|
|
|
- Τυρώ Ανδρεάδης
- 6 χρόνια πριν
- Προβολές:
Transcript
1
2
3
4
5
6
7 Supplemental Figure 1 Genechip data analysis. Three pairs of bulge and sub-bulge ORS Genechips were analyzed and compared by MASv5.0 and dchip 1.3 software algorithms in order to identify gene transcripts that were over-represented or under-represented in the bulge ORS by both algorithms. Two additional sets of Genechips were generated form the ORS regions of upper-bulge, bulge, inner-bulge, sub-bulge and suprabulbar ORS in order to precisely localize gene expression within the hair follicle. Genes that were not selectively and specifically over- or under-represented in the bulge ORS were excluded. Each Genechip pair or set was generated from an individual volunteer. Supplemental Figure 2 The correlation between mrna expression patterns and immunohistochemical findings for gene transcripts over-represented in bulge ORS. The upper panels demonstrate human hair follicle vertical sections stained with designated mabs. Scale bars: 50µm.The lower boxes indicate the Genechip signal intensities (mrna expression levels) detected for individual genes from corresponding ORS subsets. Genechips were normalized to the same parameters. The gene expression patterns correlated well with the immunohistochemical staining patterns of gene products in the ORS. Red and blue numbers indicate individual samples. P: detected as expressed or present according to MASv5.0 algorithm. A: detected as not expressed or absent according to MASv5.0 algorithm. Parenthesis; Probe set ID of Affymetrix U-133 Genechip sets. *Epidermis; Signal intensities in Genechips generated from human foreskin epidermis are shown.
8 Total RNA was isolated from epidermal sheet and RNA probes were generated by singleround amplification. Normalized to the same parameters as ORS Genechips. Supplemental Figure 3 Anti-FZD1 monoclonal antibody could detect the bulge ORS cells in fixed midfollicle cell suspension but not living mid-follicle cells. (A) Consistent with microarray and immunohistochemistry data, approximately 70% of fixed mid-follicle cells were variably FZD1 positive by FACS analysis. (B) The FZD1 hi FST hi cells in fixed midfollicle suspension represented bulge ORS cells. (C) Anti-FZD1 antibody could only detect approximately 1% of living mid-follicle cells, suggesting that FZD1 detection by this mab requires fixation. Supplemental Figure 4 Mid-follicle CD200 hi cells were CD45 lo c-kit lo and α6 integrin hi. (A) A midfollicle cell suspension was stained for CD200 and CD45. Most of CD200 hi cells were distinct form CD45 hi bone marrow derived cells. (B) A mid-follicle cell suspension was stained for CD200 and c-kit (CD117). Most of CD200 hi cells were distinct from c-kit hi melanocytes or mast cells. (C) A mid-follicle cell suspension was stained for CD200 and α6 integrin (CD49f). Most CD200 hi cells were α6 integrin hi. Gated to non-fixed living cells based on 7-AAD negative staining. Supplemental Figure 5 Genechip data sets for the CD markers identified in human hair follicle. Numbers
9 in boxes indicate the signal intensities for each CD marker (mrna expression levels) in the indicated ORS subsets. Red and blue numbers indicate individual experiments. P: detected as expressed or present according to MASv5.0 algorithm. A: detected as not expressed or absent according to MASv5.0 algorithm. Parenthesis; Probe set ID of Affymetrix U-133 Genechip sets. *Epidermis; signal intensities in Genechips generated from human foreskin epidermis. Total RNA was isolated from epidermal sheet and RNA probes were generated by single-round amplification. Normalized to the same parameters as ORS Genechips. Supplemental Figure 6 Isolated CD200 hi BNC lo bulge cells were uniform, relatively compact and smooth when compared to mid-follicle cells or CD59+ hair follicle cells. CD200 hi BNC lo bulge cells were isolated from mid-follicle cell suspension by magnetic beads selection. (A) By light microscopy, isolated living bulge CD200 hi cells (Arrows) possessed compact and smooth morphology compared to mid-follicle (arrowheads) cells or CD59 hi cells (open arrows) isolated from mid-follicle suspension. Stained with trypan blue. Original magnification 400x. (B) FACS analysis confirmed the microscopic observation. Living 7- AAD negative cells were gated. Magnetic beads were also gated out.
10 Supplemental table 1. Genes detected as up-regulated in human bulge by MAS 5.0 software Gene Probe set Mouse bulge Bulge 1 Sub-bulge 1 Fold change 1 Bulge 2 Sub-bulge 2 Fold change 2 Bulge 3 Sub-bulge 3 Fold change 3 Average Fold change trinucleotide repeat containing 9 (TNRC9) _x_at P 38.3 P P 1.6 A P 12.9 A trinucleotide repeat containing 9 (TNRC9) _x_at P 33.8 P P 2.2 A P 32.5 A deiodinase, iodothyronine, type II (DIO2) _s_at P P P 27.5 A P P Small proline-rich protein SPRK human, odontogenic keratocysts (SPRR1A) _at P P P 86.8 A P 33.9 A adenylate cyclase 2 (brain) (ADCY2) _at A A A 8.9 A A 89.6 A cystatin EM (CST6) _at P 1310 P P 201 P P P DBCCR1-like (DCCR1L) _at 82.3 P 33.2 P P 10.6 A P 20.4 A Wnt inhibitory factor-1 (WIF1) _at *** P P P P P P deiodinase, iodothyronine, type II (DIO2), transcript variant _s_at P 197 P P 91.6 P P P aldo-keto reductase family 1, member C3 (3-alpha hydroxysteroid dehydrogenase, type II) (AKR1C3) _at P 66.9 P P 53.5 P P 15 A trinucleotide repeat containing 9 (TNRC9) _x_at P 26.6 P P 15.8 A P 22.8 A dickkopf (Xenopus laevis) homolog 3 (DKK3) _s_at *,** P P P 40.1 A P 41.6 P aquaporin 3 (AQP3) 39248_at P P P 312 P P P pleckstrin homology-like domain, family A, member 1(PHLDA1) _at P 63.2 P P 27.5 A P 60.4 P calcium/calmodulin-dependent protein kinasell (CaMKllNalpha) _at 559 P P P 55.6 P P P dopachrome tautomerase (dopachrome delta-isomerase, tyrosine-related protein 2) (DCT) _at 209 P P P 21.8 A P P monooxygenase, DBH-like 1 (MOXD1) _at P 71.7 P P 69.3 P P 76.4 A estrogen receptor 1 (ESR1) _at P P P 82.8 A P 84.2 M mrna; cdna DKFZp779H233 (from clone DKFZp779H233) _at 29.4 P 27.3 A P 6.9 A P 8.9 A creatine kinase, mitochondrial 1 (ubiquitous) (CKMT1), nuclear gene encoding mitochondrial protein _s_at P 89.5 P P 61 A P 70.3 P dickkopf (Xenopus laevis) homolog 3 (DKK3) _s_at *,** P P P 669 P P P RAS guanyl releasing protein 1 (calcium and DAG-regulated) (RASGRP1) _at P 37.3 P P 33.1 A P 36.2 P sema domain, transmembrane domain (TM), and cytoplasmic domain, (semaphorin6a) (SEMA6A) _at P 68.5 P P 28.2 M P 40.7 A angiopoietin-like 2 (ANGPTL2) _at P P P 65 P P 43.1 P protein tyrosine phosphate, receptor type, D (PTPRD) _at 117 P 55.4 M A 7.6 A P 51.2 A family with sequence similarity 13, membera1 (FAM13A1) _s_at P P P 154 P P P Ral guanine nucleotide exchange factor RalGPS1A (RalGPS1A) _at P 60 P P 49.8 A P 30.3 A Forkhead box C1 (FOXC1) _at 618 P P P 99.3 P P P zinc finger, BED domain containing 1 (ZBED1) _at P P P 53.9 A P 342 P nebulette (NEBL) _s_at P 136 P P 60.9 P P 70.8 P pleiomorphic adenoma gene 1 (PLAG1) _at P P P 32.7 P P 77.1 P phosphoinositide-3-kinase, regulatory subunit, polypeptide 1 (p85 alpha) (PIK3R1) _at P P P P P P angiopoietin-like 2 (ANGPTL2) _at P P P P P P fibroblast growth factor 13 (FGF13) _s_at P 44.7 A P 36 A P 42.2 A signal peptide, CUB domain, EGF-like _s_at P 94.3 P P P P P dihydropyrimidinase-like 2 (DPYSL2) _at P P P P P P seizure related 6 homolog (mouse)-like (SEZ6L) _s_at P 74 A P 42.2 A P A decorin (DCN) _s_at P P P P P P nebulette (NEBL) _at P P P P P P LERK-5 (EPLG5, ephrin-b2, EFNB2) _s_at * P P P 38.2 A P P hypothetical protein FLJ10847 (FLJ10847) _at P P P 90.2 A P P hypothetical protein FLJ11795 (FLJ11795) _at 74.4 P 34.1 A P 55.6 A P 11.9 A SCLAC2-B _at P P P P P P serine (or cysteine) proteinase inhibitor, clade ), member 1 (SERPINF1) _at P P P 85.8 A P P dihydropyrimidinase-like 3 (DPYSL3) _s_at P P P 87.2 M P P protein tyrosine phosphate, receptor type, D (PTPRD) _at P P P 93.1 P P 115 P very long-chain acyl-coa synthetase; lipidosin (BG1) _at P P P 50.2 A M 80.3 A major histocompatibility complex, classll, DQ beta2 (HLA-DQB2) _at P 48.3 P P 89.4 P P 56.3 M transforming growth factor-beta-2 (TGFB2) _s_at * P P P 33.2 P P 91.9 P golgi phosphoprotein 4 (GOLPH4) _s_at P P P P P P phospholipase C-beta-1 (PLCB1, PLC-I) _at P P P P P P decorin (DCN) _x_at P 1447 P P P P P latrophilin 3 (LPHN3) _s_at P P P P P P abhydrolase domain containing 6 (ABDH6) 45288_at P 72.1 P P 42.2 P P 53.3 P Ca2+-dependent acivator protein for secretion 2 (CADPS2) _at 369 P P P P P P aldo-keto reductase family1, member C2 (AKR1C2) _x_at P 59.1 P P 37 A A 89.6 P met proto-oncogene (hepatocyte growth factor receptor) (MET) _at P P P 67.1 P P 80.6 P KIAA1036 protein _s_at P P P P P P Keratin15 (KRT15) _at ** P P P P P P bromodomain containing 2 (BRD2) _s_at P P P P P P growth-arrest-specific 6 (Gas6) 1598_g_at *,**(Gas1) 995 P P P 306 P P P C-terminal binding protein 2 (CTBP2) _s_at * P P P P P P dopachrome tautomerase (dopachrome delta-isomerase, tyrosine-related protein 2) (DCT) _s_at P P P 95 A P P CD200 antigen (CD200) _s_at ** P P P 86 P P P S Ribosomal Protein L21 pseudogene and an HNRNP A3 pseudogene _x_at 36.4 P 24.6 A P 14.1 A P 42.5 A glycoprotein M6B (GPM6B) _at P A P A M A latrophilin 3 (LPHN3) _s_at P P P P P P chromogranin A (parathyroid secretory protein 1) (CHGA) _s_at 76.8 P 62.3 A P 41.1 A P 65.7 A collagen, type VII, alpha 1 (epidermolysis bullosa, dystrophic, dominant and recessive) (COL7A1) _at P P P P P P solute carrier family 43, member 3 (SLC43A3) _s_at P P P P P P inhibitor of DNA binding 2, dominant negative helix-loop-helix protein (ID2) _s_at *,** A A M A A A Rho guanine nucleotide exchange factor (GEF) 10 (ARHGEF10) _s_at P P P P P P glycoprotein M6B (GPM6B) _s_at 234 P P P 51.9 P P P nuclear factor (erythroid-derived 2)-like _s_at P P P 99.6 A P P pleckstrin homology domain-containing, family A (phosphoinositide binding specific) member 1 (PLEKHA1) _at 273 P P P P P P Hypothetical LOC _x_at 41.8 P 35.9 A P 28.5 A P 13.7 A Follistatin (FST) _s_at *(Fstl) P A P A P A zinc finger protein (ZFD25) _x_at 122 P 79.3 P P 60.2 M P 37.9 P C-terminal binding protein 2 (CTBP2), transcript variant _x_at *(CTBP2) P P P P P P regulated in glioma (RIG) _s_at P P P 57.5 A P 83.8 P dual specificity phosphatase 1 (DUSP1) _s_at P P P P P P natural killer-tumor recognition sequence (NKTR) _s_at P P P P P P TDP-glucose 4,6-dehydratase (TGDS) _s_at P P P P P P A kinase (PRKA) anchor protein 1 (AKAP1) _s_at A A A A P A ubiquitin-conjugating enzyme E2E 1 (UBC4/5 homolog, yeast) _at 90.7 P 46.8 A P 48.4 A P 56.4 A chromosome 14 open reading frame 78 (C14orf78) _at 1942 P P P P P P Homo sapiens frizzled (Drosophila) homolog 1 (FZD1) _at *,**(FZD2) P P P P P P FXYD domain-containing ion transport regulator 6 (FXYD6) _at P P P P P P parvin, alpha (PARVA) _at P 74.7 A P 76.7 A P 59.7 A dystonin (DST) _s_at P P P P P P hypothetical protein LOC _at P P P A P P nuclear factor of activated T-cells 5, tonicity-resonsive (NFAT5) _s_at P P P P P P recombining binding protein supressor of hairless (Drosophila) (RBPSUH) _x_at P 751 P P P P P immunoglobulin superfamily, member 4 (IGSF4) _at *,** P P P P P 377 P nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 1 (NFATC1) _s_at * 439 P P P 297 P P P POU domain, class 2, transcription factor 1 (POU2F1) _s_at P A P 119 A P 80.2 A transient receptor potential cation channel, subfamily V, member 2) _s_at 55.1 P 22.3 A P 30.4 A P 32.5 A calmodulin 1(phosphorylase kinase, delta) (CALM1) _at P 287 P P P P P plexin A1 (PLXNA1) _at **(Plxn2) 92.9 P A P 54.6 A P 83.2 A zinc fingers ans homeoboxes 3 (ZHX3) _s_at 87.5 P 70 A P 77.2 A P 69.7 A nebulette protein (NEBL) _s_at 83.5 P 53 A P 63.8 A P 57.9 A Numbers indicate signal intensities detected for each probe sets P: "present" call, detected as expressed, A: "Absent" call, detected as not expressed. Bold: Up-regulated in both MAS 5.0 and dchip analysis Underline: "membrane" protein annotated by GOA analysis Mouse bulge column: asterisks indicate the up-regulation of same or similar genes in mouse bulge transcription profiling data * Tumbar et al. Science, , **Morris et al. Nat Biotechnol, , *** Fuchs et al. Cell , parenthesis; gene of similar function was detected as upregulated.
