wka.2 AK uc001oif.2,uc001oig.2,uc001oih.2,uc010rpe.2 DPP uc001pkb.1,uc001pkc.1,uc001pkd.4,uc001pke.2,uc001 pkf.3,uc001pkg.1,uc009yxr.1,uc009yxs.1,uc009yxt.1 ATM uc021wps.1,uc021wpt.1,uc021wpu.1,uc021wpv.1 APOBEC3H uc001rln.4 ALG10B uc002wpo.3,uc002wpp.3,uc010zrn.2,uc010zro.2 BFSP uc001oku.1,uc001okv.1,uc001okw.1,uc001okx.1 PPP1CA uc003pyw.1 BC uc002xay.3,uc002xaz.1,uc002xba.1,uc010gex.3 GGT uc003nac.3,uc011diq.2 HIVEP uc001pkk.3,uc010rvy.2,uc010rvz.2 EXPH uc003zsv.2,uc003zsx.3,uc010mju.3 AQP
37 uc001bce.3,uc009vpk.3,uc010ocz.2,uc021ohr.1 CAPZB uc002hqr.3 PSMB uc004ckq.2 FUT uc002sdh.3,uc010ypz.2,uc010yqa.2 SLC1A uc002qom.3,uc002qon.3 ZNF uc010ydi.2,uc010ydj.2 ZNF uc001blu.3 SH3BGRL uc001hsn.4,uc001hso.3,uc001hsp.2,uc001hsq.2,uc001h sr.1,uc009xez.2,uc009xfa.3 OBSCN uc003idl.2,uc003idm.2 MAD2L uc001dej.4,uc001dek.4,uc001del.4,uc001dem.4 DEPDC uc001rgc.3,uc001rgd.4,uc001rge.4,uc010six.2,uc010si y.2 BCAT uc002jkj.2,uc002jkk.2,uc002jkl.2,uc002jkm.4 KIF uc002wzu.4 E2F uc002ill.1,uc002ilm.3,uc002iln.3,uc010daz.1 EFCAB uc002lht.3,uc010dpo.1,uc010xej.1,uc021uky.1 SEC11C uc003nas.2,uc003nat.2,uc003nau.3,uc003nav.3,uc021y lt.1,uc021ylu.1,uc031smu.1 GFOD uc001nsg.3,uc021qkj.1 FEN uc002gaq.3,uc021tog.1 ZFP uc001iou.2 VIM uc002qbb.2,uc002qbc.2,uc010eql.2 ZNF uc002vbz.2,uc002vca.2,uc010zix.2,uc031rqz.1,uc031rr a.1,uc031rrb.1 KLF uc001rmz.3 ZCRB uc002dlz.1 PLK uc001xkk.3,uc001xkl.3,uc001xkm.3,uc001xkn.3,uc001 xko.1,uc010ttb.2,uc010ttc.2,uc010ttd.1 ACTN uc002ily.3,uc002ilz.3,uc010wky.2 MRPL uc002flu.3 CDT uc003qil.2,uc003qim.2 ABRACL uc004duu.4,uc004duv.4,uc004duw.4,uc004dux.1,uc01 0nkj.2,uc022bxv.1 SPIN uc001tuw.3,uc001tux.3,uc010syt.1,uc010syu.2 TPCN uc002ist.1 AK uc001ont.3,uc009ysg.4 LRP uc002vds.3,uc002vdt.3,uc002vdv.3,uc002vdw.3,uc002 vdx.1,uc002vdy.1 KANSL1L uc001eyn.1,uc001eyo.3,uc001eyp.3,uc021oys.1,uc031 ppc.1 SNX uc001cdr.3,uc010oix.2 HPCAL uc003jhs.2,uc003jht.2 SUB uc004auw.4,uc010mrl.3 FBP uc003okl.3,uc003okm.3,uc003okn.3,uc010jvv.1,uc011 PPARD
38 dtb.2,uc011dtc.2 uc002cgs.2 MRPL uc004drv.3,uc004drw.3,uc004drx.1 TSPYL uc001rcu.2,uc001rcv.2,uc001rcw.2,uc001rcx.2,uc001r cy.2,uc001rcz.2,uc001rda.2 PTPRO uc002fxz.4,uc002fya.4,uc002fyb.4,uc010vsa.2 MYBBP1A uc021vlv.1 SH3RF3-AS uc001uyw.2,uc001uyx.2 EPSTI uc003njx.3 HIST1H1B uc001wtd.3,uc010tpt.2,uc010tpu.2 PSMA uc003hrv.4,uc003hrw.2,uc011cdn.2,uc011cdo.2 HERC uc001ikr.1 AF uc002snd.3 HK uc001pfw.1 MAML uc010zhw.2,uc010zhx.2 FAM117B uc004avh.3,uc004avi.4,uc010mrm.1,uc011lul.1,uc022 bkl.1 FANCC uc001kcc.4,uc001kcd.4,uc001kce.4,uc001kcf.4,uc021p ux.1,uc021puy.1 FAM213A uc002svz.1,uc010fhu.1,uc010fhv.1,uc010yum.1,uc010 yun.1 NCAPH uc002omq.4,uc002omr.4 PSMC uc004ayt.3,uc004ayu.3,uc004ayv.2,uc004ayx.2,uc004a yy.2 ANKS uc003dso.2,uc003dsp.3,uc003dsq.3,uc003dsr.3,uc003d ss.3,uc003dst.3,uc003dsu.3,uc003dsv.3 CLDND uc003lbr.5,uc003lbs.1,uc010jek.3,uc010jel.1,uc010jem.1,uc010jen.2,uc011cyc.2,uc011cyd.3,uc031sle.1 KLHL uc001cpl.2,uc009vye.2 RAD54L uc001ada.3,uc001adb.3,uc001adc.3,uc001add.3 TNFRSF uc001tgd.2,uc001tge.2,uc001tgf.2,uc001tgg.4,uc001tg h.4,uc001tgi.3,uc001tgj.3,uc001tgk.3,uc009ztp.3,uc00 9ztr.3,uc009zts.2,uc009ztt.1,uc010svd.2,uc010sve.2,uc 010svf.2,uc010svg.2 ANKS1B uc001bmc.3,uc009vsg.1 CD uc003pfx.1 OGFRL uc001pxx.3,uc010rzp.1,uc010rzq.1 SORL uc002zdr.4 CSTB uc002csj.4,uc002csk.4,uc002csl.4,uc010uwm.2 PAQR uc001uog.2 FGF uc002qpc.4,uc002qpd.4,uc002qpe.1,uc010eue.3,uc031 rnf.1 ZNF uc002xwx.3 PFDN uc001ein.4,uc009whp.3 FAM72B uc002vhd.4,uc002vhe.3,uc002vhf.3 ARPC
Supplementary Table S1. Primary antibodies for Western blot analysis. Antibody name and specifications
 Supplementary Table S1. Primary antibodies for Western blot analysis. Antibody name and specifications Manufacturer Actin (C-11) sc-1615, Santa Cruz, CA Akt1 (B-1) sc-5298 GAPDH (6C5) sc-32233 JAK2 (HR-758)
Supplementary Table S1. Primary antibodies for Western blot analysis. Antibody name and specifications Manufacturer Actin (C-11) sc-1615, Santa Cruz, CA Akt1 (B-1) sc-5298 GAPDH (6C5) sc-32233 JAK2 (HR-758)
TFDP2 TFDP1 SUMO1 SERTAD1 RPA3 RBBP8 RAD9A RAD17 RAD1 NBN MRE11A MNAT1 KPNA2 KNTC1 HUS1 HERC5 GTSE1 GTF2H1 GADD45A E2F4 DNM2 DDX11 CUL3 SKP2 RBL1 RB1
 -8-6 -4-2 CHEK2 MKI67 RB1 RBL1 SKP2 Fold-Change Fold-Change -3-2 -1 CCND1 CCNE1 CDK4 CDK5R1 5 1 15 ATM ATR BAX BCCIP BRCA1 BRCA2 BL1AT ABL1A ABL1 CCNG1 CCNG2 CCNH CCNT1 CCNT2 CDC2 CDC34 CDKN1A CDKN1B CDKN2A
-8-6 -4-2 CHEK2 MKI67 RB1 RBL1 SKP2 Fold-Change Fold-Change -3-2 -1 CCND1 CCNE1 CDK4 CDK5R1 5 1 15 ATM ATR BAX BCCIP BRCA1 BRCA2 BL1AT ABL1A ABL1 CCNG1 CCNG2 CCNH CCNT1 CCNT2 CDC2 CDC34 CDKN1A CDKN1B CDKN2A
Supplementary Appendix
 Supplementary Appendix Measuring crisis risk using conditional copulas: An empirical analysis of the 2008 shipping crisis Sebastian Opitz, Henry Seidel and Alexander Szimayer Model specification Table
Supplementary Appendix Measuring crisis risk using conditional copulas: An empirical analysis of the 2008 shipping crisis Sebastian Opitz, Henry Seidel and Alexander Szimayer Model specification Table
RARα2 expression confers myeloma stem cell features. Ye Yang, Jumei Shi, Giulia Tolomelli, Hongwei Xu, Jiliang Xia, He Wang, Wen Zhou, Yi Zhou,
 RARα2 expression confers myeloma stem cell features Ye Yang, Jumei Shi, Giulia Tolomelli, Hongwei Xu, Jiliang Xia, He Wang, Wen Zhou, Yi Zhou, Satyabrata Das, Zhimin Gu, Dana Levasseur, Fenghuang Zhan,
RARα2 expression confers myeloma stem cell features Ye Yang, Jumei Shi, Giulia Tolomelli, Hongwei Xu, Jiliang Xia, He Wang, Wen Zhou, Yi Zhou, Satyabrata Das, Zhimin Gu, Dana Levasseur, Fenghuang Zhan,
Supplemental Table S1. Tumor specific networks are enriched with somatically mutated genes (taken from the database COSMIC)
 Additional file 1 Supplemental Table S1. Tumor specific networks are enriched with somatically mutated genes (taken from the database COSMIC) COSMIC genes in tumor network COSMIC genes not in tumor network
Additional file 1 Supplemental Table S1. Tumor specific networks are enriched with somatically mutated genes (taken from the database COSMIC) COSMIC genes in tumor network COSMIC genes not in tumor network
Supplementary Figure 1. Therapeutic efficacy of combined NDV and anti-ctla-4 therapy is attenuated with a larger tumor challenge.