11 Supplemental table 2. Genes detected as down-regulated in human bulge by MAS 5.0 software Gene Probe set Bulge 1 Sub-bulge 1 Fold change 1 Bulge 2 Sub-bulge 2 Fold change 2 Bulge 3 Sub-bulge 3 fold change 3 Average fold change microfibrillar associated protein 5 (MFAP5) _s_at 72.2P 648.8P P 471.3P A 108.1P cartilage oligomeric matrix protein (pseudoachondroplasia, epiphyseal dysplasia 1, multiple) (COMP) _s_at 113.9P P P 992.1P A P angiopoietin-like 7 (ANGPTL7) _at 628.5P P P 5226P P P endothelin receptor type B (EDNRB) _at 82.5P 264.1P P 187.6P A 44.5P aldehyde oxidase 1 (AOX1) _at 119.3A 163.8P M 149.1P A 48.9A wingless-type MMTV integration site family, member 5A (WNT5A) _at 48.2P 652.6P P 467.7P P 143.9P topoisomerase (DNA) II alpha (170kD) (TOP2A) _at 80.5P 198.4P P 469.2P A 207.7P protocadherin 8 (PCDH8) _at 39.5P 225.4P P 314.6P P 187.9P aminolevulinate, delta-, dehydratase (ALAD) _at 180.5P P P P A 638.7P calcium channel voltage-dependent alpha2/delta3 subunit (CACNA2D3) _s_at 10.2A 57.6A A 120.9A A 48.1A microfibrillar associated protein 5 (MFAP5) _at 66.7P 291.5P P 117.3P A 60P tripartite motif protein TRIM9 isoform alpha (TRIM9) _at 142.3P 429P P 658.6P P 361.6P wingless-type MMTV integration site family, member 5A (WNT5A) _s_at 248.2P P P P A 412.3P fibroblast growth factor 18 (FGF18) _x_at 63.3P 214.6P P 407P A 316P prostaglandin E receptor 4 (subtype EP4) (PTGER4) _at 141.3P 272.7P P 794.6P A 115.8P tissue inhibitor of metalloproteinase 3 (Sorsby fundus dystrophy, pseudoinflammatory) (TIMP3) _s_at 167.3P 421.1P P 652.8P A 472.3P keratin associated protein 1-3 (KRTAP1-3) _at 37.5A 97.7P A 292.6P A 97.7P fibroblast growth factor 18 (FGF18) _x_at 77.8A 234.8P P 482.1P A 318.5P matrix Gla protein (MGP) _s_at P 2617P P P P P platelet derived growth factor C (PDGFC) _at 398.9P P P P P 832.6P cell division cycle 2, G1 to S and G2 to M (CDC2) _at 69.8P 192.2P P 429.5P P 170.6P isochorismatase domain containing 1 (ISOC1) _at 299.4P P P P P 635.9P tissue inhibitor of metalloproteinase 3 (Sorsby fundus dystrophy, pseudoinflammatory) (TIMP3) _s_at 621P P P P P P syndecan 2 (heparan sulfate proteoglycan 1, cell surface-associated) (SDC2) _at 396.3P P P P P 717P ets variant gene 5 (ets-related molecule) (ETV5) _s_at 158.8P 379.7P P 384.1P P 575.2P proprotein convertase subtilisinkexin type 5 (PCSK5) _s_at 57P 112.5P P 330.3P A 58.2P hyaluronan-mediated motility receptor (RHAMM) (HMMR) _at 78.2P 173.2P P 347.3P A 188.3P EH-domain containing 3 (EHD3) _at 221.9M 484.4P P 923P P 417.6P laminin, beta 1 (LAMB1) _at 108P 261.9P P 383.3P P 116.2P growth hormone receptor (GHR) _at 103.6P 168P P 329.3P M 111.1P KIAA0101 gene product (KIAA0101) _s_at 353.7P 548.1P P P P 626.5P nucleolar and spindle associated protein 1 (NUSAP1) _at 282.7P 668.1P P P P 725P serine (or cysteine) proteinase inhibitor, clade B (ovalbumin), member 13 (SERPINB13) _s_at 134.9P 249.7P P 416.8P A 189P solute carrier family 30 (zinc transporter), member1 (SLC30A1) _at 233.3P 400.5P P P P 479.9P collagen, type XI, alpha 1 (COL11A1) 37892_at 1042P P P P P P solute carrier family 1 (glutamateneutral amino acid transporter), member 4 (SLC1A4) _s_at 242.7P 491.9P P P P 533P response gene to complement 32 (RGC32) _s_at 219.3P 435.1P P 214.2P A 132.7P lipoma HMGIC fusion partner (LHFP) _s_at 255.4P 377.5P P 727.1P A 205.5P phosphorylase, glycogen; liver (Hers disease, glycogen storage disease type VI) (PYGL) _at 278.9P 627.4P P 681.9P P 564.6P protein phosphatase 1, regulatory (inhibitor) subunit 3C (PPP1R3C) _at 397.9P 798.6P P 791.5P P 756.5P basic helix-loop-helix domain containing, class B, 3 (BHLHB3) _s_at 355.5P 497.8P P 647.8P A 467.8P glypican 4 (GPC4) _at 183.8P 278P P 397.4P P 411.5P KIAA0186 gene product (KIAA0186) _at 59A 110.7P P 178.1P A 183P MAD2 (mitotic arrest deficient, yeast, homolog)-like 1 (MAD2L1) _s_at 39.9P 135P P 245.3P P 77.6P solute carrier family 4, sodium bicarbonate cotransporter, member 7 (SLC4A7) _s_at 58.2P 165.6P P 167.5P A 40.8P Rac GTPase activating protein 1 (RACGAP1) _s_at 128.3P 295.3P P 476.9P P 193.1P carbonic anhydrase II (CA2) _at 75.5P 174.6P P 250.4P A 90.2P phospholipase A2, group IVA (cytosolic, calcium-dependent) (PLA2G4A) _at 98.9P 214.2P P 297.6P P 326.3P RAD51-interacting protein (RAD51AP) _at 39.9P 105.4P P 237.9P P 55.3P adenosine A2b receptor (ADORA2B) _at 111.1A 149.4A P 255.9P A 264.4P karyopherin alpha 2 (RAG cohort 1, importin alpha 1) (KPNA2) _s_at 71.6P 146.4P P 341.3P P 223.3P ZW10 interactor (ZWINT) _s_at 227.3P 487.8P P 821.6P P 407.3P endothelin receptor typeb (EDNRB) _s_at 189P 275.2P P 258.5P P 149.5P ribonucleotide reductase M2 polypeptide (RRM2) _at 95.8A 242P P 429.2P P 192.6P melanoma adhesion molecule (MCAM, CD146) _x_at 439P 832.8P P P P 683.5P phosphoglycerate dehydrogenase (PHGDH) _at 435.5P 686.8P P 552.5P P 1239P PRP4 pre-mrna processing factor 4 homolog (PRPF4) _at 148.7M 209.9P P 392.1P A 271.6P hypothetical protein MAC30 (MAC30) _s_at 251.1P 435.8P P 748.6P P 599.2P similar to CG9996-PA _at 193.5P 339.4P P 523.4P A 194.1P syndecan 2 (heparan sulfate proteoglycan 1, cell surface-associated, fibroglycan) (SDC2) _at 126.9P 228.6P P 297.9P M 171.8P amylo-1,6-glucosidase, 4-alpha-glucanotransferase (AGL) _s_at 472.1P 799.9P P P P 380.3P retinoic acid induced 14 (RAI14) _s_at 230P 469.5P P 358.3P P 258P influenza virus NS1A binding protein (IVNS1ABP) _at 382.1P 715.8P P 894.9P P 441.7P DNA segment on chromosome 4 (unique) 234 expressed sequence (D4S234E) _s_at 306P 354P P P P 466.3P chromosome 20 open reading frame 42 (C20orf42) _at 474P 943.2P P 1664P P 885.1P solute carrier family 39 (zinc transporter), member6 (SLC39A6) _at P P P P P P solute carrier family 1 (glutamateneutral amino acid transporter), member 4 (SLC1A4) _s_at 182.3P 332.3P P 543.9P P 302.5P adenylyl cyclase-associated protein1 (CAP1) _s_at 934.9P P P P P P mesoderm specific transcript (mouse) homolog (MEST) _at 287.1P 460.5P P 750.1P P 320.2P metastasis suppressor 1 (MTSS1) _s_at 1227P P P P P P serine (or cysteine) proteinase inhibitor, clade B (ovalbumin), member 13 (SERPINB13) _s_at 113.8P 172.5P P 236P P 63.1P chloride channel, calcium activated, family member 2 (CLCA2) _s_at 819P P P P P 725.6P homeodomain-only protein (HOP) _s_at 631.3P 848.5P P P P 898.8P glutamate-cysteine ligase, catalytic subunit (GCLC) _s_at 325.3P 743.2P P 987P P 363.2P latent transforming growth factor beta binding protein 2 (LTBP2) _at P P P P P P SMC4 (structural maintenance of chromosomes 4, yeast)-like 1 (SMC4L1) _at 177P 317.7P P 586.8P P 285.2P vav 3 oncogene (VAV3) _at 658P P P P P 741.2P tumor rejection antigen (gp96) 1 (TRA1) _s_at 885P P P P P 800.9P L-3-hydroxyacyl-Coenzyme A dehydrogenase, short chain (HADHSC) _s_at 398.6P 514.7P P P P 339.4P defender against cell death 1 (DAD1) _at 366.9P 685.1P P 892.6P P 987.1P solute carrier family 7(cationic amino acid transporter, y+ system), member 1 (SLC7A1) _s_at P P P P P 3051P tumor necrosis factor (ligand) superfamily, member 10 (TNFSF10) _at 757.4P P P 935.9P P 922.4P fibroblast growth factor receptor 3 (achondroplasia, thanatophoric dwarfism) (FGFR3) _s_at P P P P P P solute carrier family 31 (copper transporters), member 2 (SLC31A2) _at 245.3P 468.2P P 535.2P P 345.4P regulator of cytokinesis 1 (PRC1) _s_at 182.5P 262.6P P 413.6P P 377P hypothetical protein MAC30 (MAC30) _at 315.2P 400.1P P 584.2P P 584.4P MLF1 interacting protein (MLF1IP) _s_at 217.3P 292.8P P 857.1P P 285.3P N-myc downstream regulated (NDRG1) _s_at 723.2P P P P P 1993P procollagen-lysine, 2-oxoglutarate 5-dioxygenase (lysine hydroxylase) 2 (PLOD2) _s_at 166.3P 227.9P P 294.5P P 107P histone acetyltransferase 1 (HAT1) _at 159.7P 273.7P P 385.8P P 252P chitinase 3-like 1 (cartilage glycoprotein-39) (CHI3L1) _s_at 193P 325.1P P 386.9P P 331.4P activated RNA polymerase II transcription cofactor 4 (PC4) _at 208.1P 332.7P P 329.1P P 247.5P chromosome 3 open reading frame 14 (C3orf14) _at 451.1P 771.5P P 976.7P P 935.7P Ras homolog enriched in brain (RHEB) _s_at 517.7P 749P P 811.1P P 403.1P lipase A, lysosomal acid, cholesterol esterase (Wolman disease) (LIPA) _at 432.6P 742.4P P 836.2P P 514.1P FN5 protein (FN5) _s_at 427.5P 534.5P P 773.6P P 490.2P flap structure-specific endonuclease 1 (FEN1) _s_at 176P 282.5P P 453P P 252.6P leukotriene A-4 hydrolase (LTA4H) _s_at 747.1P P P P P 1047P myristoylated alanine-rich protein kinase C substrate (MARCKS, 80K-L) (MACS) _s_at 395.2P 638.2P P 743.1P P 329.5P transmembrane 4 superfamily member 1 (TM4SF1) _s_at 211P 349.8P P 531P P 319.1P PTPL1-associated RhoGAP 1 (PARG1) _at 407.2P 617.7P P 637.3P P 300.9P thymidylate synthetase (TYMS) _at 245.9P 359.2P P 648.5P P 465.9P CDC28 protein kinase 2 (CKS2) _s_at 120.4P 191.3P P 311.4P P 187.4P similar to lysosome-associated membrane glycoprotein (TSC403) _at 530.6P 838.6P P 771.2P P 559.9P nucleophosmin (nucleolar phosphoprotein B23, numatrin) (NPM1) _s_at P P P P P P influenza virus NS1A binding protein (IVNS1ABP) _s_at 144.3P 209.7P P 268.9P P 245.6P high-mobility group nucleosome binding domain 1 (HMGN1) _s_at P P P P P P proliferating cell nuclear antigen (PCNA) _at 415.1P 601.3P P 852.5P P 383.6P glyoxalase I (GLO1) _at 624.6P 945.3P P P P P von Hippel-Lindau binding protein 1 (VBP1) _at 668.3P P P P P 753.9P ATP synthase, H+ transporting, mitochondrial F0 complex, subunit b, isoform 1 (ATP5F1) _s_at P P P P P P tumor necrosis factor, alpha-induced protein 1 (endothelial) (TNFAIP1) _at 411P 474.8P P 752.1P P 670.7P profilin 2 (PFN2) _s_at P P P P P 899.9P chromosome 7 open reading frame 24 (C7orf24) _s_at 763.9P P P P P P thymopoietin (TMPO) _at 72.2P 83.2P P 183.6P P 86.8P basic transcription factor (BTF3) _x_at P P P P P P syndecan 2 (heparan sulfate proteoglycan 1, cell surface-associated, fibroglycan) (SDC2) _at 225P 312.8P P 359P P 279.1P tubulin, alpha3 tubrin, alpha, ubiquitous (TUBA3 K-ALPHA-1) _x_at P P P P P P brain acid-soluble protein 1 (BASP1) _at P P P P P P Numbers indicate signal intensities detected for each probe sets P: "present" call, detected as expressed, A: "Absent" call, detected as not expressed. Bold: Up-regulated in both MAS 5.0 and dchip analysis Underline: "membrane" protein annotated by GOA analysis
12 Supplemental table 3. Genes detected as up-regulated in human bulge by dchip1.3 software Gene Probe set Mouse bulge Sub-bulge 1 Sub-bulge 2 Sub-bulge 3 Baseline mean Bulge 1 Bulge 2 Bulge 3 Experiment mean Fold change Paired p-value vasoactive intestinal peptide (VIP) _at trinucleotide repeat containing 9 (TNRC9) _x_at trinucleotide repeat containing 9 (TNRC9) _x_at deiodinase, iodothyronine, type II _s_at trinucleotide repeat containing 9 (TNRC9) _x_at pleckstrin homology-like domain, family A, member 1 (PHLDA1) _at Forkhead box C1 (FOXC1) _at angiopoietin-like 2 (ANGPTL2) _at WNT inhibitory factor 1 (WIF1) _at *** nebulette (NEBL) _at regulated in glioma (RIG) _s_at dickkopf homolog 3 (Xenopus laevis) (DKK3) _s_at *,** hypothetical protein FLJ _at aquaporin 3(AQP3) 39248_at calcium/calmodulin-dependent protein kinase ll (CaMKllNalpha) _at dihydropyrimidinase-like 2 (DPYSL2) _at pleckstrin homology-like domain, family A, member 1 (PHLDA1) _at glycoprotein M6B (GPM6B) _at dihydropyrimidinase-like 3 (DPYSL3) _s_at decorin (DCN) _x_at serine (or cysteine) proteinase inhibitor, clade F, member 1 (SERPINF1) _at transducin-like enhancer of split 4 (E(sp1) homolog, Drosophila) (TLE4) _at Ca2+-dependent activator protein for secretion 2 (CADPS2) _at transforming growth factor, beta 2 (TGFB2) _s_at * nebulette (NEBL) _s_at keratin 15 (KRT15) _at ** adrenergic, alpha-2a-, receptor (ADRA2A) _at chromosome 14 open reading frame 78 (C14orf78) _at PTPRF interacting protein, binding protein 1 (liprin beta 1) _at dystrophin (muscular dystrophy, Duchenne and Becker types) (DMD) _at ** coagulation factor II (thrombin) receptor-like 1 (F2RL1) _at iduronate 2-sulfatase (Hunter syndrome) (IDS) _at catenin (cadherin-associated protein), delta 2 (neural plakophilin-related arm-repeat protein) (CTNND2) _s_at fibulin 1 (FBLN1) _s_at * peptidylglycine alpha-amidating monooxygenase (PAM) _x_at microphthalmia-associated transcription factor (MITF) _s_at glycoprotein M6B (GPM6B) _s_at frizzled homolog 1 (Drosophila) (FZD1) _at *,**(FZD2) cdna clone IMAGE: , partial cds _at dual specificity phosphatase 3 (vaccinia virus phosphatase VH1-related) (DUSP3) _at pyridoxal (pyridoxine, vitaminb6) kinase (PDXK) _s_at SH3 domain binding glutamic acid-rich protein like 3 (SH3BGRL3) _s_at neuroblastoma, suppression of tumorigenicity 1 (NBL1) 37005_at bone morphogenetic protein 5 (BMP5) _at nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 1 (NFATC1) _s_at *,** inositol 1,4,5-triphosphate receptor, type 3 (ITPR3) _s_at follistatin (FST) _s_at *,**(Fstl) erythrocyte membrane protein band 4.1-like 2 (EPB41L2) _s_at chromosome 15 open reading frame 25 (C15prf25) _x_at myeloid/lymphoid or mixed-lineage leukemia (trithorax homolog, Drosophila); translocated to, 3 (MLLT3) _s_at glycoprotein M6B (GPM6B) _at hypothetical protein FLJ21075 (FLJ21075) _at angiopoietin-like 2 (ANGPTL2) _at rap2 interacting protein X (RIPX) _s_at KH-type splicing regulatory protein (FUSE binding protein 2) (KHSRP) _s_at interferon regulatory factor 4 (IRF4) _at solute carrier family 12 (potassium/chloride transporters), member 4 (SLC12A4) _at **(Scl29al) collagen, type V, alpha 1 (COL5A1) _at cytochrome b-5 (CYB5) _at zinc finger protein (ZFD25) _x_at neuregulin 1 (NRG1) _s_at calmodulin 1 (phosphorylase kinase, delta) (CALM1) _at solute carrier family 11 (proton-coupled divalent metal ion transporters), member 1 (SLC11A1) _x_at hexokinase 1 (HK1) _at CD59 antigen p18-20 (CD59) _s_at KIAA0924 protein _at carcinoembryonic antigen gene (CEA) _at WW domain containing oxidoreductase (WWOX) _x_at *,**(Wwp2) transportin (importin 3, karyopherin beta 2b) (TNPO2) _s_at KIAA1128 protein (KIAA1128) _s_at interferon induced transmembrane protein 1(9-27) (IFITM1) _s_at forkhead box O3A _s_at chemokine (C-X-C motif) ligand 11 (CXCL11) _s_at *(Cxcl14) transcription elongation factor A (Sll)-like (TCEAL2) _at dopachrome tautomerase (dopachrome delta-isomerase, tyrosine-related protein 2) (DCT) _at immunoglobulin superfamily, member 4 (IGSF4) _s_at *,** serine (or cysteine) proteinase inhibitor, clade E), member 2 (SERPINE2) _at AE binding protein 1 (AEBP1) _at lethal giant larvae homolog 2 (Drosophila) (LLGL2) _s_at Glycine-, glutamate-thienylcyclohexylpiperidine binding protein (KIAA0562) _s_at adrenergic, beta-1-, receptor (ADRB1) _at CD1A antigen, a polypeptide (CD1A) _at cadherin, EGF LAG seven-pass G-type receptor 1 (flamingo homolog, Drosophila) _at apoptosis inhibitor 5 (API5) _s_at peptide YY, 2 (seminalplasmin) (PYY2) _at Numbers indicate Model Based Expression Indexes (MBEIs) detected for each probe sets Bold: Up-regulated in both MAS 5.0 and dchip analysis Underline: "membrane" protein annotated by GOA analysis Mouse bulge column: asterisks indicate the up-regulation of same or similar genes in mouse bulge transcription profiling data * Tumbar et al. Science, , **Morris et al. Nat Biotechnol, , *** Fuchs et al. Cell , parenthesis; gene of similar function was detected as upregulated.
13 Supplemental table 4. Genes detected as down-regulated in human bulge by dchip1.3 software Gene Probe set Sub-bulge1 Sub-bulge2 Sub-bulge3 Baseline mean Bulge 1 Bulge 2 Bulge 3 Experiment mean Fold change Paired p- value cartilage oligomeric matrix protein (pseudoachondroplasia, epiphyseal dysplasia 1, multiple) (COMP) _s_at cytokine receptor-like factor 1 (CRLF1) _at angiopoietin-like 7 (ANGPTL7) _at tripartite motif-containing 9 (TRIM9) _at aminolevulinate, delta-, dehydratase (ALAD) _at protocadherin 8 (PCDH8) _at collagen, type X, alpha 1 (Schmid metaphyseal chondrodysplasia) (COL10A1) _s_at solute carrier family 27 (fatty acid transporter), member 6 (SLC27A6) _at tissue inhibitor of metalloproteinase 3 (Sorsby fundus dystrophy, pseudoinflammatory) (TIMP3) _s_at nucleolar and spindle associated protein 1 (NUSAP1) _at serine (or cysteine) proteinase inhibitor, clade B (ovalbumin), member 13 (SERPINB13) _s_at platelet derived growth factor C (PDGFC) _at solute carrier family 4, sodium bicarbonate cotransporter, member 7 (SLC4A7) _s_at tissue inhibitor of metalloproteinase 3 (Sorsby fundus dystrophy, pseudoinflammatory) (TIMP3) _s_at chitinase 3-like 1 (cartilage glycoprotein-39) (CHI3L1) _at chitinase 3-like 1 (cartilage glycoprotein-39) (CHI3L1) _s_at laminin, beta 1 (LAMB1) _at ets variant gene 5 (ets-related molecule) (ETV5) _s_at cell division cycle 2, G1 to S and G2 to M (CDC2) _at phospholipase A2, group IVA (cytosolic, calcium-dependent) (PLA2G4A) _at kinesin family member 20A (KIF20A) _at GRB2-associated binding protein 2 (GAB2) _s_at protein regulator of cytokinesis 1 (PRC1) _s_at glypican 4 (GPC4) _at ribonucleotide reductase M2 polypeptide (RRM2) _at insulin-like 3 (Leydig cell) (INSL3) _at vav 3 oncogene (VAV3) _at ZW10 interactor (ZWINT) _s_at phosphorylase, glycogen; liver (Hers disease, glycogen storage disease type VI) (PYGL) _at syndecan 2 (heparan sulfate proteoglycan 1, cell surface-associated, fibroglycan) (SDC2) _at syndecan 2 (heparan sulfate proteoglycan 1, cell surface-associated, fibroglycan) (SDC2) _at karyopherin alpha 2 (RAG cohort 1, importin alpha 1) (KPNA2) _s_at protein phosphatase 1, regulatory (inhibitor) subunit 3C (PPP1R3C) _at MCM6 minichromosome maintenance deficient 6 (MIS5 homolog, S. pombe) (S. cerevisiae) (MCM6) _at flap structure-specific endonuclease 1 (FEN1) _s_at retinoic acid induced 14 (RAI14) _s_at topoisomerase (DNA) II alpha 170kDa (TOP2A) _at CS box-containing WD protein _at solute carrier family 1 (glutamate/neutral amino acid transporter), member 4 (SLC1A4) _s_at WAP four-disulfide core domain 1 (WFDC1) _at a disintegrin-like and metalloprotease with thrombospondin type 1 motif, 5 (aggrecanase-2) (ADAMTS5) _at homeodomain-only protein (HOP) _s_at chromosome 22 open reading frame 18 (C22orf18) _at thymidine kinase 1, soluble (TK1) _at fibroblast growth factor 18 (FGF18) _s_at topoisomerase (DNA) II alpha 170kDa (TOP2A) _s_at ribosomal protein L39-like (RPL39L) _at thymidylate synthetase (TYMS) _at Rac GTPase activating protein 1 (RACGAP1) _s_at putative lymphocyte G0/G1 switch gene (GOS2) _s_at collagen, type XI, alpha 1 (COL11A1) 37892_at lactamase, beta 2 (LACTB2) _at melanoma cell adhesion molecule (MCAM, CD146) _x_at chloride channel, calcium activated, family member 2 (CLCA2) _s_at latent transforming growth factor beta binding protein 2 (LTBP2) _at fibroblast growth factor 18 (FGF18) _s_at MCM2 minichromosome maintenance deficient 2, mitotin (S. cerevisiae)(mcm2) _s_at phosphodiesterase 4D interacting protein (myomegalin) (PDE4DIP) _at down-regulator of transcription 1, TBP-binding (negative cofactor 2) (DR1) _s_at ADP-ribosylation factor-like 6 interacting protein (ARL6IP) _at sorting nexin 7 (SNX7) _s_at G protein-coupled receptor 87 (GPR87) _s_at jagged 1 (Alagille syndrome) (JAG1) _s_at N-myc downstream regulated gene 1(NDRG1) _s_at adaptor-related protein complex 1, sigma 2 subunit (AP1S2) _s_at similar to Transcription factor BTF3 homolog 3 (LOC132556) _at thymosin, beta, identified in neuroblastoma cells (TMSNB) _s_at polo-like kinase (Drosophila) (PLK2) _at hairy and enhancer of split 1, (Drosophila) (HES1) _s_at neuromedin U (NMU) _at H2A histone family, member V (H2AFV) _s_at ubiquitin specific protease 4 (proto-oncogene) (USP4) _s_at carbonic anhydrase II (CA2) _at Hypothetical protein LOC (LOC339692) _at F-box protein 28 (FBXO28) _s_at core-binding factor, beta subunit (CBFB) _s_at von Hippel-Lindau binding protein 1 (VBP1) _at DEK oncogene (DNA binding) (DEK) _at squamous cell carcinoma antigen recognized by T cell (SART2) _at ubiquitin-conjugating enzyme E2A (RAD6 homolog) (UBE2A) _s_at karyopherin alpha 2 (RAG cohort 1, importin alpha 1) (KPNA2) _at endothelin receptor type B (EDNRB) _s_at FN5 protein (FN5) _s_at Ras homolog enriched in brain (RHEB) _s_at neurofibromin 1 (neurofibromatosis, von Recklinghausen disease, Watson disease) (NF1) _at B-cell CLL/lymphoma 10 (BCL10) _at nuclear receptor subfamily 4, group A, member 2 (NR4A2) _x_at NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 6, 17kDa (NDUFB6) _s_at Ras and Rab interactor 3 (RIN3) _at solute carrier family 7(cationic amino acid transporter, y+ system), member 1 (SLC7A1) _s_at down-regulator of transcription 1, TBP-binding (negative cofactor 2) (DR1) _x_at SH3-domain GRB2-like endophilin B1 (SH3GLB1) _s_at annexin A2 pseudogene 2 (ANXA2P2) _x_at microfibrillar-associated protein 2 (MFAP2) _at myeloid cell leukemia sequence 1 (BCL2-related) (MCL1) _x_at diacylglycerol kinase, theta 110kDa _at chromosome 9 open reading frame 76 (C9orf76) _at tripartite motif-containing 2 (TRIM2) _s_at adenosine kinase (ADK) _s_at CDC28 protein kinase regulatory subunit 1B (CKS1B) _s_at glycogenin 2 (GYG2) _s_at dual specificity phosphatase 6 (DuSP6) _at isopentenyl-diphosphate delta isomerase (IDI1) _x_at Rho GTPase activating protein 22 (ARHGAP22) _at mitogen-activated protein kinase 10 (MAPK10) _at ubiquitin carboxyl-terminal esterase L3 (ubiquitin thiolesterase) (UCHL3) _at peptidylprolyl isomerase D (cyclophilin D) (PPID) _x_at myotubularin related protein 2 (MTMR2) _s_at thioredoxin domain containing 9 (TXNDC9) _x_at ESTs, Moderately similar to RL39_HUMAN 60S ribosomal protein L39 [H.sapiens] _at gamma-glutamyl carboxylase (GGCX) _at DC13 protein _at Numbers indicate Model Based Expression Indexes (MBEIs) detected for each probe sets Bold: Up-regulated in both MAS 5.0 and dchip analysis Underline: "membrane" protein annotated by GOA analysis
Supplemental Table 1. Oligonucleotides used to identify H-RAS, K-RAS and N-RAS mutations and PTEN gene expression.
 Supplemental Table 1. Oligonucleotides used to identify H-RAS, K-RAS and N-RAS mutations and PTEN gene expression. Gene Exon Forward primer Reverse primer Temp ( C) HRAS 2-3 ATGACGGAATATAAGCTGGT ATGGCAAACACACACAGGAA
Supplemental Table 1. Oligonucleotides used to identify H-RAS, K-RAS and N-RAS mutations and PTEN gene expression. Gene Exon Forward primer Reverse primer Temp ( C) HRAS 2-3 ATGACGGAATATAAGCTGGT ATGGCAAACACACACAGGAA
Ο ΡΟΛΟΣ ΤΩΝ ΚΑΝΝΑΒΙΝΟΕΙ ΩΝ ΚΑΤΑ ΤΗΝ ΕΜΒΡΥΪΚΗ ΑΝΑΠΤΥΞΗ
 ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΥΓΕΙΑΣ ΤΜΗΜΑ ΙΑΤΡΙΚΗΣ Πρόγραµµα Μεταπτυχιακών Σπουδών Εφαρµογές στις Βασικές Ιατρικές Επιστήµες Εργαστήριο Ανατοµικής-Ιστολογίας-Εµβρυολογίας ΙΠΛΩΜΑΤΙΚΗ ΕΡΓΑΣΙΑ Ο ΡΟΛΟΣ ΤΩΝ ΚΑΝΝΑΒΙΝΟΕΙ
ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΥΓΕΙΑΣ ΤΜΗΜΑ ΙΑΤΡΙΚΗΣ Πρόγραµµα Μεταπτυχιακών Σπουδών Εφαρµογές στις Βασικές Ιατρικές Επιστήµες Εργαστήριο Ανατοµικής-Ιστολογίας-Εµβρυολογίας ΙΠΛΩΜΑΤΙΚΗ ΕΡΓΑΣΙΑ Ο ΡΟΛΟΣ ΤΩΝ ΚΑΝΝΑΒΙΝΟΕΙ
Η ΦΛΕΓΜΟΝΩ ΗΣ ΑΝΤΙ ΡΑΣΗ ΤΟΥ ΓΑΣΤΡΙΚΟΥ ΒΛΕΝΝΟΓΟΝΟΥ ΣΤΗ ΛΟΙΜΩΞΗ ΜΕ ΕΛΙΚΟΒΑΚΤΗΡΙ ΙΟ ΤΟΥ ΠΥΛΩΡΟΥ ΠΡΙΝ ΚΑΙ ΜΕΤΑ ΤΗ ΘΕΡΑΠΕΙΑ
 ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΥΓΕΙΑΣ ΤΜΗΜΑ ΙΑΤΡΙΚΗΣ ΤΟΜΕΑΣ ΥΓΕΙΑΣ ΕΡΓΑΣΤΗΡΙΑΚΟΣ ΙΕΥΘΥΝΤΗΣ: Ο ΚΑΘΗΓΗΤΗΣ ΓΕΩΡΓΙΟΣ ΚΑΡΚΑΒΕΛΑΣ ΠΑΝΕΠ. ΕΤΟΣ 2008-2009 Αριθµ. 2084 Η ΦΛΕΓΜΟΝΩ ΗΣ ΑΝΤΙ
ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΥΓΕΙΑΣ ΤΜΗΜΑ ΙΑΤΡΙΚΗΣ ΤΟΜΕΑΣ ΥΓΕΙΑΣ ΕΡΓΑΣΤΗΡΙΑΚΟΣ ΙΕΥΘΥΝΤΗΣ: Ο ΚΑΘΗΓΗΤΗΣ ΓΕΩΡΓΙΟΣ ΚΑΡΚΑΒΕΛΑΣ ΠΑΝΕΠ. ΕΤΟΣ 2008-2009 Αριθµ. 2084 Η ΦΛΕΓΜΟΝΩ ΗΣ ΑΝΤΙ
SABiosciences PCR Array Catalog #: PAHS-021 SA+ SCF 4h stimulaton AVG(Ct) Position Unigene Refseq Symbol Description shcontr shcontr-4h shgskβ A01
 SABiosciences PCR Array Catalog #: PAHS-021 SA+ SCF 4h stimulaton AVG(Ct) Position Unigene Refseq Symbol Description shcontr shcontr-4h shgskβ A01 Hs.1274 NM_006129 BMP1 Bone morphogenetic protein 1 25.25
SABiosciences PCR Array Catalog #: PAHS-021 SA+ SCF 4h stimulaton AVG(Ct) Position Unigene Refseq Symbol Description shcontr shcontr-4h shgskβ A01 Hs.1274 NM_006129 BMP1 Bone morphogenetic protein 1 25.25
Table S2. Sequences immunoprecipitating with anti-hif-1α
 Genome-wide analysis of HIF DNA-bdg-supplemental material Table S2. Sequences immunoprecipitatg with anti-α The table gives the chromosomal co-ordates of each anti-α immunoprecipitated locus meetg criteria
Genome-wide analysis of HIF DNA-bdg-supplemental material Table S2. Sequences immunoprecipitatg with anti-α The table gives the chromosomal co-ordates of each anti-α immunoprecipitated locus meetg criteria
Debashish Sahay. To cite this version: HAL Id: tel
 Identification of genes activated and biological markers involved in lysophosphatidic acid (LPA)-induced breast cancer metastasis through its receptor LPA1 Debashish Sahay To cite this version: Debashish
Identification of genes activated and biological markers involved in lysophosphatidic acid (LPA)-induced breast cancer metastasis through its receptor LPA1 Debashish Sahay To cite this version: Debashish
Κυτταρική Επικοινωνία Cell Communication
 Κυτταρική Επικοινωνία Cell Communication Η μοίρα του κυττάρου ρυθμίζεται από εξωτερικά σήματα (από το περιβάλλον και άλλα κύτταρα) survival division differentiation apoptosis Είδη κυτταρικής επικοινωνίας
Κυτταρική Επικοινωνία Cell Communication Η μοίρα του κυττάρου ρυθμίζεται από εξωτερικά σήματα (από το περιβάλλον και άλλα κύτταρα) survival division differentiation apoptosis Είδη κυτταρικής επικοινωνίας
Μαρία Κατσιφοδήμου. Ο ρόλος της έκκρισης HLA-G από τα ανθρώπινα έμβρυα στην επιτυχία της εξωσωματικής γονιμοποίησης. Μεταπτυχιακή Διπλωματική Εργασία
 ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΣΧΟΛΗ ΘΕΤΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΤΜΗΜΑ ΒΙΟΛΟΓΙΑΣ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ ΚΑΤΕΥΘΥΝΣΗ: ΕΦΑΡΜΟΣΜΕΝΗ ΓΕΝΕΤΙΚΗ ΚΑΙ ΒΙΟΤΕΧΝΟΛΟΓΙΑ Ο ρόλος της έκκρισης HLA-G από τα ανθρώπινα
ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΣΧΟΛΗ ΘΕΤΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΤΜΗΜΑ ΒΙΟΛΟΓΙΑΣ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ ΚΑΤΕΥΘΥΝΣΗ: ΕΦΑΡΜΟΣΜΕΝΗ ΓΕΝΕΤΙΚΗ ΚΑΙ ΒΙΟΤΕΧΝΟΛΟΓΙΑ Ο ρόλος της έκκρισης HLA-G από τα ανθρώπινα
Hvordan påvirker CO 2 post-smolt i brakkvann RAS?