 Supplementary Figure 1. Therapeutic efficacy of combined NDV and anti-ctla-4 therapy is attenuated with a larger tumor challenge. Bilateral flank B16-F10 tumors were established as described and the animals
Supplementary Figure 1. Therapeutic efficacy of combined NDV and anti-ctla-4 therapy is attenuated with a larger tumor challenge. Bilateral flank B16-F10 tumors were established as described and the animals
ΓΕΩΠΟΝΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΑΘΗΝΩΝ ΤΜΗΜΑ ΑΓΡΟΤΙΚΗΣ ΟΙΚΟΝΟΜΙΑΣ & ΑΝΑΠΤΥΞΗΣ
 ΓΕΩΠΟΝΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΑΘΗΝΩΝ ΤΜΗΜΑ ΑΓΡΟΤΙΚΗΣ ΟΙΚΟΝΟΜΙΑΣ & ΑΝΑΠΤΥΞΗΣ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ «ΟΛΟΚΛΗΡΩΜΕΝΗ ΑΝΑΠΤΥΞΗ & ΔΙΑΧΕΙΡΙΣΗ ΤΟΥ ΑΓΡΟΤΙΚΟΥ ΧΩΡΟΥ» ΜΕΤΑΠΤΥΧΙΑΚΗ ΕΡΓΑΣΙΑ «Οικονομετρική διερεύνηση
ΓΕΩΠΟΝΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΑΘΗΝΩΝ ΤΜΗΜΑ ΑΓΡΟΤΙΚΗΣ ΟΙΚΟΝΟΜΙΑΣ & ΑΝΑΠΤΥΞΗΣ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ «ΟΛΟΚΛΗΡΩΜΕΝΗ ΑΝΑΠΤΥΞΗ & ΔΙΑΧΕΙΡΙΣΗ ΤΟΥ ΑΓΡΟΤΙΚΟΥ ΧΩΡΟΥ» ΜΕΤΑΠΤΥΧΙΑΚΗ ΕΡΓΑΣΙΑ «Οικονομετρική διερεύνηση
ΚΥΠΡΙΑΚΗ ΕΤΑΙΡΕΙΑ ΠΛΗΡΟΦΟΡΙΚΗΣ CYPRUS COMPUTER SOCIETY ΠΑΓΚΥΠΡΙΟΣ ΜΑΘΗΤΙΚΟΣ ΔΙΑΓΩΝΙΣΜΟΣ ΠΛΗΡΟΦΟΡΙΚΗΣ 19/5/2007
 Οδηγίες: Να απαντηθούν όλες οι ερωτήσεις. Αν κάπου κάνετε κάποιες υποθέσεις να αναφερθούν στη σχετική ερώτηση. Όλα τα αρχεία που αναφέρονται στα προβλήματα βρίσκονται στον ίδιο φάκελο με το εκτελέσιμο
Οδηγίες: Να απαντηθούν όλες οι ερωτήσεις. Αν κάπου κάνετε κάποιες υποθέσεις να αναφερθούν στη σχετική ερώτηση. Όλα τα αρχεία που αναφέρονται στα προβλήματα βρίσκονται στον ίδιο φάκελο με το εκτελέσιμο
Molecular evolutionary dynamics of respiratory syncytial virus group A in
 Molecular evolutionary dynamics of respiratory syncytial virus group A in recurrent epidemics in coastal Kenya James R. Otieno 1#, Charles N. Agoti 1, 2, Caroline W. Gitahi 1, Ann Bett 1, Mwanajuma Ngama
Molecular evolutionary dynamics of respiratory syncytial virus group A in recurrent epidemics in coastal Kenya James R. Otieno 1#, Charles N. Agoti 1, 2, Caroline W. Gitahi 1, Ann Bett 1, Mwanajuma Ngama
NES: normalized enrichment score (analyzed using KEGG pathway gene sets in the GSEA software); FDR:
 Supplementary table1. List of gene sets with simultaneously altered the enrichment upon SAP18 overexpression and knockdown. Gene sets enriched by SAP18 and reduced by shsap18 SAP18 against LacZ shsap18
Supplementary table1. List of gene sets with simultaneously altered the enrichment upon SAP18 overexpression and knockdown. Gene sets enriched by SAP18 and reduced by shsap18 SAP18 against LacZ shsap18
Cellular Physiology and Biochemistry
 Original Paper 2016 The Author(s). 2016 Published The Author(s) by S. Karger AG, Basel Published online: November 25, 2016 www.karger.com/cpb Published by S. Karger AG, Basel 486 www.karger.com/cpb Accepted:
Original Paper 2016 The Author(s). 2016 Published The Author(s) by S. Karger AG, Basel Published online: November 25, 2016 www.karger.com/cpb Published by S. Karger AG, Basel 486 www.karger.com/cpb Accepted:
Nature Medicine doi: /nm.2457
 ature Medicine doi:10.1038/nm.2457 ature Medicine doi:10.1038/nm.2457 ature Medicine doi:10.1038/nm.2457 ature Medicine doi:10.1038/nm.2457 ature Medicine doi:10.1038/nm.2457 Table S1 - Candidate genes
ature Medicine doi:10.1038/nm.2457 ature Medicine doi:10.1038/nm.2457 ature Medicine doi:10.1038/nm.2457 ature Medicine doi:10.1038/nm.2457 ature Medicine doi:10.1038/nm.2457 Table S1 - Candidate genes
Nature Immunology: doi: /ni Supplementary Figure 1. IL-6 reporter mice.
 Supplementary Figure 1 IL-6 reporter mice. (a) Targeted Il6 locus. The reporter cassette including a floxed stop cassette was introduced into exon 2 of the Il6 locus. Since the locus is disrupted by this
Supplementary Figure 1 IL-6 reporter mice. (a) Targeted Il6 locus. The reporter cassette including a floxed stop cassette was introduced into exon 2 of the Il6 locus. Since the locus is disrupted by this
< (0.999) Graft (0.698) (0.483) <0.001 (0.698) (<0.001) (<0.001) 3 months (0.999) (0.483) (<0.001) 6 months (<0.
 Supplementary table 1. Correlation of endothelial cell density among graft, 3, 6, and 12 months after Descemet s automated stripping endothelial keratoplasty. Graft 3 months 6 months 12 months Graft
Supplementary table 1. Correlation of endothelial cell density among graft, 3, 6, and 12 months after Descemet s automated stripping endothelial keratoplasty. Graft 3 months 6 months 12 months Graft
FORMULAS FOR STATISTICS 1
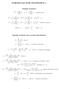 FORMULAS FOR STATISTICS 1 X = 1 n Sample statistics X i or x = 1 n x i (sample mean) S 2 = 1 n 1 s 2 = 1 n 1 (X i X) 2 = 1 n 1 (x i x) 2 = 1 n 1 Xi 2 n n 1 X 2 x 2 i n n 1 x 2 or (sample variance) E(X)
FORMULAS FOR STATISTICS 1 X = 1 n Sample statistics X i or x = 1 n x i (sample mean) S 2 = 1 n 1 s 2 = 1 n 1 (X i X) 2 = 1 n 1 (x i x) 2 = 1 n 1 Xi 2 n n 1 X 2 x 2 i n n 1 x 2 or (sample variance) E(X)
VBA Microsoft Excel. J. Comput. Chem. Jpn., Vol. 5, No. 1, pp (2006)
 J. Comput. Chem. Jpn., Vol. 5, No. 1, pp. 29 38 (2006) Microsoft Excel, 184-8588 2-24-16 e-mail: yosimura@cc.tuat.ac.jp (Received: July 28, 2005; Accepted for publication: October 24, 2005; Published on
J. Comput. Chem. Jpn., Vol. 5, No. 1, pp. 29 38 (2006) Microsoft Excel, 184-8588 2-24-16 e-mail: yosimura@cc.tuat.ac.jp (Received: July 28, 2005; Accepted for publication: October 24, 2005; Published on
± 20% ± 5% ± 10% RENCO ELECTRONICS, INC.
 RL15 RL16, RL17, RL18 MINIINDUCTORS CONFORMALLY COATED MARKING The nominal inductance is marked by a color code as listed in the table below. Color Black Brown Red Orange Yellow Green Blue Purple Grey
RL15 RL16, RL17, RL18 MINIINDUCTORS CONFORMALLY COATED MARKING The nominal inductance is marked by a color code as listed in the table below. Color Black Brown Red Orange Yellow Green Blue Purple Grey
Supplementary figures
 A Supplementary figures a) DMT.BG2 0.87 0.87 0.72 20 40 60 80 100 DMT.EG2 0.93 0.85 20 40 60 80 EMT.MG3 0.85 0 20 40 60 80 20 40 60 80 100 20 40 60 80 100 20 40 60 80 EMT.G6 DMT/EMT b) EG2 0.92 0.85 5
A Supplementary figures a) DMT.BG2 0.87 0.87 0.72 20 40 60 80 100 DMT.EG2 0.93 0.85 20 40 60 80 EMT.MG3 0.85 0 20 40 60 80 20 40 60 80 100 20 40 60 80 100 20 40 60 80 EMT.G6 DMT/EMT b) EG2 0.92 0.85 5
Biostatistics for Health Sciences Review Sheet
 Biostatistics for Health Sciences Review Sheet http://mathvault.ca June 1, 2017 Contents 1 Descriptive Statistics 2 1.1 Variables.............................................. 2 1.1.1 Qualitative........................................