 Hvordan påvirker CO 2 post-smolt i brakkvann RAS? Tom O. Nilsen,. Mota, V.C., Kolarevic, J., Ytterborg, E., Ebbesson, Lars O.E., Handeland, S.O., Stiller, K., Terjesen, B.F SUNNDALSØRA, 12. NOVEMBER 2018
Hvordan påvirker CO 2 post-smolt i brakkvann RAS? Tom O. Nilsen,. Mota, V.C., Kolarevic, J., Ytterborg, E., Ebbesson, Lars O.E., Handeland, S.O., Stiller, K., Terjesen, B.F SUNNDALSØRA, 12. NOVEMBER 2018
High mobility group 1 HMG1
 Vol. 29, pp.705 ~ 711, 2001 High mobility group 1 HMG1 13 12 20 anti-neutrophil cytoplasmic antibodies, ANCA ANCA 1982 Davies 1980 1 high mobility group HMG1 HMG2 30 kd high mobility group HMGHMG HMG1
Vol. 29, pp.705 ~ 711, 2001 High mobility group 1 HMG1 13 12 20 anti-neutrophil cytoplasmic antibodies, ANCA ANCA 1982 Davies 1980 1 high mobility group HMG1 HMG2 30 kd high mobility group HMGHMG HMG1
Μελέτη της έκφρασης του ογκοκατασταλτικού γονιδίου Cyld στον καρκίνο του μαστού
 Σχολή Θετικών Επιστημών Τμήμα Βιολογίας Πρόγραμμα Μεταπτυχιακών Σπουδών Κατεύθυνση: Εφαρμοσμένη γενετική και βιοτεχνολογία ΜΕΤΑΠΤΥΧΙΑΚΗ ΔΙΠΛΩΜΑΤΙΚΗ ΕΡΓΑΣΙΑ Μελέτη της έκφρασης του ογκοκατασταλτικού γονιδίου
Σχολή Θετικών Επιστημών Τμήμα Βιολογίας Πρόγραμμα Μεταπτυχιακών Σπουδών Κατεύθυνση: Εφαρμοσμένη γενετική και βιοτεχνολογία ΜΕΤΑΠΤΥΧΙΑΚΗ ΔΙΠΛΩΜΑΤΙΚΗ ΕΡΓΑΣΙΑ Μελέτη της έκφρασης του ογκοκατασταλτικού γονιδίου
ΠΡΟΣΚΛΗΣΗ ΕΚΔΗΛΩΣΗΣ ΕΝΔΙΑΦΕΡΟΝΤΟΣ
 1 ΕΛΛΗΝΙΚΗ ΔΗΜΟΚΡΑΤΙΑ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΡΗΤΗΣ ΕΙΔΙΚΟΣ ΛΟΓΑΡΙΑΣΜΟΣ ΑΝΑΡΤΗΤΕΑ ΣΤΟ ΔΙΑΔΙΚΤΥΟ ΠΡΟΣΚΛΗΣΗ ΕΚΔΗΛΩΣΗΣ ΕΝΔΙΑΦΕΡΟΝΤΟΣ Ηράκλειο, 04.02.2014 Αρ. πρωτ. 1054 Ο Ειδικός Λογαριασμός του Πανεπιστημίου Κρήτης
1 ΕΛΛΗΝΙΚΗ ΔΗΜΟΚΡΑΤΙΑ ΠΑΝΕΠΙΣΤΗΜΙΟ ΚΡΗΤΗΣ ΕΙΔΙΚΟΣ ΛΟΓΑΡΙΑΣΜΟΣ ΑΝΑΡΤΗΤΕΑ ΣΤΟ ΔΙΑΔΙΚΤΥΟ ΠΡΟΣΚΛΗΣΗ ΕΚΔΗΛΩΣΗΣ ΕΝΔΙΑΦΕΡΟΝΤΟΣ Ηράκλειο, 04.02.2014 Αρ. πρωτ. 1054 Ο Ειδικός Λογαριασμός του Πανεπιστημίου Κρήτης
ΙΩΑΝΝΗΣ ΚΛΑΓΚΑΣ. Χημικός, MSc ΔΙΔΑΚΤΟΡΙΚΗ ΔΙΑΤΡΙΒΗ ΥΠΟΒΛΗΘΗΚΕ ΣΤΗΝ ΙΑΤΡΙΚΗ ΣΧΟΛΗ ΤΟΥ ΑΡΙΣΤΟΤΕΛΕΙΟΥ ΠΑΝΕΠΙΣΤΗΜΙΟΥ ΘΕΣΣΑΛΟΝΙΚΗΣ
 ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΙΑΤΡΙΚΗ ΣΧΟΛΗ ΤΟΜΕΑΣ ΦΥΣΙΟΛΟΓΙΑΣ ΦΑΡΜΑΚΟΛΟΓΙΑΣ B ΕΡΓΑΣΤΗΡΙΟ ΦΑΡΜΑΚΟΛΟΓΙΑΣ ΔΙΕΥΘΥΝΤΗΣ: ΚΑΘΗΓΗΤΗΣ ΓΙΩΡΓΟΣ ΚΑΡΑΚΙΟΥΛΑΚΗΣ ΠΑΝΕΠ. ΕΤΟΣ 2008-2009 ΑΡΙΘΜ. 2376 Η ΕΠΙΔΡΑΣΗ
ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΙΑΤΡΙΚΗ ΣΧΟΛΗ ΤΟΜΕΑΣ ΦΥΣΙΟΛΟΓΙΑΣ ΦΑΡΜΑΚΟΛΟΓΙΑΣ B ΕΡΓΑΣΤΗΡΙΟ ΦΑΡΜΑΚΟΛΟΓΙΑΣ ΔΙΕΥΘΥΝΤΗΣ: ΚΑΘΗΓΗΤΗΣ ΓΙΩΡΓΟΣ ΚΑΡΑΚΙΟΥΛΑΚΗΣ ΠΑΝΕΠ. ΕΤΟΣ 2008-2009 ΑΡΙΘΜ. 2376 Η ΕΠΙΔΡΑΣΗ
Supplemental Figure 1. Pathways associated with the most significantly altered FGF-8b-regulated
 Supplemental Figure legends Supplemental Figure 1. Pathways associated with the most significantly altered FGF-8b-regulated genes. a) Mitotic roles of Polo-like kinase. b) Cell cycle regulation by BTG
Supplemental Figure legends Supplemental Figure 1. Pathways associated with the most significantly altered FGF-8b-regulated genes. a) Mitotic roles of Polo-like kinase. b) Cell cycle regulation by BTG
ΥΠΕΡΙΝΣΟΥΛΙΝΑΙΜΙΑ ΚΑΙ ΑΝΤΙΣΤΑΣΗ ΣΤΗΝ ΙΝΣΟΥΛΙΝΗ ΣΕ ΠΑΧΥΣΑΡΚΑ ΠΑΙΔΙΑ ΚΑΙ ΕΦΗΒΟΥΣ ΜΕ ΠΡΩΙΜΗ ΑΔΡΕΝΑΡΧΗ
 ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΙΑΤΡΙΚΗ ΣΧΟΛΗ ΤΟΜΕΑΣ ΒΙΟΛΟΓΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΚΑΙ ΠΡΟΛΗΠΤΙΚΗΣ ΙΑΤΡΙΚΗΣ ΕΡΓΑΣΤΗΡΙΟ ΒΙΟΧΗΜΕΙΑΣ ΔΙΕΥΘΥΝΤΡΙΑ: Η ΚΑΘΗΓΗΤΡΙΑ Ν. ΒΑΒΑΤΣΗ-ΧΡΙΣΤΑΚΗ ΠΑΝΕΠ. ΕΤΟΣ 2009-2010 Αριθμ.2456
ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΙΑΤΡΙΚΗ ΣΧΟΛΗ ΤΟΜΕΑΣ ΒΙΟΛΟΓΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΚΑΙ ΠΡΟΛΗΠΤΙΚΗΣ ΙΑΤΡΙΚΗΣ ΕΡΓΑΣΤΗΡΙΟ ΒΙΟΧΗΜΕΙΑΣ ΔΙΕΥΘΥΝΤΡΙΑ: Η ΚΑΘΗΓΗΤΡΙΑ Ν. ΒΑΒΑΤΣΗ-ΧΡΙΣΤΑΚΗ ΠΑΝΕΠ. ΕΤΟΣ 2009-2010 Αριθμ.2456
Table S1A. Generic EMT signature for tumour
 Table S1A. Generic EMT signature for tumour Index Gene Symbol Gene Title GO Molecular Function Weight Weight.Zt ransform ed Category In Generic Cell Line EMT Signature = 1 1 KRT19 Keratin, type I cytoskeletal
Table S1A. Generic EMT signature for tumour Index Gene Symbol Gene Title GO Molecular Function Weight Weight.Zt ransform ed Category In Generic Cell Line EMT Signature = 1 1 KRT19 Keratin, type I cytoskeletal
Κυτταρική Επικοινωνία Cell Communication
 Κυτταρική Επικοινωνία Cell Communication Κυτταρική Επικοινωνία Είναι η ικανότητα του κυττάρου να αποκρίνεται σε εξωτερικά σήματα (σήματα από το περιβάλλον του ή άλλα κύτταρα) Τα σήματα αυτά μπορεί να είναι
Κυτταρική Επικοινωνία Cell Communication Κυτταρική Επικοινωνία Είναι η ικανότητα του κυττάρου να αποκρίνεται σε εξωτερικά σήματα (σήματα από το περιβάλλον του ή άλλα κύτταρα) Τα σήματα αυτά μπορεί να είναι
NES: normalized enrichment score (analyzed using KEGG pathway gene sets in the GSEA software); FDR:
 Supplementary table1. List of gene sets with simultaneously altered the enrichment upon SAP18 overexpression and knockdown. Gene sets enriched by SAP18 and reduced by shsap18 SAP18 against LacZ shsap18
Supplementary table1. List of gene sets with simultaneously altered the enrichment upon SAP18 overexpression and knockdown. Gene sets enriched by SAP18 and reduced by shsap18 SAP18 against LacZ shsap18
Supplemental Figures and Tables
 KLF2 and KLF4 control endothelial identity and vascular integrity Panjamaporn Sangwung 1, 2,#, Guangjin Zhou 1,#, Lalitha Nayak 1,3, E. Ricky Chan 4, Sandeep Kumar 5, Dong-Won Kang 5, Rongli Zhang 1, Xudong
KLF2 and KLF4 control endothelial identity and vascular integrity Panjamaporn Sangwung 1, 2,#, Guangjin Zhou 1,#, Lalitha Nayak 1,3, E. Ricky Chan 4, Sandeep Kumar 5, Dong-Won Kang 5, Rongli Zhang 1, Xudong
Overlapping sets of transcripts from host and non-host interactions of tomato are expressed early during non-host resistance
 POJ 7(1):19-27 (2014) ISSN:1836-3644 Overlapping sets of transcripts from host and non-host interactions of tomato are expressed early during non-host resistance Battepati Uma and Appa Rao Podile* Supplementary
POJ 7(1):19-27 (2014) ISSN:1836-3644 Overlapping sets of transcripts from host and non-host interactions of tomato are expressed early during non-host resistance Battepati Uma and Appa Rao Podile* Supplementary
ΜΟΡΙΑΚΕΣ ΜΕΘΟΔΟΙ ΚΡΙΤΗΡΙΑ ΕΠΙΛΟΓΗΣ ΑΞΙΟΛΟΓΗΣΗ
 - - ό ό : Ω ί ό ί ώ.. ά ή ί ός ή ς ύ ί ί ή έ ώ ό. ή ς ά ς ής ό ός ά ς ή έ ός ή ς ί ής ί ς. ό έ ώ - ώ ή ής ή ς- ί ά ά ς ί ς. ά ί ί έ έ ά ύ ή, ό ί ά, ό ό ά έ ά ής ί ύ George Wald ή ί έ ς ί ύ ό ς ί ς ά έ
- - ό ό : Ω ί ό ί ώ.. ά ή ί ός ή ς ύ ί ί ή έ ώ ό. ή ς ά ς ής ό ός ά ς ή έ ός ή ς ί ής ί ς. ό έ ώ - ώ ή ής ή ς- ί ά ά ς ί ς. ά ί ί έ έ ά ύ ή, ό ί ά, ό ό ά έ ά ής ί ύ George Wald ή ί έ ς ί ύ ό ς ί ς ά έ
Legends for Supplemental Data
 Legends for Supplemental Data Supplemental Fig. 1. Top molecular and cellular functions identified by Ingenuity Pathway Analysis after 6 h (left) and 24 h (right) of exposure of mouse pancreatic islets
Legends for Supplemental Data Supplemental Fig. 1. Top molecular and cellular functions identified by Ingenuity Pathway Analysis after 6 h (left) and 24 h (right) of exposure of mouse pancreatic islets
SUPPLEMENTAL MATERIAL
 SUPPLEMENTAL MATERIAL 1. Methods MicroRNA TaqMan Low Density Array MiRNAs were reverse transcribed and amplified using the multiplex RT TaqMan MicroRNA Low Density Array (TLDA) (Life Technologies). Global
SUPPLEMENTAL MATERIAL 1. Methods MicroRNA TaqMan Low Density Array MiRNAs were reverse transcribed and amplified using the multiplex RT TaqMan MicroRNA Low Density Array (TLDA) (Life Technologies). Global
Data-Driven Discovery of Extravasation Pathway in Circulating Tumor Cells
 Data-Driven Discovery of Extravasation Pathway in Circulating Tumor Cells Yadavalli S. 1, Jayaram S. 1,2, Manda SS. 1,3, Madugundu AK. 1,3, Nayakanti DS. 1, Tuan TZ. 6, Bhat R. 4, Rangarajan A. 4, Chatterjee
Data-Driven Discovery of Extravasation Pathway in Circulating Tumor Cells Yadavalli S. 1, Jayaram S. 1,2, Manda SS. 1,3, Madugundu AK. 1,3, Nayakanti DS. 1, Tuan TZ. 6, Bhat R. 4, Rangarajan A. 4, Chatterjee
ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΙΑΤΡΙΚΗ ΣΧΟΛΗ. ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥ ΩΝ «Ιατρική Ερευνητική Μεθοδολογία» ΙΠΛΩΜΑΤΙΚΗ ΕΡΓΑΣΙΑ
 ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΙΑΤΡΙΚΗ ΣΧΟΛΗ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥ ΩΝ «Ιατρική Ερευνητική Μεθοδολογία» ΙΠΛΩΜΑΤΙΚΗ ΕΡΓΑΣΙΑ Νόσος Pompe: Νεότερα κλινικά, γενετικά, διαγνωστικά και θεραπευτικά
ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΙΑΤΡΙΚΗ ΣΧΟΛΗ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥ ΩΝ «Ιατρική Ερευνητική Μεθοδολογία» ΙΠΛΩΜΑΤΙΚΗ ΕΡΓΑΣΙΑ Νόσος Pompe: Νεότερα κλινικά, γενετικά, διαγνωστικά και θεραπευτικά
Hepatic Stellate Cells: Multifunctional mesenchymal cells of the Liver
 Nagoya Med. J., 137 Hepatic Stellate Cells: Multifunctional mesenchymal cells of the Liver KAZUO IKEDA Department of Anatomy and Cell Biology, Graduate School of Medical Sciences, Nagoya City University
Nagoya Med. J., 137 Hepatic Stellate Cells: Multifunctional mesenchymal cells of the Liver KAZUO IKEDA Department of Anatomy and Cell Biology, Graduate School of Medical Sciences, Nagoya City University
CPT. Tsuchiya. beta. quantitative RT PCR QIAGEN IGFBP. Fect Transfection Reagent sirna. RT PCR RNA Affymetrix GeneChip Expression Array
 7 PTEN p Tsuchiya DNA Hepatocyte nuclear factor beta HNF beta HNFbeta IGFBP GLUT G Pase MODY maturity onset diabetes of the young Tsuchiya HNFbeta HNFbeta HNFbeta sirna CPT HNFbeta HNFbeta TOVG KOC ESMCAS
7 PTEN p Tsuchiya DNA Hepatocyte nuclear factor beta HNF beta HNFbeta IGFBP GLUT G Pase MODY maturity onset diabetes of the young Tsuchiya HNFbeta HNFbeta HNFbeta sirna CPT HNFbeta HNFbeta TOVG KOC ESMCAS
Differentially expressed genes of HEK293/HeLa Myst4 sirna vs. controls.