Biostatistics for Health Sciences Review Sheet http://mathvault.ca June 1, 2017 Contents 1 Descriptive Statistics 2 1.1 Variables.............................................. 2 1.1.1 Qualitative........................................
encouraged to use the Version of Record that, when published, will replace this version. The most /BCJ
 Biochemical Journal: this is an Accepted Manuscript, not the final Version of Record. You are encouraged to use the Version of Record that, when published, will replace this version. The most up-to-date
Biochemical Journal: this is an Accepted Manuscript, not the final Version of Record. You are encouraged to use the Version of Record that, when published, will replace this version. The most up-to-date
Si + Al Mg Fe + Mn +Ni Ca rim Ca p.f.u
 .6.5. y = -.4x +.8 R =.9574 y = - x +.14 R =.9788 y = -.4 x +.7 R =.9896 Si + Al Fe + Mn +Ni y =.55 x.36 R =.9988.149.148.147.146.145..88 core rim.144 4 =.6 ±.6 4 =.6 ±.18.84.88 p.f.u..86.76 y = -3.9 x
.6.5. y = -.4x +.8 R =.9574 y = - x +.14 R =.9788 y = -.4 x +.7 R =.9896 Si + Al Fe + Mn +Ni y =.55 x.36 R =.9988.149.148.147.146.145..88 core rim.144 4 =.6 ±.6 4 =.6 ±.18.84.88 p.f.u..86.76 y = -3.9 x
Does anemia contribute to end-organ dysfunction in ICU patients Statistical Analysis
 Does anemia contribute to end-organ dysfunction in ICU patients Statistical Analysis Xue Han, MPH and Matt Shotwell, PhD Department of Biostatistics Vanderbilt University School of Medicine March 14, 2014
Does anemia contribute to end-organ dysfunction in ICU patients Statistical Analysis Xue Han, MPH and Matt Shotwell, PhD Department of Biostatistics Vanderbilt University School of Medicine March 14, 2014
Οι επιδόσεις Ελλήνων στο Mini Mental State Examination με βάση την ηλικία και τη νοητική κατάσταση από την παιδική στην τρίτη ηλικία.
 CLINICAL TRIALS ΚΛΙΝΙΚΕΣ ΜΕΛΕΤΕΣ 25 Οι επιδόσεις Ελλήνων στο Mini Mental State Examination με βάση την ηλικία και τη νοητική κατάσταση από την παιδική στην τρίτη ηλικία. Τσάνταλη, Ε. 1,2,3, Οικονομίδης,
CLINICAL TRIALS ΚΛΙΝΙΚΕΣ ΜΕΛΕΤΕΣ 25 Οι επιδόσεις Ελλήνων στο Mini Mental State Examination με βάση την ηλικία και τη νοητική κατάσταση από την παιδική στην τρίτη ηλικία. Τσάνταλη, Ε. 1,2,3, Οικονομίδης,
5.4 The Poisson Distribution.
 The worst thing you can do about a situation is nothing. Sr. O Shea Jackson 5.4 The Poisson Distribution. Description of the Poisson Distribution Discrete probability distribution. The random variable
The worst thing you can do about a situation is nothing. Sr. O Shea Jackson 5.4 The Poisson Distribution. Description of the Poisson Distribution Discrete probability distribution. The random variable
Aquinas College. Edexcel Mathematical formulae and statistics tables DO NOT WRITE ON THIS BOOKLET
 Aquinas College Edexcel Mathematical formulae and statistics tables DO NOT WRITE ON THIS BOOKLET Pearson Edexcel Level 3 Advanced Subsidiary and Advanced GCE in Mathematics and Further Mathematics Mathematical
Aquinas College Edexcel Mathematical formulae and statistics tables DO NOT WRITE ON THIS BOOKLET Pearson Edexcel Level 3 Advanced Subsidiary and Advanced GCE in Mathematics and Further Mathematics Mathematical
 22 .5 Real consumption.5 Real residential investment.5.5.5 965 975 985 995 25.5 965 975 985 995 25.5 Real house prices.5 Real fixed investment.5.5.5 965 975 985 995 25.5 965 975 985 995 25.3 Inflation
22 .5 Real consumption.5 Real residential investment.5.5.5 965 975 985 995 25.5 965 975 985 995 25.5 Real house prices.5 Real fixed investment.5.5.5 965 975 985 995 25.5 965 975 985 995 25.3 Inflation
Τα ηπατικά επίπεδα του FOXP3 mrna στη χρόνια ηπατίτιδα Β εξαρτώνται από την έκφραση των οδών Fas/FasL και PD-1/PD-L1
 Τα ηπατικά επίπεδα του FOXP3 mrna στη χρόνια ηπατίτιδα Β εξαρτώνται από την έκφραση των οδών Fas/FasL και PD-1/PD-L1 Γ. Γερμανίδης, Ν. Αργέντου, Θ. Βασιλειάδης, Κ. Πατσιαούρα, Κ. Μαντζούκης, A. Θεοχαρίδου,
Τα ηπατικά επίπεδα του FOXP3 mrna στη χρόνια ηπατίτιδα Β εξαρτώνται από την έκφραση των οδών Fas/FasL και PD-1/PD-L1 Γ. Γερμανίδης, Ν. Αργέντου, Θ. Βασιλειάδης, Κ. Πατσιαούρα, Κ. Μαντζούκης, A. Θεοχαρίδου,
Μαντζούνη, Πιπερίγκου, Χατζή. ΒΙΟΣΤΑΤΙΣΤΙΚΗ Εργαστήριο 5 ο
 Κατανομές Στατιστικών Συναρτήσεων Δύο δείγματα από κανονική κατανομή Έστω Χ= ( Χ, Χ,..., Χ ) τ.δ. από Ν( µ, σ ) μεγέθους n και 1 n 1 1 Y = (Y, Y,...,Y ) τ.δ. από Ν( µ, σ ) 1 n 1 Χ Y ( µ µ ) S σ Τ ( Χ,Y)
Κατανομές Στατιστικών Συναρτήσεων Δύο δείγματα από κανονική κατανομή Έστω Χ= ( Χ, Χ,..., Χ ) τ.δ. από Ν( µ, σ ) μεγέθους n και 1 n 1 1 Y = (Y, Y,...,Y ) τ.δ. από Ν( µ, σ ) 1 n 1 Χ Y ( µ µ ) S σ Τ ( Χ,Y)
A strategy for the identification of combinatorial bioactive compounds. contributing to the holistic effect of herbal medicines
 1 2 Supplementary information 3 4 A strategy for the identification of combinatorial bioactive compounds contributing to the holistic effect of herbal medicines 5 6 Fang Long 1, Hua Yang 1, Yanmin Xu,
1 2 Supplementary information 3 4 A strategy for the identification of combinatorial bioactive compounds contributing to the holistic effect of herbal medicines 5 6 Fang Long 1, Hua Yang 1, Yanmin Xu,
Math 6 SL Probability Distributions Practice Test Mark Scheme
 Math 6 SL Probability Distributions Practice Test Mark Scheme. (a) Note: Award A for vertical line to right of mean, A for shading to right of their vertical line. AA N (b) evidence of recognizing symmetry
Math 6 SL Probability Distributions Practice Test Mark Scheme. (a) Note: Award A for vertical line to right of mean, A for shading to right of their vertical line. AA N (b) evidence of recognizing symmetry
Supplementary Information 1.
 Supplementary Information 1. Fig. S1. Correlations between litter-derived-c and N (percent of initial input) and Al-/Fe- (hydr)oxides dissolved by ammonium oxalate (AO); a) 0 10 cm; b) 10 20 cm; c) 20
Supplementary Information 1. Fig. S1. Correlations between litter-derived-c and N (percent of initial input) and Al-/Fe- (hydr)oxides dissolved by ammonium oxalate (AO); a) 0 10 cm; b) 10 20 cm; c) 20
Supplementary Fig. 1. Nature Medicine: doi: /nm z Donor # BB-z. 28- IL2RB-z (YXXQ) 28-IL2RBz (YXXQ) Control MFI: 1866 MFI:
 Supplementary Fig. 1 a Donor # 28-IL2RB-z 28- IL2RB-z Control MFI: 1866 1394 612 1297 11 MFI: 1794 1294 566 1225 11 MFI: 1433 1157 327 1117 98.5 MFI: 1463 1224 352 111 126 MFI: 1441 1232 497 1268 151 MFI:
Supplementary Fig. 1 a Donor # 28-IL2RB-z 28- IL2RB-z Control MFI: 1866 1394 612 1297 11 MFI: 1794 1294 566 1225 11 MFI: 1433 1157 327 1117 98.5 MFI: 1463 1224 352 111 126 MFI: 1441 1232 497 1268 151 MFI:
Supporting Information
 Electronic Supplementary Material (ESI) for Organic & Biomolecular Chemistry. This journal is The Royal Society of Chemistry 2018 Supporting Information Crotonols A and B, Two Rare Tigliane Diterpenoid
Electronic Supplementary Material (ESI) for Organic & Biomolecular Chemistry. This journal is The Royal Society of Chemistry 2018 Supporting Information Crotonols A and B, Two Rare Tigliane Diterpenoid
PHOS π 0 analysis, for production, R AA, and Flow analysis, LHC11h
 PHOS π, ask PHOS π analysis, for production, R AA, and Flow analysis, Henrik Qvigstad henrik.qvigstad@fys.uio.no University of Oslo --5 PHOS π, ask ask he task we use, AliaskPiFlow was written prior, for
PHOS π, ask PHOS π analysis, for production, R AA, and Flow analysis, Henrik Qvigstad henrik.qvigstad@fys.uio.no University of Oslo --5 PHOS π, ask ask he task we use, AliaskPiFlow was written prior, for
The effect of curcumin on the stability of Aβ. dimers
 The effect of curcumin on the stability of Aβ dimers Li Na Zhao, See-Wing Chiu, Jérôme Benoit, Lock Yue Chew,, and Yuguang Mu, School of Physical and Mathematical Sciences, Nanyang Technological University,
The effect of curcumin on the stability of Aβ dimers Li Na Zhao, See-Wing Chiu, Jérôme Benoit, Lock Yue Chew,, and Yuguang Mu, School of Physical and Mathematical Sciences, Nanyang Technological University,
ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΥΠΡΟΥ ΣΧΟΛΗ ΓΕΩΠΟΝΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΒΙΟΤΕΧΝΟΛΟΓΙΑΣ ΚΑΙ ΕΠΙΣΤΗΜΗΣ ΤΡΟΦΙΜΩΝ. Πτυχιακή εργασία
 ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΥΠΡΟΥ ΣΧΟΛΗ ΓΕΩΠΟΝΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΒΙΟΤΕΧΝΟΛΟΓΙΑΣ ΚΑΙ ΕΠΙΣΤΗΜΗΣ ΤΡΟΦΙΜΩΝ Πτυχιακή εργασία ΜΕΛΕΤΗ ΠΟΛΥΦΑΙΝΟΛΩΝ ΚΑΙ ΑΝΤΙΟΞΕΙΔΩΤΙΚΗΣ ΙΚΑΝΟΤΗΤΑΣ ΣΟΚΟΛΑΤΑΣ Αναστασία Σιάντωνα Λεμεσός
ΤΕΧΝΟΛΟΓΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΥΠΡΟΥ ΣΧΟΛΗ ΓΕΩΠΟΝΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΒΙΟΤΕΧΝΟΛΟΓΙΑΣ ΚΑΙ ΕΠΙΣΤΗΜΗΣ ΤΡΟΦΙΜΩΝ Πτυχιακή εργασία ΜΕΛΕΤΗ ΠΟΛΥΦΑΙΝΟΛΩΝ ΚΑΙ ΑΝΤΙΟΞΕΙΔΩΤΙΚΗΣ ΙΚΑΝΟΤΗΤΑΣ ΣΟΚΟΛΑΤΑΣ Αναστασία Σιάντωνα Λεμεσός
Electronic Supplementary Information (ESI)
 Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2016 Electronic Supplementary Information (ESI) Cyclopentadienyl iron dicarbonyl (CpFe(CO) 2 ) derivatives
Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2016 Electronic Supplementary Information (ESI) Cyclopentadienyl iron dicarbonyl (CpFe(CO) 2 ) derivatives
CHAPTER 25 SOLVING EQUATIONS BY ITERATIVE METHODS
 CHAPTER 5 SOLVING EQUATIONS BY ITERATIVE METHODS EXERCISE 104 Page 8 1. Find the positive root of the equation x + 3x 5 = 0, correct to 3 significant figures, using the method of bisection. Let f(x) =
CHAPTER 5 SOLVING EQUATIONS BY ITERATIVE METHODS EXERCISE 104 Page 8 1. Find the positive root of the equation x + 3x 5 = 0, correct to 3 significant figures, using the method of bisection. Let f(x) =
519.22(07.07) 78 : ( ) /.. ; c (07.07) , , 2008
 .. ( ) 2008 519.22(07.07) 78 : ( ) /.. ;. : -, 2008. 38 c. ( ) STATISTICA.,. STATISTICA.,. 519.22(07.07),.., 2008.., 2008., 2008 2 ... 4 1...5...5 2...14...14 3...27...27 3 ,, -. " ", :,,,... STATISTICA.,,,.
.. ( ) 2008 519.22(07.07) 78 : ( ) /.. ;. : -, 2008. 38 c. ( ) STATISTICA.,. STATISTICA.,. 519.22(07.07),.., 2008.., 2008., 2008 2 ... 4 1...5...5 2...14...14 3...27...27 3 ,, -. " ", :,,,... STATISTICA.,,,.
Group 2 Methotrexate 7.5 mg/week, increased to 15 mg/week after 4 weeks. Methotrexate 7.5 mg/week, increased to 15 mg/week after 4 weeks
 Group 1 Methotrexate 7.5 mg/week, increased to 15 mg/week after 4 weeks Group 2 Methotrexate 7.5 mg/week, increased to 15 mg/week after 4 weeks Sulfasalazine 2000-3000 mg/day Leflunomide 20 mg/day Infliximab
Group 1 Methotrexate 7.5 mg/week, increased to 15 mg/week after 4 weeks Group 2 Methotrexate 7.5 mg/week, increased to 15 mg/week after 4 weeks Sulfasalazine 2000-3000 mg/day Leflunomide 20 mg/day Infliximab
Potential Dividers. 46 minutes. 46 marks. Page 1 of 11
 Potential Dividers 46 minutes 46 marks Page 1 of 11 Q1. In the circuit shown in the figure below, the battery, of negligible internal resistance, has an emf of 30 V. The pd across the lamp is 6.0 V and
Potential Dividers 46 minutes 46 marks Page 1 of 11 Q1. In the circuit shown in the figure below, the battery, of negligible internal resistance, has an emf of 30 V. The pd across the lamp is 6.0 V and
HOMEWORK 4 = G. In order to plot the stress versus the stretch we define a normalized stretch:
 HOMEWORK 4 Problem a For the fast loading case, we want to derive the relationship between P zz and λ z. We know that the nominal stress is expressed as: P zz = ψ λ z where λ z = λ λ z. Therefore, applying
HOMEWORK 4 Problem a For the fast loading case, we want to derive the relationship between P zz and λ z. We know that the nominal stress is expressed as: P zz = ψ λ z where λ z = λ λ z. Therefore, applying
Single Stock Analysis Stock Pair Analysis Portfolio Dates Portfolio Dates Correl Maturity VolRatio Ref Stock Correl Ref Stock
 Single Stock Analysis Stock Pair Analysis Portfolio Dates Portfolio Dates orrel Maturity Date Monday, December 21, 2009 Date Monday, December 21, 2009 30 Percentile Date Friday, December 19, 2008 Percentile
Single Stock Analysis Stock Pair Analysis Portfolio Dates Portfolio Dates orrel Maturity Date Monday, December 21, 2009 Date Monday, December 21, 2009 30 Percentile Date Friday, December 19, 2008 Percentile
Figure 3 Three observations (Vp, Vs and density isosurfaces) intersecting in the PLF space. Solutions exist at the two indicated points.