 Supplemental data - Content Supplemental figures Figure 1 Figure 2 Figure 3 Figure 4 Western blot analysis of MYST4. Expression pattern of MYST4 isoforms. Differentially expressed genes of HEK293/HeLa
Supplemental data - Content Supplemental figures Figure 1 Figure 2 Figure 3 Figure 4 Western blot analysis of MYST4. Expression pattern of MYST4 isoforms. Differentially expressed genes of HEK293/HeLa
5- CACGAAACTACCTTCAACTCC-3 beta actin-r 5- CATACTCCTGCTTGCTGATC-3 GAPDH-F GAPDH-R
 Table S1 Primers used in this work LncPPARδ-F 5- AGGATCCCAAGGCAGAAGTT -3 LncPPARδ-R 5- GAATAGAAAGGCGGGAAAGG -3 PPARδ-F 5- CACCCAGTGTCCTTCCATCT -3 PPARδ-R 5- CAGAATTGGGTGTGCTGCTT -3 ADRP-F 5- ACAACCGAGTGTGGTGACTC
Table S1 Primers used in this work LncPPARδ-F 5- AGGATCCCAAGGCAGAAGTT -3 LncPPARδ-R 5- GAATAGAAAGGCGGGAAAGG -3 PPARδ-F 5- CACCCAGTGTCCTTCCATCT -3 PPARδ-R 5- CAGAATTGGGTGTGCTGCTT -3 ADRP-F 5- ACAACCGAGTGTGGTGACTC
Γενετικά νοσήµατα: Μοριακή βάση και διαγνωστική
 Μαθήματα Παθολογικής Ανατομικής: Η ώρα του ειδικευόμενου 13 Aπριλίου 2011 Γενετικά νοσήµατα: Μοριακή βάση και διαγνωστική Ντίνα Τηνιακού Εργ/ριο Ιστολογίας & Εµβρυολογίας Ιατρική Σχολή Πανεπιστηµίου Aθηνών
Μαθήματα Παθολογικής Ανατομικής: Η ώρα του ειδικευόμενου 13 Aπριλίου 2011 Γενετικά νοσήµατα: Μοριακή βάση και διαγνωστική Ντίνα Τηνιακού Εργ/ριο Ιστολογίας & Εµβρυολογίας Ιατρική Σχολή Πανεπιστηµίου Aθηνών
Human angiogenin is a potent cytotoxin in the absence of ribonuclease inhibitor
 Human angiogenin is a potent cytotoxin in the absence of ribonuclease inhibitor SYDNEY P. THOMAS, 1 TRISH T. HOANG, 2 VALERIE T. RESSLER, 4 and RONALD T. RAINES 2,3,4 1 Graduate Program in Cell & Molecular
Human angiogenin is a potent cytotoxin in the absence of ribonuclease inhibitor SYDNEY P. THOMAS, 1 TRISH T. HOANG, 2 VALERIE T. RESSLER, 4 and RONALD T. RAINES 2,3,4 1 Graduate Program in Cell & Molecular
A rearrangement is established as a barrier in parapatry. The accumulation of incompatibilites starts. Chrom 1. Rearranged
 Figure S1. An ancestral species splits in two according to the model discussed in Navarro and Barton (Evolution. In press. 2003). A rearrangement in chromosome 1 was involved in the process. From the moment
Figure S1. An ancestral species splits in two according to the model discussed in Navarro and Barton (Evolution. In press. 2003). A rearrangement in chromosome 1 was involved in the process. From the moment
ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΓΕΩΠΟΝΙΚΗ ΣΧΟΛΗ ΤΟΜΕΑΣ ΕΠΙΣΤΗΜΗΣ ΚΑΙ ΤΕΧΝΟΛΟΓΙΑΣ ΤΡΟΦΙΜΩΝ ΜΑΡΙΑΣ ΦΩΤΙΟΥ ΠΤΥΧΙΟΥΧΟΥ ΓΕΩΠΟΝΟΥ
 ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΓΕΩΠΟΝΙΚΗ ΣΧΟΛΗ ΤΟΜΕΑΣ ΕΠΙΣΤΗΜΗΣ ΚΑΙ ΤΕΧΝΟΛΟΓΙΑΣ ΤΡΟΦΙΜΩΝ ΜΑΡΙΑΣ ΦΩΤΙΟΥ ΠΤΥΧΙΟΥΧΟΥ ΓΕΩΠΟΝΟΥ Συγκέντρωση των ελεύθερων αµινοξέων στο αµνιακό υγρό σε σχέση µε την εβδοµάδα
ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΓΕΩΠΟΝΙΚΗ ΣΧΟΛΗ ΤΟΜΕΑΣ ΕΠΙΣΤΗΜΗΣ ΚΑΙ ΤΕΧΝΟΛΟΓΙΑΣ ΤΡΟΦΙΜΩΝ ΜΑΡΙΑΣ ΦΩΤΙΟΥ ΠΤΥΧΙΟΥΧΟΥ ΓΕΩΠΟΝΟΥ Συγκέντρωση των ελεύθερων αµινοξέων στο αµνιακό υγρό σε σχέση µε την εβδοµάδα
BD Cytometric Bead Array Flex Set System
 BD Cytometric Bead Array Flex Set System BD Cytometric Bead Array System BD Biosciences Pharmingen ELISA Cytometric Bead Array CBA System Capture Caputure Serum Supernatant Lysate PE -Detection 1 2 PE
BD Cytometric Bead Array Flex Set System BD Cytometric Bead Array System BD Biosciences Pharmingen ELISA Cytometric Bead Array CBA System Capture Caputure Serum Supernatant Lysate PE -Detection 1 2 PE
encouraged to use the Version of Record that, when published, will replace this version. The most /BCJ BIOCHEMICAL JOURNAL
 Biochemical Journal: this is an Accepted Manuscript, not the final Version of Record. You are encouraged to use the Version of Record that, when published, will replace this version. The most up-to-date
Biochemical Journal: this is an Accepted Manuscript, not the final Version of Record. You are encouraged to use the Version of Record that, when published, will replace this version. The most up-to-date
ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ ΣΤΙΣ «ΚΛΙΝΙΚΕΣ ΚΑΙ ΚΛΙΝΙΚΟΕΡΓΑΣΤΗΡΙΑΚΕΣ ΙΑΤΡΙΚΕΣ ΕΙΔΙΚΟΤΗΤΕΣ»
 ΠΑΝΕΠΙΣΤΗΜΙΟ ΠΑΤΡΩΝ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΥΓΕΙΑΣ ΤΜΗΜΑ ΙΑΤΡΙΚΗΣ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ ΣΤΙΣ «ΚΛΙΝΙΚΕΣ ΚΑΙ ΚΛΙΝΙΚΟΕΡΓΑΣΤΗΡΙΑΚΕΣ ΙΑΤΡΙΚΕΣ ΕΙΔΙΚΟΤΗΤΕΣ» ΑΡΝΗΤΙΚΗ ΡΥΘΜΙΣΗ ΤΗΣ ΜΕΤΑΒΙΒΑΣΗΣ ΤΟΥ ΣΗΜΑΤΟΣ ΤΗΣ
ΠΑΝΕΠΙΣΤΗΜΙΟ ΠΑΤΡΩΝ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΥΓΕΙΑΣ ΤΜΗΜΑ ΙΑΤΡΙΚΗΣ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ ΣΤΙΣ «ΚΛΙΝΙΚΕΣ ΚΑΙ ΚΛΙΝΙΚΟΕΡΓΑΣΤΗΡΙΑΚΕΣ ΙΑΤΡΙΚΕΣ ΕΙΔΙΚΟΤΗΤΕΣ» ΑΡΝΗΤΙΚΗ ΡΥΘΜΙΣΗ ΤΗΣ ΜΕΤΑΒΙΒΑΣΗΣ ΤΟΥ ΣΗΜΑΤΟΣ ΤΗΣ
< (0.999) Graft (0.698) (0.483) <0.001 (0.698) (<0.001) (<0.001) 3 months (0.999) (0.483) (<0.001) 6 months (<0.
 Supplementary table 1. Correlation of endothelial cell density among graft, 3, 6, and 12 months after Descemet s automated stripping endothelial keratoplasty. Graft 3 months 6 months 12 months Graft
Supplementary table 1. Correlation of endothelial cell density among graft, 3, 6, and 12 months after Descemet s automated stripping endothelial keratoplasty. Graft 3 months 6 months 12 months Graft
Christopher Stephen Inchley, 2015
 Christopher Stephen Inchley, 2015 Series of dissertations submitted to the Faculty of Medicine, University of Oslo No. 2009 ISBN 978-82-8264-990-2 All rights reserved. No part of this publication may be
Christopher Stephen Inchley, 2015 Series of dissertations submitted to the Faculty of Medicine, University of Oslo No. 2009 ISBN 978-82-8264-990-2 All rights reserved. No part of this publication may be
Long intergenic non-coding RNA expression signature in human breast cancer
 Long intergenic non-coding RNA expression signature in human breast cancer Yanfeng Zhang 1,5,#, Erin K. Wagner 2,#, Xingyi Guo 1, Isaac May 3, Qiuyin Cai 1, Wei Zheng 1, Chunyan He 2,4,*, Jirong Long 1,*
Long intergenic non-coding RNA expression signature in human breast cancer Yanfeng Zhang 1,5,#, Erin K. Wagner 2,#, Xingyi Guo 1, Isaac May 3, Qiuyin Cai 1, Wei Zheng 1, Chunyan He 2,4,*, Jirong Long 1,*
Διάλεξη 5. Μπράλιου Γεωργία, PhD, Τμήμα Πληροφορικής με Εφαρμογές στη Βιοϊατρική Πανεπιστήμιο Θεσσαλίας
 Διάλεξη 5 Δομή μεμβρανών (πάχος 5nm, 50 άτομα) Μεμβρανική μεταφορά Φραγμοί Μεμβρανικές πρωτεΐνες Λειτουργίες μεμβρανικών πρωτεϊνών Δίαυλοι-μεταφορείς-δυναμικό της μεμβράνης G protein coupled receptors
Διάλεξη 5 Δομή μεμβρανών (πάχος 5nm, 50 άτομα) Μεμβρανική μεταφορά Φραγμοί Μεμβρανικές πρωτεΐνες Λειτουργίες μεμβρανικών πρωτεϊνών Δίαυλοι-μεταφορείς-δυναμικό της μεμβράνης G protein coupled receptors
Supplementary Information Titles
 Supplementary Information Titles Journal: Nature Neuroscience Article Title: Corresponding Author: Wnt-mediated activation of NeuroD1 and retroelements during Tomoko Kuwabara adult neurogenesis Supplementary
Supplementary Information Titles Journal: Nature Neuroscience Article Title: Corresponding Author: Wnt-mediated activation of NeuroD1 and retroelements during Tomoko Kuwabara adult neurogenesis Supplementary
Supplementary Table 1. Primer sequences used for RT-qPCR validation of the microarray data. Primer. direction. Forward
 Supplementary Table 1. Primer sequences used for RT-qPCR validation of the microarray data Gene Glyceraldehyde 3-phosphate dehydrogenase (Gapdh) Cell death-inducing DFFAlike effector a (Cidea) Peroxisome
Supplementary Table 1. Primer sequences used for RT-qPCR validation of the microarray data Gene Glyceraldehyde 3-phosphate dehydrogenase (Gapdh) Cell death-inducing DFFAlike effector a (Cidea) Peroxisome
TFDP2 TFDP1 SUMO1 SERTAD1 RPA3 RBBP8 RAD9A RAD17 RAD1 NBN MRE11A MNAT1 KPNA2 KNTC1 HUS1 HERC5 GTSE1 GTF2H1 GADD45A E2F4 DNM2 DDX11 CUL3 SKP2 RBL1 RB1
 -8-6 -4-2 CHEK2 MKI67 RB1 RBL1 SKP2 Fold-Change Fold-Change -3-2 -1 CCND1 CCNE1 CDK4 CDK5R1 5 1 15 ATM ATR BAX BCCIP BRCA1 BRCA2 BL1AT ABL1A ABL1 CCNG1 CCNG2 CCNH CCNT1 CCNT2 CDC2 CDC34 CDKN1A CDKN1B CDKN2A
-8-6 -4-2 CHEK2 MKI67 RB1 RBL1 SKP2 Fold-Change Fold-Change -3-2 -1 CCND1 CCNE1 CDK4 CDK5R1 5 1 15 ATM ATR BAX BCCIP BRCA1 BRCA2 BL1AT ABL1A ABL1 CCNG1 CCNG2 CCNH CCNT1 CCNT2 CDC2 CDC34 CDKN1A CDKN1B CDKN2A
The effect of curcumin on the stability of Aβ. dimers
 The effect of curcumin on the stability of Aβ dimers Li Na Zhao, See-Wing Chiu, Jérôme Benoit, Lock Yue Chew,, and Yuguang Mu, School of Physical and Mathematical Sciences, Nanyang Technological University,
The effect of curcumin on the stability of Aβ dimers Li Na Zhao, See-Wing Chiu, Jérôme Benoit, Lock Yue Chew,, and Yuguang Mu, School of Physical and Mathematical Sciences, Nanyang Technological University,
ΚΥΤΤΑΡΙΚΟΣ ΘΑΝΑΤΟΣ ΚΑΙ ΑΝΑΓΕΝΝΗΣΗ Β-ΚΥΤΤΑΡΟΥ ΝΕΟΤΕΡΑ ΔΕΔΟΜΕΝΑ ΓΙΑ ΤΗ ΘΕΡΑΠΕΙΑ ΤΟΥ ΣΑΚΧΑΡΩΔΗ ΔΙΑΒΗΤΗ
 ΚΥΤΤΑΡΙΚΟΣ ΘΑΝΑΤΟΣ ΚΑΙ ΑΝΑΓΕΝΝΗΣΗ Β-ΚΥΤΤΑΡΟΥ ΝΕΟΤΕΡΑ ΔΕΔΟΜΕΝΑ ΓΙΑ ΤΗ ΘΕΡΑΠΕΙΑ ΤΟΥ ΣΑΚΧΑΡΩΔΗ ΔΙΑΒΗΤΗ ΚΑΛΛΙΟΠΗ ΚΩΤΣΑ ΕΠΙΚ. ΚΑΘΗΓΗΤΡΙΑ ΕΝΔΟΚΡΙΝΟΛΟΓΙΑΣ ΙΑΤΡΙΚΗ ΣΧΟΛΗ Α.Π.Θ ΤΜΗΜΑ ΕΝΔΟΚΡΙΝΟΛΟΓΙΑΣ-ΔΙΑΒΗΤΟΛΟΓΙΚΟ
ΚΥΤΤΑΡΙΚΟΣ ΘΑΝΑΤΟΣ ΚΑΙ ΑΝΑΓΕΝΝΗΣΗ Β-ΚΥΤΤΑΡΟΥ ΝΕΟΤΕΡΑ ΔΕΔΟΜΕΝΑ ΓΙΑ ΤΗ ΘΕΡΑΠΕΙΑ ΤΟΥ ΣΑΚΧΑΡΩΔΗ ΔΙΑΒΗΤΗ ΚΑΛΛΙΟΠΗ ΚΩΤΣΑ ΕΠΙΚ. ΚΑΘΗΓΗΤΡΙΑ ΕΝΔΟΚΡΙΝΟΛΟΓΙΑΣ ΙΑΤΡΙΚΗ ΣΧΟΛΗ Α.Π.Θ ΤΜΗΜΑ ΕΝΔΟΚΡΙΝΟΛΟΓΙΑΣ-ΔΙΑΒΗΤΟΛΟΓΙΚΟ
Supplemental Table S1. Tumor specific networks are enriched with somatically mutated genes (taken from the database COSMIC)
 Additional file 1 Supplemental Table S1. Tumor specific networks are enriched with somatically mutated genes (taken from the database COSMIC) COSMIC genes in tumor network COSMIC genes not in tumor network
Additional file 1 Supplemental Table S1. Tumor specific networks are enriched with somatically mutated genes (taken from the database COSMIC) COSMIC genes in tumor network COSMIC genes not in tumor network
Chalkou I. C. [PROJECT] Ανάθεση εργασιών.