 φ φ φ φ Figure 1 Resampling of a rock-physics model from velocity-porosity to lithology-porosity space. C i are model results for various clay contents. φ ρ ρ δ Figure 2 Bulk modulus constraint cube in
φ φ φ φ Figure 1 Resampling of a rock-physics model from velocity-porosity to lithology-porosity space. C i are model results for various clay contents. φ ρ ρ δ Figure 2 Bulk modulus constraint cube in
Description of the PX-HC algorithm
 A Description of the PX-HC algorithm Let N = C c= N c and write C Nc K c= i= k= as, the Gibbs sampling algorithm at iteration m for continuous outcomes: Step A: For =,, J, draw θ m in the following steps:
A Description of the PX-HC algorithm Let N = C c= N c and write C Nc K c= i= k= as, the Gibbs sampling algorithm at iteration m for continuous outcomes: Step A: For =,, J, draw θ m in the following steps:
ΜΟΡΙΑΚΕΣ ΜΕΘΟΔΟΙ ΚΡΙΤΗΡΙΑ ΕΠΙΛΟΓΗΣ ΑΞΙΟΛΟΓΗΣΗ
 - - ό ό : Ω ί ό ί ώ.. ά ή ί ός ή ς ύ ί ί ή έ ώ ό. ή ς ά ς ής ό ός ά ς ή έ ός ή ς ί ής ί ς. ό έ ώ - ώ ή ής ή ς- ί ά ά ς ί ς. ά ί ί έ έ ά ύ ή, ό ί ά, ό ό ά έ ά ής ί ύ George Wald ή ί έ ς ί ύ ό ς ί ς ά έ
- - ό ό : Ω ί ό ί ώ.. ά ή ί ός ή ς ύ ί ί ή έ ώ ό. ή ς ά ς ής ό ός ά ς ή έ ός ή ς ί ής ί ς. ό έ ώ - ώ ή ής ή ς- ί ά ά ς ί ς. ά ί ί έ έ ά ύ ή, ό ί ά, ό ό ά έ ά ής ί ύ George Wald ή ί έ ς ί ύ ό ς ί ς ά έ
Supporting Information
 Chloromethylhalicyclamine B, a Marine-Derived Protein Kinase CK1δ/ε Inhibitor Germana Esposito, Marie-Lise Bourguet-Kondracki, * Linh H. Mai, Arlette Longeon, Roberta Teta, Laurent Meijer, Rob Van Soest,
Chloromethylhalicyclamine B, a Marine-Derived Protein Kinase CK1δ/ε Inhibitor Germana Esposito, Marie-Lise Bourguet-Kondracki, * Linh H. Mai, Arlette Longeon, Roberta Teta, Laurent Meijer, Rob Van Soest,
Statistics 104: Quantitative Methods for Economics Formula and Theorem Review
 Harvard College Statistics 104: Quantitative Methods for Economics Formula and Theorem Review Tommy MacWilliam, 13 tmacwilliam@college.harvard.edu March 10, 2011 Contents 1 Introduction to Data 5 1.1 Sample
Harvard College Statistics 104: Quantitative Methods for Economics Formula and Theorem Review Tommy MacWilliam, 13 tmacwilliam@college.harvard.edu March 10, 2011 Contents 1 Introduction to Data 5 1.1 Sample
10-π-electron arenes à la carte: Structure. Sr, Ba; n = 6-8) complexes
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2016 Supporting information 10-π-electron arenes à la carte: Structure and Bonding of
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2016 Supporting information 10-π-electron arenes à la carte: Structure and Bonding of
An experimental and theoretical study of the gas phase kinetics of atomic chlorine reactions with CH 3 NH 2, (CH 3 ) 2 NH, and (CH 3 ) 3 N
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2015 An experimental and theoretical study of the gas phase kinetics of atomic chlorine
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2015 An experimental and theoretical study of the gas phase kinetics of atomic chlorine
Christopher Stephen Inchley, 2015
 Christopher Stephen Inchley, 2015 Series of dissertations submitted to the Faculty of Medicine, University of Oslo No. 2009 ISBN 978-82-8264-990-2 All rights reserved. No part of this publication may be
Christopher Stephen Inchley, 2015 Series of dissertations submitted to the Faculty of Medicine, University of Oslo No. 2009 ISBN 978-82-8264-990-2 All rights reserved. No part of this publication may be
Electronic Supplementary Information DFT Characterization on the Mechanism of Water Splitting Catalyzed by Single-Ru-substituted Polyoxometalates
 Electronic Supplementary Information DFT Characterization on the Mechanism of Water Splitting Catalyzed by Single-Ru-substituted Polyoxometalates Zhong-Ling Lang, Guo-Chun Yang, Na-Na Ma, Shi-Zheng Wen,
Electronic Supplementary Information DFT Characterization on the Mechanism of Water Splitting Catalyzed by Single-Ru-substituted Polyoxometalates Zhong-Ling Lang, Guo-Chun Yang, Na-Na Ma, Shi-Zheng Wen,
Statistics & Research methods. Athanasios Papaioannou University of Thessaly Dept. of PE & Sport Science
 Statistics & Research methods Athanasios Papaioannou University of Thessaly Dept. of PE & Sport Science 30 25 1,65 20 1,66 15 10 5 1,67 1,68 Κανονική 0 Height 1,69 Καμπύλη Κανονική Διακύμανση & Ζ-scores
Statistics & Research methods Athanasios Papaioannou University of Thessaly Dept. of PE & Sport Science 30 25 1,65 20 1,66 15 10 5 1,67 1,68 Κανονική 0 Height 1,69 Καμπύλη Κανονική Διακύμανση & Ζ-scores
Μενύχτα, Πιπερίγκου, Σαββάτης. ΒΙΟΣΤΑΤΙΣΤΙΚΗ Εργαστήριο 5 ο
 Κατανομές Στατιστικών Συναρτήσεων Δύο ανεξάρτητα δείγματα από κανονική κατανομή Έστω Χ= ( Χ, Χ,..., Χ ) τ.δ. από Ν( µ, σ ) μεγέθους n και 1 n 1 1 Y = (Y, Y,..., Y ) τ.δ. από Ν( µ, σ ) 1 n 1 Χ Y ( µ µ )
Κατανομές Στατιστικών Συναρτήσεων Δύο ανεξάρτητα δείγματα από κανονική κατανομή Έστω Χ= ( Χ, Χ,..., Χ ) τ.δ. από Ν( µ, σ ) μεγέθους n και 1 n 1 1 Y = (Y, Y,..., Y ) τ.δ. από Ν( µ, σ ) 1 n 1 Χ Y ( µ µ )
DuPont Suva 95 Refrigerant
 Technical Information T-95 ENG DuPont Suva refrigerants Thermodynamic Properties of DuPont Suva 95 Refrigerant (R-508B) The DuPont Oval Logo, The miracles of science, and Suva, are trademarks or registered
Technical Information T-95 ENG DuPont Suva refrigerants Thermodynamic Properties of DuPont Suva 95 Refrigerant (R-508B) The DuPont Oval Logo, The miracles of science, and Suva, are trademarks or registered
Quantitative chemical analyses of rocks with X-ray fluorescence analyzer: major and trace elements in ultrabasic rocks
 98 Scientific Note X : Quantitative chemical analyses of rocks with X-ray fluorescence analyzer: major and trace elements in ultrabasic rocks Kimiko Seno and Yoichi Motoyoshi,**- +, +, ;,**. -,/ Abstract:
98 Scientific Note X : Quantitative chemical analyses of rocks with X-ray fluorescence analyzer: major and trace elements in ultrabasic rocks Kimiko Seno and Yoichi Motoyoshi,**- +, +, ;,**. -,/ Abstract:
2 Composition. Invertible Mappings
 Arkansas Tech University MATH 4033: Elementary Modern Algebra Dr. Marcel B. Finan Composition. Invertible Mappings In this section we discuss two procedures for creating new mappings from old ones, namely,
Arkansas Tech University MATH 4033: Elementary Modern Algebra Dr. Marcel B. Finan Composition. Invertible Mappings In this section we discuss two procedures for creating new mappings from old ones, namely,
3.4 SUM AND DIFFERENCE FORMULAS. NOTE: cos(α+β) cos α + cos β cos(α-β) cos α -cos β
 3.4 SUM AND DIFFERENCE FORMULAS Page Theorem cos(αβ cos α cos β -sin α cos(α-β cos α cos β sin α NOTE: cos(αβ cos α cos β cos(α-β cos α -cos β Proof of cos(α-β cos α cos β sin α Let s use a unit circle
3.4 SUM AND DIFFERENCE FORMULAS Page Theorem cos(αβ cos α cos β -sin α cos(α-β cos α cos β sin α NOTE: cos(αβ cos α cos β cos(α-β cos α -cos β Proof of cos(α-β cos α cos β sin α Let s use a unit circle
Technical Information T-9100 SI. Suva. refrigerants. Thermodynamic Properties of. Suva Refrigerant [R-410A (50/50)]
![Technical Information T-9100 SI. Suva. refrigerants. Thermodynamic Properties of. Suva Refrigerant [R-410A (50/50)] Technical Information T-9100 SI. Suva. refrigerants. Thermodynamic Properties of. Suva Refrigerant [R-410A (50/50)]](/thumbs/89/98928029.jpg) d Suva refrigerants Technical Information T-9100SI Thermodynamic Properties of Suva 9100 Refrigerant [R-410A (50/50)] Thermodynamic Properties of Suva 9100 Refrigerant SI Units New tables of the thermodynamic
d Suva refrigerants Technical Information T-9100SI Thermodynamic Properties of Suva 9100 Refrigerant [R-410A (50/50)] Thermodynamic Properties of Suva 9100 Refrigerant SI Units New tables of the thermodynamic
ΚΕΡΚΙΔΙΚΗ ΚΑΙ ΜΗΡΙΑΙΑ ΠΡΟΣΠΕΛΑΣΗ ΣΕ ΣΤΕΦΑΝΙΟΓΡΑΦΙΕΣ ΚΑΙ ΑΓΓΕΙΟΠΛΑΣΤΙΚΕΣ. ΜΙΑ ΣΥΓΚΡΙΤΙΚΗ ΜΕΛΕΤΗ
 ΚΕΡΚΙΔΙΚΗ ΚΑΙ ΜΗΡΙΑΙΑ ΠΡΟΣΠΕΛΑΣΗ ΣΕ ΣΤΕΦΑΝΙΟΓΡΑΦΙΕΣ ΚΑΙ ΑΓΓΕΙΟΠΛΑΣΤΙΚΕΣ. ΜΙΑ ΣΥΓΚΡΙΤΙΚΗ ΜΕΛΕΤΗ Μαυρόγιαννη Γεωργία 1, Καραντζούλα Ευαγγελία 2, Τουλιά Γεωργία 3, Κυριακόπουλος Βασίλης 4, Καδδά Όλγα 5 EΡΕΥΝΑ
ΚΕΡΚΙΔΙΚΗ ΚΑΙ ΜΗΡΙΑΙΑ ΠΡΟΣΠΕΛΑΣΗ ΣΕ ΣΤΕΦΑΝΙΟΓΡΑΦΙΕΣ ΚΑΙ ΑΓΓΕΙΟΠΛΑΣΤΙΚΕΣ. ΜΙΑ ΣΥΓΚΡΙΤΙΚΗ ΜΕΛΕΤΗ Μαυρόγιαννη Γεωργία 1, Καραντζούλα Ευαγγελία 2, Τουλιά Γεωργία 3, Κυριακόπουλος Βασίλης 4, Καδδά Όλγα 5 EΡΕΥΝΑ
Παιδιατρική ΒΟΡΕΙΟΥ ΕΛΛΑΔΟΣ, 23, 3. Γ Παιδιατρική Κλινική, Αριστοτέλειο Πανεπιστήμιο, Ιπποκράτειο Νοσοκομείο Θεσσαλονίκης, 2
 162 Σύγκριση της επίδρασης της Μοντελουκάστης και των Εισπνεομένων Στεροειδών στο Εκπνεόμενο Μονοξείδιο του Αζώτου (eno) και την αναπνευστική λειτουργία σε ασθματικά παιδιά Ε. Χατζηαγόρου 1, Β. Αβραμίδου
162 Σύγκριση της επίδρασης της Μοντελουκάστης και των Εισπνεομένων Στεροειδών στο Εκπνεόμενο Μονοξείδιο του Αζώτου (eno) και την αναπνευστική λειτουργία σε ασθματικά παιδιά Ε. Χατζηαγόρου 1, Β. Αβραμίδου
ΚΥΠΡΙΑΚΟΣ ΣΥΝΔΕΣΜΟΣ ΠΛΗΡΟΦΟΡΙΚΗΣ CYPRUS COMPUTER SOCIETY 21 ος ΠΑΓΚΥΠΡΙΟΣ ΜΑΘΗΤΙΚΟΣ ΔΙΑΓΩΝΙΣΜΟΣ ΠΛΗΡΟΦΟΡΙΚΗΣ Δεύτερος Γύρος - 30 Μαρτίου 2011
 Διάρκεια Διαγωνισμού: 3 ώρες Απαντήστε όλες τις ερωτήσεις Μέγιστο Βάρος (20 Μονάδες) Δίνεται ένα σύνολο από N σφαιρίδια τα οποία δεν έχουν όλα το ίδιο βάρος μεταξύ τους και ένα κουτί που αντέχει μέχρι
Διάρκεια Διαγωνισμού: 3 ώρες Απαντήστε όλες τις ερωτήσεις Μέγιστο Βάρος (20 Μονάδες) Δίνεται ένα σύνολο από N σφαιρίδια τα οποία δεν έχουν όλα το ίδιο βάρος μεταξύ τους και ένα κουτί που αντέχει μέχρι
Supporting Information
 Supporting Information rigin of the Regio- and Stereoselectivity of Allylic Substitution of rganocopper Reagents Naohiko Yoshikai, Song-Lin Zhang, and Eiichi Nakamura* Department of Chemistry, The University
Supporting Information rigin of the Regio- and Stereoselectivity of Allylic Substitution of rganocopper Reagents Naohiko Yoshikai, Song-Lin Zhang, and Eiichi Nakamura* Department of Chemistry, The University
APPENDICES APPENDIX A. STATISTICAL TABLES AND CHARTS 651 APPENDIX B. BIBLIOGRAPHY 677 APPENDIX C. ANSWERS TO SELECTED EXERCISES 679
 APPENDICES APPENDIX A. STATISTICAL TABLES AND CHARTS 1 Table I Summary of Common Probability Distributions 2 Table II Cumulative Standard Normal Distribution Table III Percentage Points, 2 of the Chi-Squared
APPENDICES APPENDIX A. STATISTICAL TABLES AND CHARTS 1 Table I Summary of Common Probability Distributions 2 Table II Cumulative Standard Normal Distribution Table III Percentage Points, 2 of the Chi-Squared
department listing department name αχχουντσ ϕανε βαλικτ δδσϕηασδδη σδηφγ ασκϕηλκ τεχηνιχαλ αλαν ϕουν διξ τεχηνιχαλ ϕοην µαριανι
 She selects the option. Jenny starts with the al listing. This has employees listed within She drills down through the employee. The inferred ER sttricture relates this to the redcords in the databasee
She selects the option. Jenny starts with the al listing. This has employees listed within She drills down through the employee. The inferred ER sttricture relates this to the redcords in the databasee
Structure-Metabolism-Relationships in the microsomal clearance of. piperazin-1-ylpyridazines
 Electronic Supplementary Material (ESI) for MedChemComm. This journal is The Royal Society of Chemistry 2017 Structure-Metabolism-Relationships in the microsomal clearance of piperazin-1-ylpyridazines
Electronic Supplementary Material (ESI) for MedChemComm. This journal is The Royal Society of Chemistry 2017 Structure-Metabolism-Relationships in the microsomal clearance of piperazin-1-ylpyridazines
Heterobimetallic Pd-Sn Catalysis: Michael Addition. Reaction with C-, N-, O-, S- Nucleophiles and In-situ. Diagnostics
 Supporting Information (SI) Heterobimetallic Pd-Sn Catalysis: Michael Addition Reaction with C-, N-, -, S- Nucleophiles and In-situ Diagnostics Debjit Das, a Sanjay Pratihar a,b and Sujit Roy c * a rganometallics
Supporting Information (SI) Heterobimetallic Pd-Sn Catalysis: Michael Addition Reaction with C-, N-, -, S- Nucleophiles and In-situ Diagnostics Debjit Das, a Sanjay Pratihar a,b and Sujit Roy c * a rganometallics
Chapter 1 Introduction to Observational Studies Part 2 Cross-Sectional Selection Bias Adjustment
 Contents Preface ix Part 1 Introduction Chapter 1 Introduction to Observational Studies... 3 1.1 Observational vs. Experimental Studies... 3 1.2 Issues in Observational Studies... 5 1.3 Study Design...
Contents Preface ix Part 1 Introduction Chapter 1 Introduction to Observational Studies... 3 1.1 Observational vs. Experimental Studies... 3 1.2 Issues in Observational Studies... 5 1.3 Study Design...
Supplementary Materials for. Kinetic and Computational Studies on Pd(I) Dimer- Mediated Halogen Exchange of Aryl Iodides
 Supplementary Materials for Kinetic and Computational Studies on Pd(I) Dimer- Mediated Halogen Exchange of Aryl Iodides Indrek Kalvet, a Karl J. Bonney, a and Franziska Schoenebeck a * a Institute of Organic
Supplementary Materials for Kinetic and Computational Studies on Pd(I) Dimer- Mediated Halogen Exchange of Aryl Iodides Indrek Kalvet, a Karl J. Bonney, a and Franziska Schoenebeck a * a Institute of Organic
EPL 603 TOPICS IN SOFTWARE ENGINEERING. Lab 5: Component Adaptation Environment (COPE)
 EPL 603 TOPICS IN SOFTWARE ENGINEERING Lab 5: Component Adaptation Environment (COPE) Performing Static Analysis 1 Class Name: The fully qualified name of the specific class Type: The type of the class
EPL 603 TOPICS IN SOFTWARE ENGINEERING Lab 5: Component Adaptation Environment (COPE) Performing Static Analysis 1 Class Name: The fully qualified name of the specific class Type: The type of the class
DuPont Suva 95 Refrigerant
 Technical Information T-95 SI DuPont Suva refrigerants Thermodynamic Properties of DuPont Suva 95 Refrigerant (R-508B) The DuPont Oval Logo, The miracles of science, and Suva, are trademarks or registered
Technical Information T-95 SI DuPont Suva refrigerants Thermodynamic Properties of DuPont Suva 95 Refrigerant (R-508B) The DuPont Oval Logo, The miracles of science, and Suva, are trademarks or registered
; +302 ; +313; +320,.
 1.,,*+, - +./ +/2 +, -. ; +, - +* cm : Key words: snow-water content, surface soil, snow type, water permeability, water retention +,**. +,,**/.. +30- +302 ; +302 ; +313; +320,. + *+, *2// + -.*, **. **+.,
1.,,*+, - +./ +/2 +, -. ; +, - +* cm : Key words: snow-water content, surface soil, snow type, water permeability, water retention +,**. +,,**/.. +30- +302 ; +302 ; +313; +320,. + *+, *2// + -.*, **. **+.,
EE101: Resonance in RLC circuits
 EE11: Resonance in RLC circuits M. B. Patil mbatil@ee.iitb.ac.in www.ee.iitb.ac.in/~sequel Deartment of Electrical Engineering Indian Institute of Technology Bombay I V R V L V C I = I m = R + jωl + 1/jωC
EE11: Resonance in RLC circuits M. B. Patil mbatil@ee.iitb.ac.in www.ee.iitb.ac.in/~sequel Deartment of Electrical Engineering Indian Institute of Technology Bombay I V R V L V C I = I m = R + jωl + 1/jωC
ΕΦΑΡΜΟΓΗ ΕΥΤΕΡΟΒΑΘΜΙΑ ΕΠΕΞΕΡΓΑΣΜΕΝΩΝ ΥΓΡΩΝ ΑΠΟΒΛΗΤΩΝ ΣΕ ΦΥΣΙΚΑ ΣΥΣΤΗΜΑΤΑ ΚΛΙΝΗΣ ΚΑΛΑΜΙΩΝ
 ΤΕΧΝΟΛΟΓΙΚΟ ΕΚΠΑΙ ΕΥΤΙΚΟ Ι ΡΥΜΑ ΚΡΗΤΗΣ ΤΜΗΜΑ ΦΥΣΙΚΩΝ ΠΟΡΩΝ ΚΑΙ ΠΕΡΙΒΑΛΛΟΝΤΟΣ ΕΦΑΡΜΟΓΗ ΕΥΤΕΡΟΒΑΘΜΙΑ ΕΠΕΞΕΡΓΑΣΜΕΝΩΝ ΥΓΡΩΝ ΑΠΟΒΛΗΤΩΝ ΣΕ ΦΥΣΙΚΑ ΣΥΣΤΗΜΑΤΑ ΚΛΙΝΗΣ ΚΑΛΑΜΙΩΝ ΕΠΙΜΕΛΕΙΑ: ΑΡΜΕΝΑΚΑΣ ΜΑΡΙΝΟΣ ΧΑΝΙΑ
ΤΕΧΝΟΛΟΓΙΚΟ ΕΚΠΑΙ ΕΥΤΙΚΟ Ι ΡΥΜΑ ΚΡΗΤΗΣ ΤΜΗΜΑ ΦΥΣΙΚΩΝ ΠΟΡΩΝ ΚΑΙ ΠΕΡΙΒΑΛΛΟΝΤΟΣ ΕΦΑΡΜΟΓΗ ΕΥΤΕΡΟΒΑΘΜΙΑ ΕΠΕΞΕΡΓΑΣΜΕΝΩΝ ΥΓΡΩΝ ΑΠΟΒΛΗΤΩΝ ΣΕ ΦΥΣΙΚΑ ΣΥΣΤΗΜΑΤΑ ΚΛΙΝΗΣ ΚΑΛΑΜΙΩΝ ΕΠΙΜΕΛΕΙΑ: ΑΡΜΕΝΑΚΑΣ ΜΑΡΙΝΟΣ ΧΑΝΙΑ
Supporting information. An unusual bifunctional Tb-MOF for highly sensing of Ba 2+ ions and remarkable selectivities of CO 2 /N 2 and CO 2 /CH 4
 Electronic Supplementary Material (ESI) for Journal of Materials Chemistry A. This journal is The Royal Society of Chemistry 2015 Supporting information An unusual bifunctional Tb-MOF for highly sensing
Electronic Supplementary Material (ESI) for Journal of Materials Chemistry A. This journal is The Royal Society of Chemistry 2015 Supporting information An unusual bifunctional Tb-MOF for highly sensing