![Chalkou I. C. [PROJECT] Ανάθεση εργασιών. Chalkou I. C. [PROJECT] Ανάθεση εργασιών.](/thumbs/29/13219375.jpg) Πληροφορική της Υγείας 2014 Chalkou I. C. [PROJECT] Ανάθεση εργασιών. Περιεχόμενα 1. Ομάδα ΣΤ... 3 1.1 ΜΑΡΚΟΠΟΥΛΟΥ- ΣΠΥΡΟΠΟΥΛΟΥ -ΚΩΝΣΤΑΝΤΟΠΟΥΛΟΥ... 3 1.2 ΜΑΡΚΟΣ- ΚΟΥΤΣΟΠΟΥΛΟΣ ΑΥΓΕΡΗ - ΜΠΟΥΖΑΛΑ... 3 1.3
Πληροφορική της Υγείας 2014 Chalkou I. C. [PROJECT] Ανάθεση εργασιών. Περιεχόμενα 1. Ομάδα ΣΤ... 3 1.1 ΜΑΡΚΟΠΟΥΛΟΥ- ΣΠΥΡΟΠΟΥΛΟΥ -ΚΩΝΣΤΑΝΤΟΠΟΥΛΟΥ... 3 1.2 ΜΑΡΚΟΣ- ΚΟΥΤΣΟΠΟΥΛΟΣ ΑΥΓΕΡΗ - ΜΠΟΥΖΑΛΑ... 3 1.3
encouraged to use the Version of Record that, when published, will replace this version. The most /BCJ
 Biochemical Journal: this is an Accepted Manuscript, not the final Version of Record. You are encouraged to use the Version of Record that, when published, will replace this version. The most up-to-date
Biochemical Journal: this is an Accepted Manuscript, not the final Version of Record. You are encouraged to use the Version of Record that, when published, will replace this version. The most up-to-date
Supporting Information. Multigenerational Disruption of Thyroid Endocrine System in Marine Medaka
 Supporting Information Multigenerational Disruption of Thyroid Endocrine System in Marine Medaka after A Life-Cycle Exposure to Perfluorobutane Sulfonate (PFBS) Lianguo Chen,, Chenyan Hu #, Mirabelle M.
Supporting Information Multigenerational Disruption of Thyroid Endocrine System in Marine Medaka after A Life-Cycle Exposure to Perfluorobutane Sulfonate (PFBS) Lianguo Chen,, Chenyan Hu #, Mirabelle M.
JCVI Locus Protein Name. JCVI Role Category JCVI Locus (LT2) Annotation name
 Additional File 4. Detailed information on all 41 genes that showed evidence for positive selection from positive selection analysis, in which gene Alignment Primary Gene Genbank Locus JCVI Locus Protein
Additional File 4. Detailed information on all 41 genes that showed evidence for positive selection from positive selection analysis, in which gene Alignment Primary Gene Genbank Locus JCVI Locus Protein
ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΤΜΗΜΑ ΟΔΟΝΤΙΑΤΡΙΚΗΣ ΕΡΓΑΣΤΗΡΙΟ ΟΔΟΝΤΙΚΗΣ ΚΑΙ ΑΝΩΤΕΡΑΣ ΠΡΟΣΘΕΤΙΚΗΣ
 ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΤΜΗΜΑ ΟΔΟΝΤΙΑΤΡΙΚΗΣ ΕΡΓΑΣΤΗΡΙΟ ΟΔΟΝΤΙΚΗΣ ΚΑΙ ΑΝΩΤΕΡΑΣ ΠΡΟΣΘΕΤΙΚΗΣ ΣΥΓΚΡΙΤΙΚΗ ΜΕΛΕΤΗ ΤΗΣ ΣΥΓΚΡΑΤΗΤΙΚΗΣ ΙΚΑΝΟΤΗΤΑΣ ΟΡΙΣΜΕΝΩΝ ΠΡΟΚΑΤΑΣΚΕΥΑΣΜΕΝΩΝ ΣΥΝΔΕΣΜΩΝ ΑΚΡΙΒΕΙΑΣ
ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΤΜΗΜΑ ΟΔΟΝΤΙΑΤΡΙΚΗΣ ΕΡΓΑΣΤΗΡΙΟ ΟΔΟΝΤΙΚΗΣ ΚΑΙ ΑΝΩΤΕΡΑΣ ΠΡΟΣΘΕΤΙΚΗΣ ΣΥΓΚΡΙΤΙΚΗ ΜΕΛΕΤΗ ΤΗΣ ΣΥΓΚΡΑΤΗΤΙΚΗΣ ΙΚΑΝΟΤΗΤΑΣ ΟΡΙΣΜΕΝΩΝ ΠΡΟΚΑΤΑΣΚΕΥΑΣΜΕΝΩΝ ΣΥΝΔΕΣΜΩΝ ΑΚΡΙΒΕΙΑΣ
Figure S1: Deletion of Dlx3 in the NC A) LacZ staining of Dlx3 LacZ/WT mice (a) and Wnt1- cre:r26r LacZ mice (b) at E11.5. The arrowhead points at
 Figure S1: Deletion of Dlx3 in the NC A) LacZ staining of Dlx3 LacZ/WT mice (a) and Wnt1- cre:r26r LacZ mice (b) at E11.5. The arrowhead points at Dlx3 expression in the branchial arches. B) PCR and Southern
Figure S1: Deletion of Dlx3 in the NC A) LacZ staining of Dlx3 LacZ/WT mice (a) and Wnt1- cre:r26r LacZ mice (b) at E11.5. The arrowhead points at Dlx3 expression in the branchial arches. B) PCR and Southern
RHOA inactivation enhances Wnt signaling and promotes colorectal cancer
 RHOA and colorectal cancer Paulo Rodrigues et al 2014 SUPPLEMENTARY INFORMATION RHOA inactivation enhances Wnt signaling and promotes colorectal cancer Paulo Rodrigues, Irati Macaya, Sarah Bazzocco, Rocco
RHOA and colorectal cancer Paulo Rodrigues et al 2014 SUPPLEMENTARY INFORMATION RHOA inactivation enhances Wnt signaling and promotes colorectal cancer Paulo Rodrigues, Irati Macaya, Sarah Bazzocco, Rocco
«Συντήρηση αχλαδιών σε νερό. υπό την παρουσία σπόρων σιναπιού (Sinapis arvensis).»
 ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΓΕΩΠΟΝΙΚΗ ΣΧΟΛΗ ΤΟΜΕΑΣ ΕΠΙΣΤΗΜΗΣ & ΤΕΧΝΟΛΟΓΙΑΣ ΤΡΟΦΙΜΩΝ ΕΛΕΝΗ Π. ΠΑΠΑΤΣΑΡΟΥΧΑ Πτυχιούχος Τεχνολόγος Τροφίμων της Γεωπονικής Σχολής (Α.Π.Θ.) «Συντήρηση αχλαδιών σε
ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΓΕΩΠΟΝΙΚΗ ΣΧΟΛΗ ΤΟΜΕΑΣ ΕΠΙΣΤΗΜΗΣ & ΤΕΧΝΟΛΟΓΙΑΣ ΤΡΟΦΙΜΩΝ ΕΛΕΝΗ Π. ΠΑΠΑΤΣΑΡΟΥΧΑ Πτυχιούχος Τεχνολόγος Τροφίμων της Γεωπονικής Σχολής (Α.Π.Θ.) «Συντήρηση αχλαδιών σε
Cellular Physiology and Biochemistry
 Original Paper 2016 The Author(s). 2016 Published The Author(s) by S. Karger AG, Basel Published online: November 25, 2016 www.karger.com/cpb Published by S. Karger AG, Basel 486 www.karger.com/cpb Accepted:
Original Paper 2016 The Author(s). 2016 Published The Author(s) by S. Karger AG, Basel Published online: November 25, 2016 www.karger.com/cpb Published by S. Karger AG, Basel 486 www.karger.com/cpb Accepted:
«ΑΓΡΟΤΟΥΡΙΣΜΟΣ ΚΑΙ ΤΟΠΙΚΗ ΑΝΑΠΤΥΞΗ: Ο ΡΟΛΟΣ ΤΩΝ ΝΕΩΝ ΤΕΧΝΟΛΟΓΙΩΝ ΣΤΗΝ ΠΡΟΩΘΗΣΗ ΤΩΝ ΓΥΝΑΙΚΕΙΩΝ ΣΥΝΕΤΑΙΡΙΣΜΩΝ»
 I ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΣΧΟΛΗ ΝΟΜΙΚΩΝ ΟΙΚΟΝΟΜΙΚΩΝ ΚΑΙ ΠΟΛΙΤΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΤΜΗΜΑ ΟΙΚΟΝΟΜΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ ΣΤΗΝ «ΔΙΟΙΚΗΣΗ ΚΑΙ ΟΙΚΟΝΟΜΙΑ» ΚΑΤΕΥΘΥΝΣΗ: ΟΙΚΟΝΟΜΙΚΗ
I ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΣΧΟΛΗ ΝΟΜΙΚΩΝ ΟΙΚΟΝΟΜΙΚΩΝ ΚΑΙ ΠΟΛΙΤΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΤΜΗΜΑ ΟΙΚΟΝΟΜΙΚΩΝ ΕΠΙΣΤΗΜΩΝ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ ΣΤΗΝ «ΔΙΟΙΚΗΣΗ ΚΑΙ ΟΙΚΟΝΟΜΙΑ» ΚΑΤΕΥΘΥΝΣΗ: ΟΙΚΟΝΟΜΙΚΗ
Αξιολόγηση των Φασματικού Διαχωρισμού στην Διάκριση Διαφορετικών Τύπων Εδάφους ΔΙΠΛΩΜΑΤΙΚΗ ΕΡΓΑΣΙΑ. Σπίγγος Γεώργιος
 ΕΘΝΙΚΟ ΜΕΤΣΟΒΙΟ ΠΟΛΥΤΕΧΝΕΙΟ ΤΜΗΜΑ ΑΓΡΟΝΟΜΩΝ ΤΟΠΟΓΡΑΦΩΝ ΜΗΧΑΝΙΚΩΝ ΤΟΜΕΑΣ ΤΟΠΟΓΡΑΦΙΑΣ-ΕΡΓΑΣΤΗΡΙΟ ΤΗΛΕΠΙΣΚΟΠΗΣΗΣ Αξιολόγηση των Φασματικού Διαχωρισμού στην Διάκριση Διαφορετικών Τύπων Εδάφους ΔΙΠΛΩΜΑΤΙΚΗ
ΕΘΝΙΚΟ ΜΕΤΣΟΒΙΟ ΠΟΛΥΤΕΧΝΕΙΟ ΤΜΗΜΑ ΑΓΡΟΝΟΜΩΝ ΤΟΠΟΓΡΑΦΩΝ ΜΗΧΑΝΙΚΩΝ ΤΟΜΕΑΣ ΤΟΠΟΓΡΑΦΙΑΣ-ΕΡΓΑΣΤΗΡΙΟ ΤΗΛΕΠΙΣΚΟΠΗΣΗΣ Αξιολόγηση των Φασματικού Διαχωρισμού στην Διάκριση Διαφορετικών Τύπων Εδάφους ΔΙΠΛΩΜΑΤΙΚΗ
Supporting Information
 Supporting Information Figure S1. The functionality of the tagged Arp6 and Swr1 was confirmed by monitoring cell growth and sensitivity to hydeoxyurea (HU). Five-fold serial dilutions of each strain were
Supporting Information Figure S1. The functionality of the tagged Arp6 and Swr1 was confirmed by monitoring cell growth and sensitivity to hydeoxyurea (HU). Five-fold serial dilutions of each strain were
 ΙΑΤΡΙΚΗ ΣΧΟΛΗ ΤΟΜΕΑΣ ΦΥΣΙΟΛΟΓΙΑΣ ΦΑΡΜΑΚΟΛΟΓΙΑΣ ΕΡΓΑΣΤΗΡΙΟ ΠΕΙΡΑΜΑΤΙΚΗΣ ΦΑΡΜΑΚΟΛΟΓΙΑΣ ΙΕΥΘΥΝΤΡΙΑ: ΚΑΘΗΓΗΤΡΙΑ ΒΑΣΙΛΙΚΗ ΜΗΡΤΣΟΥ-ΦΙ ΑΝΗ ΠΑΝΕΠ. ΕΤΟΣ 2005-2006 ΑΡΙΘΜ. 1844 ΙΕΡΕΥΝΗΣΗ ΤΟΥ ΡΟΛΟΥ ΑΜΙΝΟΞΕΩΝ, ΠΕΠΤΙ
ΙΑΤΡΙΚΗ ΣΧΟΛΗ ΤΟΜΕΑΣ ΦΥΣΙΟΛΟΓΙΑΣ ΦΑΡΜΑΚΟΛΟΓΙΑΣ ΕΡΓΑΣΤΗΡΙΟ ΠΕΙΡΑΜΑΤΙΚΗΣ ΦΑΡΜΑΚΟΛΟΓΙΑΣ ΙΕΥΘΥΝΤΡΙΑ: ΚΑΘΗΓΗΤΡΙΑ ΒΑΣΙΛΙΚΗ ΜΗΡΤΣΟΥ-ΦΙ ΑΝΗ ΠΑΝΕΠ. ΕΤΟΣ 2005-2006 ΑΡΙΘΜ. 1844 ΙΕΡΕΥΝΗΣΗ ΤΟΥ ΡΟΛΟΥ ΑΜΙΝΟΞΕΩΝ, ΠΕΠΤΙ
High genetic similarity between Polish and North European Scots pine (Pinus sylvestris L.) populations at nuclear gene loci
 SUPPLEMENTARY MATERIAL Witold Wachowiak 1,2,3*, Błażej Wόjkiewicz 2, Stephen Cavers 3, Andrzej Lewandowski 2 High genetic similarity between Polish and North European Scots pine (Pinus sylvestris L.) populations
SUPPLEMENTARY MATERIAL Witold Wachowiak 1,2,3*, Błażej Wόjkiewicz 2, Stephen Cavers 3, Andrzej Lewandowski 2 High genetic similarity between Polish and North European Scots pine (Pinus sylvestris L.) populations
ΕΦΑΡΜΟΓΗ ΕΥΤΕΡΟΒΑΘΜΙΑ ΕΠΕΞΕΡΓΑΣΜΕΝΩΝ ΥΓΡΩΝ ΑΠΟΒΛΗΤΩΝ ΣΕ ΦΥΣΙΚΑ ΣΥΣΤΗΜΑΤΑ ΚΛΙΝΗΣ ΚΑΛΑΜΙΩΝ
 ΤΕΧΝΟΛΟΓΙΚΟ ΕΚΠΑΙ ΕΥΤΙΚΟ Ι ΡΥΜΑ ΚΡΗΤΗΣ ΤΜΗΜΑ ΦΥΣΙΚΩΝ ΠΟΡΩΝ ΚΑΙ ΠΕΡΙΒΑΛΛΟΝΤΟΣ ΕΦΑΡΜΟΓΗ ΕΥΤΕΡΟΒΑΘΜΙΑ ΕΠΕΞΕΡΓΑΣΜΕΝΩΝ ΥΓΡΩΝ ΑΠΟΒΛΗΤΩΝ ΣΕ ΦΥΣΙΚΑ ΣΥΣΤΗΜΑΤΑ ΚΛΙΝΗΣ ΚΑΛΑΜΙΩΝ ΕΠΙΜΕΛΕΙΑ: ΑΡΜΕΝΑΚΑΣ ΜΑΡΙΝΟΣ ΧΑΝΙΑ
ΤΕΧΝΟΛΟΓΙΚΟ ΕΚΠΑΙ ΕΥΤΙΚΟ Ι ΡΥΜΑ ΚΡΗΤΗΣ ΤΜΗΜΑ ΦΥΣΙΚΩΝ ΠΟΡΩΝ ΚΑΙ ΠΕΡΙΒΑΛΛΟΝΤΟΣ ΕΦΑΡΜΟΓΗ ΕΥΤΕΡΟΒΑΘΜΙΑ ΕΠΕΞΕΡΓΑΣΜΕΝΩΝ ΥΓΡΩΝ ΑΠΟΒΛΗΤΩΝ ΣΕ ΦΥΣΙΚΑ ΣΥΣΤΗΜΑΤΑ ΚΛΙΝΗΣ ΚΑΛΑΜΙΩΝ ΕΠΙΜΕΛΕΙΑ: ΑΡΜΕΝΑΚΑΣ ΜΑΡΙΝΟΣ ΧΑΝΙΑ
Growth Factors/Wachstumsfaktoren. Human (Hu) Cytokines and Growth Factors/Humane (Hu) Cytokine und Wachstumsfaktoren
 Growth Factors/Wachstumsfaktoren Human (Hu) Cytokines and Growth Factors/Humane (Hu) Cytokine und Wachstumsfaktoren 4-1BBr Hu 4-1BB Receptor rec. CB-1010100 5 μg CB-1010101 20 μg CB-1010102 1 mg Acrp30
Growth Factors/Wachstumsfaktoren Human (Hu) Cytokines and Growth Factors/Humane (Hu) Cytokine und Wachstumsfaktoren 4-1BBr Hu 4-1BB Receptor rec. CB-1010100 5 μg CB-1010101 20 μg CB-1010102 1 mg Acrp30
Structural and Biochemical Studies of Cotranslational Protein Transport across the Endoplasmic Reticulum
 Structural and Biochemical Studies of Cotranslational Protein Transport across the Endoplasmic Reticulum Inaugural-Dissertation zur Erlangung des akademischen Grades Doktor der Naturwissenschaften - Dr.