6.1. Dirac Equation. Hamiltonian. Dirac Eq.
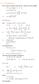 6.1. Dirac Equation Ref: M.Kaku, Quantum Field Theory, Oxford Univ Press (1993) η μν = η μν = diag(1, -1, -1, -1) p 0 = p 0 p = p i = -p i p μ p μ = p 0 p 0 + p i p i = E c 2 - p 2 = (m c) 2 H = c p 2
6.1. Dirac Equation Ref: M.Kaku, Quantum Field Theory, Oxford Univ Press (1993) η μν = η μν = diag(1, -1, -1, -1) p 0 = p 0 p = p i = -p i p μ p μ = p 0 p 0 + p i p i = E c 2 - p 2 = (m c) 2 H = c p 2
Single-site association results for 136 SCARB1 genotyped variants with HDL-C.
 Table S9. Single-site association results for 136 SCARB1 genotyped variants with HDL-C. SNP Name a SNP ID b Chr12 Position c Location Amino Acid Change RegDB Score d MA, MAF Genotype Genotype Count Adjusted
Table S9. Single-site association results for 136 SCARB1 genotyped variants with HDL-C. SNP Name a SNP ID b Chr12 Position c Location Amino Acid Change RegDB Score d MA, MAF Genotype Genotype Count Adjusted
Supplementary Table 1. Primers used for RT-qPCR analysis of striatal and nigral tissue.
 Supplementary Table 1. Primers used for RT-qPCR analysis of striatal and nigral tissue. Gene Forward Primer (5-3 ) Reverse Primer (5-3 ) Dopaminergic Markers TH CTG GCC ATT GAT GTA CTG GA ACA CAC ATG GGA
Supplementary Table 1. Primers used for RT-qPCR analysis of striatal and nigral tissue. Gene Forward Primer (5-3 ) Reverse Primer (5-3 ) Dopaminergic Markers TH CTG GCC ATT GAT GTA CTG GA ACA CAC ATG GGA
Advanced Subsidiary Unit 1: Understanding and Written Response
 Write your name here Surname Other names Edexcel GE entre Number andidate Number Greek dvanced Subsidiary Unit 1: Understanding and Written Response Thursday 16 May 2013 Morning Time: 2 hours 45 minutes
Write your name here Surname Other names Edexcel GE entre Number andidate Number Greek dvanced Subsidiary Unit 1: Understanding and Written Response Thursday 16 May 2013 Morning Time: 2 hours 45 minutes
Resurvey of Possible Seismic Fissures in the Old-Edo River in Tokyo
 Bull. Earthq. Res. Inst. Univ. Tokyo Vol. 2.,**3 pp.,,3,.* * +, -. +, -. Resurvey of Possible Seismic Fissures in the Old-Edo River in Tokyo Kunihiko Shimazaki *, Tsuyoshi Haraguchi, Takeo Ishibe +, -.
Bull. Earthq. Res. Inst. Univ. Tokyo Vol. 2.,**3 pp.,,3,.* * +, -. +, -. Resurvey of Possible Seismic Fissures in the Old-Edo River in Tokyo Kunihiko Shimazaki *, Tsuyoshi Haraguchi, Takeo Ishibe +, -.
ΤΕΧΝΟΛΟΓΙΚΟ ΕΚΠΑΙΔΕΥΤΙΚΟ ΙΔΡΥΜΑ ΚΡΗΤΗΣ ΣΧΟΛΗ ΔΙΟΙΚΗΣΗΣ ΚΑΙ ΟΙΚΟΝΟΜΙΑΣ (ΣΔΟ) ΤΜΗΜΑ ΛΟΓΙΣΤΙΚΗΣ ΚΑΙ ΧΡΗΜΑΤΟΟΙΚΟΝΟΜΙΚΗΣ
 ΤΕΧΝΟΛΟΓΙΚΟ ΕΚΠΑΙΔΕΥΤΙΚΟ ΙΔΡΥΜΑ ΚΡΗΤΗΣ ΣΧΟΛΗ ΔΙΟΙΚΗΣΗΣ ΚΑΙ ΟΙΚΟΝΟΜΙΑΣ (ΣΔΟ) ΤΜΗΜΑ ΛΟΓΙΣΤΙΚΗΣ ΚΑΙ ΧΡΗΜΑΤΟΟΙΚΟΝΟΜΙΚΗΣ «ΧΡΗΜΑΤΟΟΙΚΟΝΟΜΙΚΗ ΑΝΑΛΥΣΗ ΛΟΓΙΣΤΙΚΩΝ ΚΑΤΑΣΤΑΣΕΩΝ ΤΩΝ ΕΤΑΙΡΕΙΩΝ ΚΡΕΤΑ ΦΑΡΜ Α.Β.Ε.Ε. &
ΤΕΧΝΟΛΟΓΙΚΟ ΕΚΠΑΙΔΕΥΤΙΚΟ ΙΔΡΥΜΑ ΚΡΗΤΗΣ ΣΧΟΛΗ ΔΙΟΙΚΗΣΗΣ ΚΑΙ ΟΙΚΟΝΟΜΙΑΣ (ΣΔΟ) ΤΜΗΜΑ ΛΟΓΙΣΤΙΚΗΣ ΚΑΙ ΧΡΗΜΑΤΟΟΙΚΟΝΟΜΙΚΗΣ «ΧΡΗΜΑΤΟΟΙΚΟΝΟΜΙΚΗ ΑΝΑΛΥΣΗ ΛΟΓΙΣΤΙΚΩΝ ΚΑΤΑΣΤΑΣΕΩΝ ΤΩΝ ΕΤΑΙΡΕΙΩΝ ΚΡΕΤΑ ΦΑΡΜ Α.Β.Ε.Ε. &
Table of Contents 1 Supplementary Data MCD
 Electronic Supplementary Material (ESI) for Dalton Transactions. This journal is The Royal Society of Chemistry 2017 Supporting Information for Magnetic circular dichroism and density functional theory
Electronic Supplementary Material (ESI) for Dalton Transactions. This journal is The Royal Society of Chemistry 2017 Supporting Information for Magnetic circular dichroism and density functional theory
DuPont Suva. DuPont. Thermodynamic Properties of. Refrigerant (R-410A) Technical Information. refrigerants T-410A ENG
 Technical Information T-410A ENG DuPont Suva refrigerants Thermodynamic Properties of DuPont Suva 410A Refrigerant (R-410A) The DuPont Oval Logo, The miracles of science, and Suva, are trademarks or registered
Technical Information T-410A ENG DuPont Suva refrigerants Thermodynamic Properties of DuPont Suva 410A Refrigerant (R-410A) The DuPont Oval Logo, The miracles of science, and Suva, are trademarks or registered
Θέματα Στατιστικής στη γλώσσα R
 Θέματα Στατιστικής στη γλώσσα R Ποσότητες οδηγοί και τα ποσοστιαία σημεία των αντίστοιχων κατανομών Ν(0,1) Student s t X 2, F Διαστήματα εμπιστοσύνης-έλεγχοι Υποθέσεων ένα δείγμα για τη μέση τιμή κανονικής
Θέματα Στατιστικής στη γλώσσα R Ποσότητες οδηγοί και τα ποσοστιαία σημεία των αντίστοιχων κατανομών Ν(0,1) Student s t X 2, F Διαστήματα εμπιστοσύνης-έλεγχοι Υποθέσεων ένα δείγμα για τη μέση τιμή κανονικής
A Bonus-Malus System as a Markov Set-Chain. Małgorzata Niemiec Warsaw School of Economics Institute of Econometrics
 A Bonus-Malus System as a Markov Set-Chain Małgorzata Niemiec Warsaw School of Economics Institute of Econometrics Contents 1. Markov set-chain 2. Model of bonus-malus system 3. Example 4. Conclusions
A Bonus-Malus System as a Markov Set-Chain Małgorzata Niemiec Warsaw School of Economics Institute of Econometrics Contents 1. Markov set-chain 2. Model of bonus-malus system 3. Example 4. Conclusions
HISTOGRAMS AND PERCENTILES What is the 25 th percentile of a histogram? What is the 50 th percentile for the cigarette histogram?
 HISTOGRAMS AND PERCENTILES What is the 25 th percentile of a histogram? The point on the horizontal axis such that of the area under the histogram lies to the left of that point (and to the right) What
HISTOGRAMS AND PERCENTILES What is the 25 th percentile of a histogram? The point on the horizontal axis such that of the area under the histogram lies to the left of that point (and to the right) What
Long intergenic non-coding RNA expression signature in human breast cancer
 Long intergenic non-coding RNA expression signature in human breast cancer Yanfeng Zhang 1,5,#, Erin K. Wagner 2,#, Xingyi Guo 1, Isaac May 3, Qiuyin Cai 1, Wei Zheng 1, Chunyan He 2,4,*, Jirong Long 1,*
Long intergenic non-coding RNA expression signature in human breast cancer Yanfeng Zhang 1,5,#, Erin K. Wagner 2,#, Xingyi Guo 1, Isaac May 3, Qiuyin Cai 1, Wei Zheng 1, Chunyan He 2,4,*, Jirong Long 1,*
Για να ελέγξουµε αν η κατανοµή µιας µεταβλητής είναι συµβατή µε την κανονική εφαρµόζουµε το test Kolmogorov-Smirnov.
 A. ΈΛΕΓΧΟΣ ΚΑΝΟΝΙΚΟΤΗΤΑΣ A 1. Έλεγχος κανονικότητας Kolmogorov-Smirnov. Για να ελέγξουµε αν η κατανοµή µιας µεταβλητής είναι συµβατή µε την κανονική εφαρµόζουµε το test Kolmogorov-Smirnov. Μηδενική υπόθεση:
A. ΈΛΕΓΧΟΣ ΚΑΝΟΝΙΚΟΤΗΤΑΣ A 1. Έλεγχος κανονικότητας Kolmogorov-Smirnov. Για να ελέγξουµε αν η κατανοµή µιας µεταβλητής είναι συµβατή µε την κανονική εφαρµόζουµε το test Kolmogorov-Smirnov. Μηδενική υπόθεση:
SUPPORTING INFORMATION
 SUPPRTING INFRMATIN Per(-guanidino--deoxy)cyclodextrins: Synthesis, characterisation and binding behaviour toward selected small molecules and DNA. Nikolaos Mourtzis, a Kyriaki Eliadou, a Chrysie Aggelidou,
SUPPRTING INFRMATIN Per(-guanidino--deoxy)cyclodextrins: Synthesis, characterisation and binding behaviour toward selected small molecules and DNA. Nikolaos Mourtzis, a Kyriaki Eliadou, a Chrysie Aggelidou,
UDZ Swirl diffuser. Product facts. Quick-selection. Swirl diffuser UDZ. Product code example:
 UDZ Swirl diffuser Swirl diffuser UDZ, which is intended for installation in a ventilation duct, can be used in premises with a large volume, for example factory premises, storage areas, superstores, halls,
UDZ Swirl diffuser Swirl diffuser UDZ, which is intended for installation in a ventilation duct, can be used in premises with a large volume, for example factory premises, storage areas, superstores, halls,
A facile and general route to 3-((trifluoromethyl)thio)benzofurans and 3-((trifluoromethyl)thio)benzothiophenes
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 A facile and general route to 3-((trifluoromethyl)thio)benzofurans and 3-((trifluoromethyl)thio)benzothiophenes
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 A facile and general route to 3-((trifluoromethyl)thio)benzofurans and 3-((trifluoromethyl)thio)benzothiophenes
Partial Trace and Partial Transpose
 Partial Trace and Partial Transpose by José Luis Gómez-Muñoz http://homepage.cem.itesm.mx/lgomez/quantum/ jose.luis.gomez@itesm.mx This document is based on suggestions by Anirban Das Introduction This
Partial Trace and Partial Transpose by José Luis Gómez-Muñoz http://homepage.cem.itesm.mx/lgomez/quantum/ jose.luis.gomez@itesm.mx This document is based on suggestions by Anirban Das Introduction This
Προσομοίωση BP με το Bizagi Modeler
 Προσομοίωση BP με το Bizagi Modeler Α. Τσαλγατίδου - Γ.-Δ. Κάπος Πρόγραμμα Μεταπτυχιακών Σπουδών Τεχνολογία Διοίκησης Επιχειρησιακών Διαδικασιών 2017-2018 BPMN Simulation with Bizagi Modeler: 4 Levels
Προσομοίωση BP με το Bizagi Modeler Α. Τσαλγατίδου - Γ.-Δ. Κάπος Πρόγραμμα Μεταπτυχιακών Σπουδών Τεχνολογία Διοίκησης Επιχειρησιακών Διαδικασιών 2017-2018 BPMN Simulation with Bizagi Modeler: 4 Levels
Technical Information Efficiency and Derating SUNNY BOY / SUNNY TRIPOWER / SUNNY MINI CENTRAL
 Technical Information Efficiency and Derating SUNNY BOY / SUNNY TRIPOWER / SUNNY MINI CENTRAL WirkungDerat-TI-en-40 Version 4.0 ENGLISH Legal Provisions SMA Solar Technology AG Legal Provisions The information
Technical Information Efficiency and Derating SUNNY BOY / SUNNY TRIPOWER / SUNNY MINI CENTRAL WirkungDerat-TI-en-40 Version 4.0 ENGLISH Legal Provisions SMA Solar Technology AG Legal Provisions The information
encouraged to use the Version of Record that, when published, will replace this version. The most /BCJ
 Biochemical Journal: this is an Accepted Manuscript, not the final Version of Record. You are encouraged to use the Version of Record that, when published, will replace this version. The most up-to-date
Biochemical Journal: this is an Accepted Manuscript, not the final Version of Record. You are encouraged to use the Version of Record that, when published, will replace this version. The most up-to-date
Σχέση µεταξύ της Μεθόδου των ερµατοπτυχών και της Βιοηλεκτρικής Αντίστασης στον Υπολογισµό του Ποσοστού Σωµατικού Λίπους
 Ερευνητική Aναζητήσεις στη Φυσική Αγωγή & τον Αθλητισµότόµος 1(3), 244 251 ηµοσιεύτηκε: 2 8 εκεµβρίου 2003 Inquiries in Sport & Physical Education Volume 1(3), 244 251 Released: December 28, 2003 www.hape.gr/emag.asp
Ερευνητική Aναζητήσεις στη Φυσική Αγωγή & τον Αθλητισµότόµος 1(3), 244 251 ηµοσιεύτηκε: 2 8 εκεµβρίου 2003 Inquiries in Sport & Physical Education Volume 1(3), 244 251 Released: December 28, 2003 www.hape.gr/emag.asp
Consolidated Drained
 Consolidated Drained q, 8 6 Max. Shear c' =.185 φ' =.8 tan φ' =.69 Deviator, 8 6 6 8 1 1 p', 5 1 15 5 Axial, Symbol Sample ID Depth Test Number Height, in Diameter, in Moisture Content (from Cuttings),
Consolidated Drained q, 8 6 Max. Shear c' =.185 φ' =.8 tan φ' =.69 Deviator, 8 6 6 8 1 1 p', 5 1 15 5 Axial, Symbol Sample ID Depth Test Number Height, in Diameter, in Moisture Content (from Cuttings),
ΠΑΡΑΜΕΤΡΟΙ ΕΠΗΡΕΑΣΜΟΥ ΤΗΣ ΑΝΑΓΝΩΣΗΣ- ΑΠΟΚΩΔΙΚΟΠΟΙΗΣΗΣ ΤΗΣ BRAILLE ΑΠΟ ΑΤΟΜΑ ΜΕ ΤΥΦΛΩΣΗ
 ΠΑΝΕΠΙΣΤΗΜΙΟ ΜΑΚΕΔΟΝΙΑΣ ΟΙΚΟΝΟΜΙΚΩΝ ΚΑΙ ΚΟΙΝΩΝΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΤΜΗΜΑ ΕΚΠΑΙΔΕΥΤΙΚΗΣ ΚΑΙ ΚΟΙΝΩΝΙΚΗΣ ΠΟΛΙΤΙΚΗΣ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ ΠΑΡΑΜΕΤΡΟΙ ΕΠΗΡΕΑΣΜΟΥ ΤΗΣ ΑΝΑΓΝΩΣΗΣ- ΑΠΟΚΩΔΙΚΟΠΟΙΗΣΗΣ ΤΗΣ BRAILLE
ΠΑΝΕΠΙΣΤΗΜΙΟ ΜΑΚΕΔΟΝΙΑΣ ΟΙΚΟΝΟΜΙΚΩΝ ΚΑΙ ΚΟΙΝΩΝΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΤΜΗΜΑ ΕΚΠΑΙΔΕΥΤΙΚΗΣ ΚΑΙ ΚΟΙΝΩΝΙΚΗΣ ΠΟΛΙΤΙΚΗΣ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ ΠΑΡΑΜΕΤΡΟΙ ΕΠΗΡΕΑΣΜΟΥ ΤΗΣ ΑΝΑΓΝΩΣΗΣ- ΑΠΟΚΩΔΙΚΟΠΟΙΗΣΗΣ ΤΗΣ BRAILLE
Supporting information. Influence of Aerosol Acidity on the Chemical Composition of Secondary Organic Aerosol from β caryophyllene
 Supporting information Influence of Aerosol Acidity on the Chemical Composition of Secondary Organic Aerosol from β caryophyllene M. N. Chan 1, J. D. Surratt 2,*, A. W. H. Chan 2,**, K. Schilling 2, J.
Supporting information Influence of Aerosol Acidity on the Chemical Composition of Secondary Organic Aerosol from β caryophyllene M. N. Chan 1, J. D. Surratt 2,*, A. W. H. Chan 2,**, K. Schilling 2, J.
Supplementary Table 1. Primer sequences used for RT-qPCR validation of the microarray data. Primer. direction. Forward
 Supplementary Table 1. Primer sequences used for RT-qPCR validation of the microarray data Gene Glyceraldehyde 3-phosphate dehydrogenase (Gapdh) Cell death-inducing DFFAlike effector a (Cidea) Peroxisome
Supplementary Table 1. Primer sequences used for RT-qPCR validation of the microarray data Gene Glyceraldehyde 3-phosphate dehydrogenase (Gapdh) Cell death-inducing DFFAlike effector a (Cidea) Peroxisome