Structural and Biochemical Studies of Cotranslational Protein Transport across the Endoplasmic Reticulum Inaugural-Dissertation zur Erlangung des akademischen Grades Doktor der Naturwissenschaften - Dr.
Απαραίτητες προϋποθέσεις για την επιτυχία των μεταμοσχεύσεων
 Απαραίτητες προϋποθέσεις για την επιτυχία των μεταμοσχεύσεων Αικατερίνη ΣταυροπουΛου-Γκιόκα Διευθύντρια Ανοσολογικού-Εθνικού Κέντρου Ιστοσυμβατότητας, ΓΠΝ Αθηνών «Γεώργιος Γεννηματάς» μεταμόσχευση οργάνων
Απαραίτητες προϋποθέσεις για την επιτυχία των μεταμοσχεύσεων Αικατερίνη ΣταυροπουΛου-Γκιόκα Διευθύντρια Ανοσολογικού-Εθνικού Κέντρου Ιστοσυμβατότητας, ΓΠΝ Αθηνών «Γεώργιος Γεννηματάς» μεταμόσχευση οργάνων
Supplementary figures for MED13 dependent signaling from the heart confers leanness by enhancing metabolism in adipose tissue and liver.
 Supplementary figures for MED13 dependent signaling from the heart confers leanness by enhancing metabolism in adipose tissue and liver. Content: Figure S1. Cardiac overexpression of MED13 increases metabolic
Supplementary figures for MED13 dependent signaling from the heart confers leanness by enhancing metabolism in adipose tissue and liver. Content: Figure S1. Cardiac overexpression of MED13 increases metabolic
encouraged to use the Version of Record that, when published, will replace this version. The most /BCJ
 Biochemical Journal: this is an Accepted Manuscript, not the final Version of Record. You are encouraged to use the Version of Record that, when published, will replace this version. The most up-to-date
Biochemical Journal: this is an Accepted Manuscript, not the final Version of Record. You are encouraged to use the Version of Record that, when published, will replace this version. The most up-to-date
JCVI Role Category_1 JCVI Role Category_2. histidinol phosphatase-related protein Unknown function NT01ST0303 AAL19211 STM0248
 Additional Table 3. Detailed information on all 81 genes that showed evidence for positive selection from initial positive selection analysis, in Protein Name Gene name JCVI Role Category_1 JCVI Role Category_2
Additional Table 3. Detailed information on all 81 genes that showed evidence for positive selection from initial positive selection analysis, in Protein Name Gene name JCVI Role Category_1 JCVI Role Category_2
[1] P Q. Fig. 3.1
![[1] P Q. Fig. 3.1 [1] P Q. Fig. 3.1](/thumbs/79/80362156.jpg) 1 (a) Define resistance....... [1] (b) The smallest conductor within a computer processing chip can be represented as a rectangular block that is one atom high, four atoms wide and twenty atoms long. One
1 (a) Define resistance....... [1] (b) The smallest conductor within a computer processing chip can be represented as a rectangular block that is one atom high, four atoms wide and twenty atoms long. One
Chinese Bulletin of Life Sciences. NTAL/LAB: immuno-regulatory role in lymphocyte development and function. WANG Ying
 17 3 2005 6 Chinese Bulletin of Life Sciences Vol. 17, No. 3 Jun., 2005 1004-0374(2005)03-0251-05 NTAL/LAB 200025 NTAL/LAB B FcγRI FcεRI NTAL/LAB NTAL/LAB ; NTAL/LAB; Q25; Q51 A NTAL/LAB: immuno-regulatory
17 3 2005 6 Chinese Bulletin of Life Sciences Vol. 17, No. 3 Jun., 2005 1004-0374(2005)03-0251-05 NTAL/LAB 200025 NTAL/LAB B FcγRI FcεRI NTAL/LAB NTAL/LAB ; NTAL/LAB; Q25; Q51 A NTAL/LAB: immuno-regulatory
Mouse Gene 2.0 ST Array Shn3. Runx2 Shn3. Runx2 Shn3 Runx2. adult. (Schnurri, Shn)3/Hivep3. Runx2 Shn/Hivep. ST2 BMP-2 Shn3
 adult (Schnurri, Shn)3/Hivep3 Runx2 Shn/Hivep Shn3 Shn3 Shn3 Shn3 Shn1 Shn2 Shn3 gain-of-function loss-of-function Shn3 in vivo mimic MC3T3-E1 ST-2 ATDC5 C3H10T1/2 ALP RT-PCR alcian blue RT-PCR Affymetrix
adult (Schnurri, Shn)3/Hivep3 Runx2 Shn/Hivep Shn3 Shn3 Shn3 Shn3 Shn1 Shn2 Shn3 gain-of-function loss-of-function Shn3 in vivo mimic MC3T3-E1 ST-2 ATDC5 C3H10T1/2 ALP RT-PCR alcian blue RT-PCR Affymetrix
Gene Primer sequence (5 to 3 ) Size (bp) E (%) GenBank
 Table S1 Primers used for quantitative real time PCR Gene Primer sequence (5 to 3 ) Size (bp) E (%) GenBank accession no: scd a got2a a ube2h a gpx4b a mknk2b a hsd11b2 b eef1a c b2m d F: ACCCGGAAGTCATCGAGAGA
Table S1 Primers used for quantitative real time PCR Gene Primer sequence (5 to 3 ) Size (bp) E (%) GenBank accession no: scd a got2a a ube2h a gpx4b a mknk2b a hsd11b2 b eef1a c b2m d F: ACCCGGAAGTCATCGAGAGA
Jesse Maassen and Mark Lundstrom Purdue University November 25, 2013
 Notes on Average Scattering imes and Hall Factors Jesse Maassen and Mar Lundstrom Purdue University November 5, 13 I. Introduction 1 II. Solution of the BE 1 III. Exercises: Woring out average scattering
Notes on Average Scattering imes and Hall Factors Jesse Maassen and Mar Lundstrom Purdue University November 5, 13 I. Introduction 1 II. Solution of the BE 1 III. Exercises: Woring out average scattering
Supplementary Table 1. Construct List with key Biophysical Properties of the expression
 SPINE Benchmark Target ID Well Code Tag (N or C) Fusion MW (kda) Fusion pi Cleavage site Prot MW (Da) Prot pi 1 A1 OPPF 2585 N-His6 21.75 6.41 Protease 3C 19.77 5.43 2 B1 OPPF 2586 N-His6 15.6 6.27 Protease
SPINE Benchmark Target ID Well Code Tag (N or C) Fusion MW (kda) Fusion pi Cleavage site Prot MW (Da) Prot pi 1 A1 OPPF 2585 N-His6 21.75 6.41 Protease 3C 19.77 5.43 2 B1 OPPF 2586 N-His6 15.6 6.27 Protease
CHAPTER 25 SOLVING EQUATIONS BY ITERATIVE METHODS
 CHAPTER 5 SOLVING EQUATIONS BY ITERATIVE METHODS EXERCISE 104 Page 8 1. Find the positive root of the equation x + 3x 5 = 0, correct to 3 significant figures, using the method of bisection. Let f(x) =
CHAPTER 5 SOLVING EQUATIONS BY ITERATIVE METHODS EXERCISE 104 Page 8 1. Find the positive root of the equation x + 3x 5 = 0, correct to 3 significant figures, using the method of bisection. Let f(x) =
Contents Part I Psychoneuroimmunology and Systems Biology Mechanisms 1 From Psychoneuroimmunology to Personalized, Systems, and Dynamical Medicine
 Contents Part I Psychoneuroimmunology and Systems Biology Mechanisms 1 From Psychoneuroimmunology to Personalized, Systems, and Dynamical Medicine... 3 1.1 Psychoneuroimmunology (PNI) and Systems Biology...
Contents Part I Psychoneuroimmunology and Systems Biology Mechanisms 1 From Psychoneuroimmunology to Personalized, Systems, and Dynamical Medicine... 3 1.1 Psychoneuroimmunology (PNI) and Systems Biology...
Example Sheet 3 Solutions
 Example Sheet 3 Solutions. i Regular Sturm-Liouville. ii Singular Sturm-Liouville mixed boundary conditions. iii Not Sturm-Liouville ODE is not in Sturm-Liouville form. iv Regular Sturm-Liouville note
Example Sheet 3 Solutions. i Regular Sturm-Liouville. ii Singular Sturm-Liouville mixed boundary conditions. iii Not Sturm-Liouville ODE is not in Sturm-Liouville form. iv Regular Sturm-Liouville note
ΕΘΝΙΚΟ ΜΕΤΣΟΒΙΟ ΠΟΛΥΤΕΧΝΕΙΟ
 ΕΘΝΙΚΟ ΜΕΤΣΟΒΙΟ ΠΟΛΥΤΕΧΝΕΙΟ ΣΧΟΛΗ ΗΛΕΚΤΡΟΛΟΓΩΝ ΜΗΧΑΝΙΚΩΝ ΚΑΙ ΜΗΧΑΝΙΚΩΝ ΥΠΟΛΟΓΙΣΤΩΝ ΤΟΜΕΑΣ ΗΛΕΚΤΡΟΜΑΓΝΗΤΙΚΩΝ ΕΦΑΡΜΟΓΩΝ ΗΛΕΚΤΡΟΟΠΤΙΚΗΣ & ΗΛΕΚΤΡΟΝΙΚΩΝ ΥΛΙΚΩΝ Μελέτη Επίδρασης Υπεριώδους Ακτινοβολίας σε Λεπτά
ΕΘΝΙΚΟ ΜΕΤΣΟΒΙΟ ΠΟΛΥΤΕΧΝΕΙΟ ΣΧΟΛΗ ΗΛΕΚΤΡΟΛΟΓΩΝ ΜΗΧΑΝΙΚΩΝ ΚΑΙ ΜΗΧΑΝΙΚΩΝ ΥΠΟΛΟΓΙΣΤΩΝ ΤΟΜΕΑΣ ΗΛΕΚΤΡΟΜΑΓΝΗΤΙΚΩΝ ΕΦΑΡΜΟΓΩΝ ΗΛΕΚΤΡΟΟΠΤΙΚΗΣ & ΗΛΕΚΤΡΟΝΙΚΩΝ ΥΛΙΚΩΝ Μελέτη Επίδρασης Υπεριώδους Ακτινοβολίας σε Λεπτά
3.5.4 Ο αυξητικός παράγοντας BMP... 24 3.5.5 Ο αυξητικός παράγοντας FGF... 25 3.5.6 Αυξητικός παράγοντας PDGF (Platelet Derived Growth Factor)...
 Ευχαριστίες Θα ήθελα να ευχαριστήσω τον επιβλέποντα καθηγητή Κο Χριστόφορο Προβατίδη για την αμέριστη υποστήριξη που μου προσέφερε και τις πολύτιμες γνώσεις που μου μετέδωσε κατά την εκπόνηση της Μεταπτυχιακής
Ευχαριστίες Θα ήθελα να ευχαριστήσω τον επιβλέποντα καθηγητή Κο Χριστόφορο Προβατίδη για την αμέριστη υποστήριξη που μου προσέφερε και τις πολύτιμες γνώσεις που μου μετέδωσε κατά την εκπόνηση της Μεταπτυχιακής
Abstract... I. Zusammenfassung... II. 1 Aim of the work Introduction Short overview of Chinese hamster ovary cell lines...
 III Contents I II III Abstract... I Zusammenfassung... II Contents... III 1 Aim of the work... 1 2 Introduction... 2 2.1 Short overview of Chinese hamster ovary cell lines... 2 2.2 Unspecific mutations:
III Contents I II III Abstract... I Zusammenfassung... II Contents... III 1 Aim of the work... 1 2 Introduction... 2 2.1 Short overview of Chinese hamster ovary cell lines... 2 2.2 Unspecific mutations:
Chapter 1. Fingolimod attenuates ceramide induced blood-brain barrier dysfunction in multiple sclerosis by targeting reactive astrocytes
 Chapter 1 Fingolimod attenuates ceramide induced blood-brain barrier dysfunction in multiple sclerosis by targeting reactive astrocytes 1 1 1 1 CHAPTER SUMMARY FINGOLIMOD ATTENUATES CERAMIDE PRODUCTION
Chapter 1 Fingolimod attenuates ceramide induced blood-brain barrier dysfunction in multiple sclerosis by targeting reactive astrocytes 1 1 1 1 CHAPTER SUMMARY FINGOLIMOD ATTENUATES CERAMIDE PRODUCTION
http / / cjbmb. bjmu. edu. cn Chinese Journal of Biochemistry and Molecular Biology Gab2 SHP2 / Ras / ERK PI3K / AKT Gab2 Signaling in Breast Cancer
 ISSN 1007-7626 CN 11-3870 / Q http / / cjbmb bjmu edu cn Chinese Journal of Biochemistry and Molecular Biology 2011 4 27 4 300 ~ 304 Gab2 * 310058 Gab2 Gabs Gab2 SH2 SHP2 / Ras / ERK PI3K / AKT GAB2 ErbB2
ISSN 1007-7626 CN 11-3870 / Q http / / cjbmb bjmu edu cn Chinese Journal of Biochemistry and Molecular Biology 2011 4 27 4 300 ~ 304 Gab2 * 310058 Gab2 Gabs Gab2 SH2 SHP2 / Ras / ERK PI3K / AKT GAB2 ErbB2
Sar1B-mediated Regulation of Triglyceride and Cholesterol
 FIGURE S1. Comparing Sar1A- and Sar1B-flag Over-expression on RNA Profiles of McArdle RH7777 cells. RNA was prepared from two Sar1B:H79G-flag cell lines and two control cell lines plus two Sar1B-flag and
FIGURE S1. Comparing Sar1A- and Sar1B-flag Over-expression on RNA Profiles of McArdle RH7777 cells. RNA was prepared from two Sar1B:H79G-flag cell lines and two control cell lines plus two Sar1B-flag and
In vitro και in vivo φαρμακοκινητική ανάλυση των παραγώγων ανθρακινόνης σε φυτικά σκευάσματα
 ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΤΜΗΜΑ ΦΑΡΜΑΚΕΥΤΙΚΗΣ ΤΟΜΕΑΣ ΦΑΡΜΑΚΟΓΝΩΣΙΑΣ ΦΑΡΜΑΚΟΛΟΓΙΑΣ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ ΚΑΤΕΥΘΥΝΣΗ ΦΑΡΜΑΚΟΛΟΓΙΑ ΚΑΙ ΘΕΡΑΠΕΥΤΙΚΗ In vitro και in vivo φαρμακοκινητική
ΑΡΙΣΤΟΤΕΛΕΙΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΟΝΙΚΗΣ ΤΜΗΜΑ ΦΑΡΜΑΚΕΥΤΙΚΗΣ ΤΟΜΕΑΣ ΦΑΡΜΑΚΟΓΝΩΣΙΑΣ ΦΑΡΜΑΚΟΛΟΓΙΑΣ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ ΚΑΤΕΥΘΥΝΣΗ ΦΑΡΜΑΚΟΛΟΓΙΑ ΚΑΙ ΘΕΡΑΠΕΥΤΙΚΗ In vitro και in vivo φαρμακοκινητική
19 Ενδοκρινολόγος. Φάρµακα που στοχεύουν το β- κύτταρο ΕΥΡΥΔΙΚΗ ΠΑΠΑΔΟΠΟΥΛΟΥ ΕΙΣΑΓΩΓΗ ΜΗΧΑΝΙΣΜΟΣ ΔΡΑΣΗΣ
 Φάρµακα που στοχεύουν το β- κύτταρο ΕΥΡΥΔΙΚΗ ΠΑΠΑΔΟΠΟΥΛΟΥ 19 Ενδοκρινολόγος ΕΙΣΑΓΩΓΗ Οι σουλφονυλουρίες είναι η πρώτη κατηγορία υπογλυκαιμικών φαρμάκων που χορηγήθηκαν για την αντιμετώπιση του διαβήτη
Φάρµακα που στοχεύουν το β- κύτταρο ΕΥΡΥΔΙΚΗ ΠΑΠΑΔΟΠΟΥΛΟΥ 19 Ενδοκρινολόγος ΕΙΣΑΓΩΓΗ Οι σουλφονυλουρίες είναι η πρώτη κατηγορία υπογλυκαιμικών φαρμάκων που χορηγήθηκαν για την αντιμετώπιση του διαβήτη
 22 .5 Real consumption.5 Real residential investment.5.5.5 965 975 985 995 25.5 965 975 985 995 25.5 Real house prices.5 Real fixed investment.5.5.5 965 975 985 995 25.5 965 975 985 995 25.3 Inflation
22 .5 Real consumption.5 Real residential investment.5.5.5 965 975 985 995 25.5 965 975 985 995 25.5 Real house prices.5 Real fixed investment.5.5.5 965 975 985 995 25.5 965 975 985 995 25.3 Inflation
Figure S1. Localization of GiOR-1 with truncated N-terminal domain of 29 amino acid residues (ΔL-GiOR-1-HA).
 Figure S1. Localization of GiOR-1 with truncated N-terminal domain of 29 amino acid residues (ΔL-GiOR-1-HA). A L-GiOR-1-HA Tom40 DAPI merge DIC B LYS CYT ORG ORG Trypsin - - - + L-GiOR-1-HA Figure S2A.
Figure S1. Localization of GiOR-1 with truncated N-terminal domain of 29 amino acid residues (ΔL-GiOR-1-HA). A L-GiOR-1-HA Tom40 DAPI merge DIC B LYS CYT ORG ORG Trypsin - - - + L-GiOR-1-HA Figure S2A.
Εισαγωγή στις πρωτεΐνες Δομή πρωτεϊνών Ταξινόμηση βάσει δομής Βάσεις με δομές πρωτεϊνών Ευθυγράμμιση δομών Πρόβλεψη 2D δομής Πρόβλεψη 3D δομής
 Εισαγωγή στις πρωτεΐνες Δομή πρωτεϊνών Ταξινόμηση βάσει δομής Βάσεις με δομές πρωτεϊνών Ευθυγράμμιση δομών Πρόβλεψη 2D δομής Πρόβλεψη 3D δομής Τι είναι η πρωτεΐνη Τι εννοούμε με δομή πρωτεϊνών Οικογένειες
Εισαγωγή στις πρωτεΐνες Δομή πρωτεϊνών Ταξινόμηση βάσει δομής Βάσεις με δομές πρωτεϊνών Ευθυγράμμιση δομών Πρόβλεψη 2D δομής Πρόβλεψη 3D δομής Τι είναι η πρωτεΐνη Τι εννοούμε με δομή πρωτεϊνών Οικογένειες
Asrani et al Supplementary Figure S1: Epidermal-specific mtorc1 loss-of-function models show impaired epidermal differentiation.
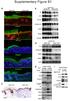 Asrani et al Supplementary Figure S1: Epidermal-specific mtorc1 loss-of-function models show impaired epidermal differentiation. (A) Rheb cko P0 epidermis fails to express spinous (Keratin 1, 10) and granular
Asrani et al Supplementary Figure S1: Epidermal-specific mtorc1 loss-of-function models show impaired epidermal differentiation. (A) Rheb cko P0 epidermis fails to express spinous (Keratin 1, 10) and granular
Figures Supplementary Figure 1. Tandem MS of yeast eif2b.
 Figures Supplementary Figure 1. Tandem MS of yeast eif2b. Tandem MS of the 56+ charge state (shaded blue) of the eif2b dimer showing that two α subunits ( at 2000 m/z) dissociate from the dimer when collision
Figures Supplementary Figure 1. Tandem MS of yeast eif2b. Tandem MS of the 56+ charge state (shaded blue) of the eif2b dimer showing that two α subunits ( at 2000 m/z) dissociate from the dimer when collision
Electronic Supplementary Information (ESI)
 Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2016 Electronic Supplementary Information (ESI) Cyclopentadienyl iron dicarbonyl (CpFe(CO) 2 ) derivatives
Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2016 Electronic Supplementary Information (ESI) Cyclopentadienyl iron dicarbonyl (CpFe(CO) 2 ) derivatives
Science of Sericulture
 Science of Sericulture 2012 38 1 0065-0069 ISSN 0257-4799 CN 32-1115 /S E-mail CYKE@ chinajournal. net. cn 215123 25 0 ~ 24 h GSH GSSG GSH + 2GSSG GSH /GSSG S- GST TPX 23% GR 57% GSH TPX GST GSSG 61% GSH
Science of Sericulture 2012 38 1 0065-0069 ISSN 0257-4799 CN 32-1115 /S E-mail CYKE@ chinajournal. net. cn 215123 25 0 ~ 24 h GSH GSSG GSH + 2GSSG GSH /GSSG S- GST TPX 23% GR 57% GSH TPX GST GSSG 61% GSH
ΓΕΩΠΟΝΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΑΘΗΝΩΝ ΤΜΗΜΑ ΑΓΡΟΤΙΚΗΣ ΟΙΚΟΝΟΜΙΑΣ & ΑΝΑΠΤΥΞΗΣ
 ΓΕΩΠΟΝΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΑΘΗΝΩΝ ΤΜΗΜΑ ΑΓΡΟΤΙΚΗΣ ΟΙΚΟΝΟΜΙΑΣ & ΑΝΑΠΤΥΞΗΣ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ «ΟΛΟΚΛΗΡΩΜΕΝΗ ΑΝΑΠΤΥΞΗ & ΔΙΑΧΕΙΡΙΣΗ ΤΟΥ ΑΓΡΟΤΙΚΟΥ ΧΩΡΟΥ» ΜΕΤΑΠΤΥΧΙΑΚΗ ΕΡΓΑΣΙΑ «Οικονομετρική διερεύνηση
ΓΕΩΠΟΝΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΑΘΗΝΩΝ ΤΜΗΜΑ ΑΓΡΟΤΙΚΗΣ ΟΙΚΟΝΟΜΙΑΣ & ΑΝΑΠΤΥΞΗΣ ΠΡΟΓΡΑΜΜΑ ΜΕΤΑΠΤΥΧΙΑΚΩΝ ΣΠΟΥΔΩΝ «ΟΛΟΚΛΗΡΩΜΕΝΗ ΑΝΑΠΤΥΞΗ & ΔΙΑΧΕΙΡΙΣΗ ΤΟΥ ΑΓΡΟΤΙΚΟΥ ΧΩΡΟΥ» ΜΕΤΑΠΤΥΧΙΑΚΗ ΕΡΓΑΣΙΑ «Οικονομετρική διερεύνηση
Linking FOXO3, NCOA3, and TCF7L2 to Ras pathway phenotypes through a genomewide forward genetic screen in human colorectal cancer cells
 1 2 3 4 5 6 7 8 9 Linking FOXO3, NCOA3, and TCF7L2 to Ras pathway phenotypes through a genomewide forward genetic screen in human colorectal cancer cells Authors: Snehangshu Kundu 1, Muhammad Akhtar Ali
1 2 3 4 5 6 7 8 9 Linking FOXO3, NCOA3, and TCF7L2 to Ras pathway phenotypes through a genomewide forward genetic screen in human colorectal cancer cells Authors: Snehangshu Kundu 1, Muhammad Akhtar Ali
SEN TRONIC AG 3-2 7 0 0 7 A 3 57 3 3 AB 93 :, C,! D 0 7 % 0 7 3 3 93 : 3 A 5 93 :
 # 3-270 07A35733 AB93:,C,!D 07% 0733 93: 3A593:!"#$%% &%&''()*%'+,-. &%&''(/*%'+0. 1*23 '4# 54/%6%7%53 *323 %7 77# %%3#% 8908/"/*55 :1$;/ = 7?@ > 7= 7 %! "$!"#$%&#%'(%%)*#$%&#%'(%#++#,-."/-0-1222"/-0-1
# 3-270 07A35733 AB93:,C,!D 07% 0733 93: 3A593:!"#$%% &%&''()*%'+,-. &%&''(/*%'+0. 1*23 '4# 54/%6%7%53 *323 %7 77# %%3#% 8908/"/*55 :1$;/ = 7?@ > 7= 7 %! "$!"#$%&#%'(%%)*#$%&#%'(%#++#,-."/-0-1222"/-0-1
UDZ Swirl diffuser. Product facts. Quick-selection. Swirl diffuser UDZ. Product code example:
 UDZ Swirl diffuser Swirl diffuser UDZ, which is intended for installation in a ventilation duct, can be used in premises with a large volume, for example factory premises, storage areas, superstores, halls,
UDZ Swirl diffuser Swirl diffuser UDZ, which is intended for installation in a ventilation duct, can be used in premises with a large volume, for example factory premises, storage areas, superstores, halls,
I ΙΑΙΤΕΡΟΤΗΤΕΣ ΤΗΣ ΙΑΠΙΣΤΕΥΣΗΣ ΤΩΝ ΜΙΚΡΟΒΙΟΛΟΓΙΚΩΝ ΕΡΓΑΣΤΗΡΙΩΝ ΝΕΡΟΥ ΚΑΤΑ ISO 17025. ΠΗΓΕΣ ΑΝΟΜΟΙΟΓΕΝΕΙΑΣ ΚΑΙ ΛΑΘΩΝ
 I ΙΑΙΤΕΡΟΤΗΤΕΣ ΤΗΣ ΙΑΠΙΣΤΕΥΣΗΣ ΤΩΝ ΜΙΚΡΟΒΙΟΛΟΓΙΚΩΝ ΕΡΓΑΣΤΗΡΙΩΝ ΝΕΡΟΥ ΚΑΤΑ ISO 17025. Ρ. Αθηνά Μαυρίδου Καθ. Μικροβιολογίας ΤΕΙ Αθήνας ΠΗΓΕΣ ΑΝΟΜΟΙΟΓΕΝΕΙΑΣ ΚΑΙ ΛΑΘΩΝ Φυσική ανοµοιογένεια νερού στην προέλευση
I ΙΑΙΤΕΡΟΤΗΤΕΣ ΤΗΣ ΙΑΠΙΣΤΕΥΣΗΣ ΤΩΝ ΜΙΚΡΟΒΙΟΛΟΓΙΚΩΝ ΕΡΓΑΣΤΗΡΙΩΝ ΝΕΡΟΥ ΚΑΤΑ ISO 17025. Ρ. Αθηνά Μαυρίδου Καθ. Μικροβιολογίας ΤΕΙ Αθήνας ΠΗΓΕΣ ΑΝΟΜΟΙΟΓΕΝΕΙΑΣ ΚΑΙ ΛΑΘΩΝ Φυσική ανοµοιογένεια νερού στην προέλευση
Τα ηπατικά επίπεδα του FOXP3 mrna στη χρόνια ηπατίτιδα Β εξαρτώνται από την έκφραση των οδών Fas/FasL και PD-1/PD-L1
 Τα ηπατικά επίπεδα του FOXP3 mrna στη χρόνια ηπατίτιδα Β εξαρτώνται από την έκφραση των οδών Fas/FasL και PD-1/PD-L1 Γ. Γερμανίδης, Ν. Αργέντου, Θ. Βασιλειάδης, Κ. Πατσιαούρα, Κ. Μαντζούκης, A. Θεοχαρίδου,
Τα ηπατικά επίπεδα του FOXP3 mrna στη χρόνια ηπατίτιδα Β εξαρτώνται από την έκφραση των οδών Fas/FasL και PD-1/PD-L1 Γ. Γερμανίδης, Ν. Αργέντου, Θ. Βασιλειάδης, Κ. Πατσιαούρα, Κ. Μαντζούκης, A. Θεοχαρίδου,
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Valk PJM, Verhaak RGW, Beijen MA, et al. Prognostically Useful
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Valk PJM, Verhaak RGW, Beijen MA, et al. Prognostically Useful
Group 2 Methotrexate 7.5 mg/week, increased to 15 mg/week after 4 weeks. Methotrexate 7.5 mg/week, increased to 15 mg/week after 4 weeks
 Group 1 Methotrexate 7.5 mg/week, increased to 15 mg/week after 4 weeks Group 2 Methotrexate 7.5 mg/week, increased to 15 mg/week after 4 weeks Sulfasalazine 2000-3000 mg/day Leflunomide 20 mg/day Infliximab
Group 1 Methotrexate 7.5 mg/week, increased to 15 mg/week after 4 weeks Group 2 Methotrexate 7.5 mg/week, increased to 15 mg/week after 4 weeks Sulfasalazine 2000-3000 mg/day Leflunomide 20 mg/day Infliximab
Μελέτη της αντιμεταλλαξιγόνου δράσης φλαβονοειδών του φυτού Lotus Edulis με τη μέθοδο Ames test
 ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΙΑΣ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΥΓΕΙΑΣ ΤΜΗΜΑ ΒΙΟΧΗΜΕΙΑΣ & ΒΙΟΤΕΧΝΟΛΟΓΙΑΣ ΔΙΠΛΩΜΑΤΙΚΗ ΕΡΓΑΣΙΑ Μελέτη της αντιμεταλλαξιγόνου δράσης φλαβονοειδών του φυτού Lotus Edulis με τη μέθοδο Ames test Επιμέλεια
ΠΑΝΕΠΙΣΤΗΜΙΟ ΘΕΣΣΑΛΙΑΣ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΥΓΕΙΑΣ ΤΜΗΜΑ ΒΙΟΧΗΜΕΙΑΣ & ΒΙΟΤΕΧΝΟΛΟΓΙΑΣ ΔΙΠΛΩΜΑΤΙΚΗ ΕΡΓΑΣΙΑ Μελέτη της αντιμεταλλαξιγόνου δράσης φλαβονοειδών του φυτού Lotus Edulis με τη μέθοδο Ames test Επιμέλεια
